Abstract
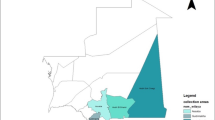
The objective of study aimed to analyze the genetic diversity and genetic structure of Moroccan landraces of sorghum (Sorghum bicolor L. Moench). Samples were obtained from 34 fields in the northern regions, specifically from Larache (13 fields), Tangier (11 fields), Chefchaouen (5 fields), and Tetouan (5 fields). A total of 398 individual plants were collected, including three accessions from each of the five cultivated races of sorghum (bicolor, durra, caudatum, guinea, and kafir) from the World Collection. Two molecular marker methods, inter-simple sequence repeat (ISSR) and amplified fragment length polymorphism (AFLP), were employed. Four ISSR primers produced 185 reproducible bands, of which 177 (96%) exhibited polymorphism. Three AFLP primer combinations were used to detect 226 markers, out of which 198 (88%) were polymorphic. The average polymorphic information (PIC) values for ISSR and AFLP markers were calculated to be 0.98 and 0.97, respectively. Both markers revealed a higher relative genetic differentiation, with values of 0.5595 (ISSR) and 0.6216 (AFLP), while the gene flow (Nm) between fields was determined to be 0.3937 (ISSR) and 0.3043 (AFLP). Analysis of Molecular Variance (AMOVA) confirmed a greater variation within the field (57%). Classification of the Moroccan landraces based on Jaccard’s similarity index led to the identification of two primary genetic pools, which was further supported by the unweighted pair group method with arithmetic mean (UPGMA) clustering and STRUCTURE analysis. The Mantel test revealed lack of correlation between genetic structure and microgeographical distribution. The ISSR and AFLP data suggest that the bicolor, durra, and caudatum races are more genetically related to the Moroccan landraces of sorghum than kafir and guinea.








Similar content being viewed by others
References
Adugna A (2014) Analysis of in situ diversity and population structure in Ethiopian cultivated Sorghum bicolor (L.) landraces using phenotypic traits and SSR markers. Springerplus 3(1):1–14
Al-Turki T, Basahi MA (2015) Assessment of ISSR based molecular genetic diversity of Hassawi rice in Saudi Arabia. Saudi J Biol Sci 22(5):591–599
Bao J, Corke H, Sun M (2006) Analysis of genetic diversity and relationships in waxy rice (Oryza sativa L.) using AFLP and ISSR markers. Genet Resour Crop Evol 53:323–330
Barrett B, Kidwell K (1998) AFLP-based genetic diversity assessment among wheat cultivars from the Pacific Northwest. Crop Sci 38(5):1261–1271
Basahi M (2015) ISSR-based analysis of genetic diversity among sorghum landraces growing in some parts of Saudi Arabia and Yemen. C R Biol 338(11):723–727. https://doi.org/10.1016/j.crvi.2015.09.003
Bernatzky R, Tanksley SD (1986) Toward a saturated linkage map in tomato based on isozymes and random cDNA sequences. Genetics 112(4):887–898. https://doi.org/10.1093/genetics/112.4.887
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32(3):314–331
Bucheyeki TL, Gwanama C, Mgonja M, Chisi M, Folkertsma R, Mutegi R (2009) Genetic variability characterisation of Tanzania sorghum landraces based on simple sequence repeats (SSRs) molecular and morphological markers. Afr Crop Sci J 17(2):71–86
Can ND, Yoshida T (1999) Genotypic and phenotypic variances and covariances in early maturing grain sorghum in a double cropping. Plant Prod Sci 2(1):67–70
Chakraborty S, Thakare I, Ravikiran R, Nikam V, Trivedi R, Sasidharan N, Jadeja G (2011) Assessment of diversity using RAPD and ISSR markers in Sorghum varieties across Gujarat, India. Electron J Plant Breed 2(4):488–493
Cuevas HE, Rosa-Valentin G, Hayes CM, Rooney WL, Hoffmann L (2017) Genomic characterization of a core set of the USDA-NPGS Ethiopian sorghum germplasm collection: implications for germplasm conservation, evaluation, and utilization in crop improvement. BMC Genom 18(1):108. https://doi.org/10.1186/s12864-016-3475-7
Deu M, Hamon P, Dufour P, D’hont A, Lanaud C, Chantereau J (1995) Mitochondrial DNA diversity in wild and cultivated sorghum. Genome 38(4):635–645
Djè Y, Ater M, Lefèbvre C, Vekemans X (1998) Patterns of morphological and allozyme variation in sorghum landraces of Northwestern Morocco. Genet Resour Crop Evol 45:541–548
Djè Y, Forcioli D, Ater M, Lefèbvre C, Vekemans X (1999) Assessing population genetic structure of sorghum landraces from North-western Morocco using allozyme and microsatellite markers. Theor Appl Genet 99:157–163
Djè Y, Heuertz M, Lefebvre C, Vekemans X (2000) Assessment of genetic diversity within and among germplasm accessions in cultivated sorghum using microsatellite markers. Theor Appl Genet 100:918–925
Djè Y, Heuertz M, Ater M, Lefèbvre C, Vekemans X (2004) In situ estimation of outcrossing rate in sorghum landraces using microsatellite markers. Euphytica Neth J Plant Breed 138:205–212
do Amaral Júnior AT, de Oliveira ÉC, Gonçalves LSA, Scapim CA, Candido LS, da Conceição Silva TR et al (2011) Assessment of genetic diversity among maize accessions using inter simple sequence repeats (ISSR) markers. Afr J Biotechnol 10(69):15462–15469
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4(2):359–361. https://doi.org/10.1007/s12686-011-9548-7
Elangovan M, Kiran Babu P, Seetharama N, Patil J (2014) Genetic diversity and heritability characters associated in sweet sorghum [Sorghum bicolor (L.) Moench]. Sugar Tech Int J Sugar Crops Relat Ind 16(2):200–210
Enyew M, Feyissa T, Carlsson AS, Tesfaye K, Hammenhag C, Geleta M (2022) Genetic diversity and population structure of sorghum [Sorghum bicolor (L.) moench] accessions as revealed by single nucleotide polymorphism markers. Front Plant Sci 12:3110
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software structure: a simulation study. Mol Ecol 14(8):2611–2620. https://doi.org/10.1111/j.1365-294X.2005.02553.x
Excoffier L, Smouse PE, Quattro JM (1992) Analysis of molecular variance inferred from metric distances among DNA haplotypes: application to human mitochondrial DNA restriction data. Genetics 131(2):479–491. https://doi.org/10.1093/genetics/131.2.479
FAO-STAT (2023) Food and Agriculture Organization of the United Nations
Federici MT, Vaughan D, Norihiko T, Kaga A, Xin WW, Koji D et al (2001) Analysis of Uruguayan weedy rice genetic diversity using AFLP molecular markers. Electron J Biotechnol 4(3):5–6
Geleta N, Labuschagne MT, Viljoen CD (2006) Genetic diversity analysis in sorghum germplasm as estimated by AFLP, SSR and morpho-agronomical markers. Biodivers Conserv 15(10):3251–3265. https://doi.org/10.1007/s10531-005-0313-7
Giordani W, Scapim CA, Ruas PM, de Fátima Ruas C, Contreras-Soto R, Coan M et al (2019) Genetic diversity, population structure and AFLP markers associated with maize reaction to southern rust. Bragantia : Boletim Técnico Do Instituto Agronômico Do Estado De São Paulo 78:183–196
Hamza NB, Idris AE, Elmunsor II, Ibrahim AIA, Abuali AI (2016) Drought tolerance assessment in grain sorghum (Sorghum bicolor [L.] Moench) genotypes using agro-morphological traits and DNA markers. Int J Plant Breed Genet 10(3):125–131. https://doi.org/10.3923/ijpbg.2016.125.131
Hariprasanna K, Rakshit S (2016) Economic importance of sorghum. The Sorghum Genome 1–25
Harlan JR, Wet JMJ (1972) A simplified classification of cultivated sorghum 1. Crop Sci 12(2):172–176. https://doi.org/10.2135/cropsci1972.0011183X001200020005x
Heuertz M, Ater M, Lefèbvre C, Vekemans X et al (2007) Évaluation de la diversité morphologique des variétés traditionnelles de sorgho du Nord-ouest du Maroc. Biotechnologie, Agronomie, Société Et Environnement/biotechnology, Agronomy, Society and Environment 11(1):39–46
Hodkinson TR (2002) Characterization of a genetic resource collection for miscanthus (Saccharinae, Andropogoneae, Poaceae) using AFLP and ISSR PCR. Ann Bot 89(5):627–636. https://doi.org/10.1093/aob/mcf091
ICRISAT (1996) The world sorghum and millet economies: facts, trends and outlook. Food & Agriculture Org
Ismail NA, Rafii M, Mahmud T, Hanafi M, Miah G (2016) Molecular markers: a potential resource for ginger genetic diversity studies. Mol Biol Rep 43:1347–1358
Kadiri M, Ater M (1997) Diversité des variétés «locales» du sorgho grain (Sorghum bicolor Moench L.) au nord ouest du Maroc. Ressources phytogénétiques et développement durable. Actes Rabat, Maroc, 203–218
Mantel N (1967) The detection of disease clustering and a generalized regression approach. Can Res 27(2):209–220
Medraoui L, Ater M, Benlhabib O, Msikine D, Filali-Maltouf A (2007) Evaluation of genetic variability of sorghum (Sorghum bicolor L. Moench) in northwestern Morocco by ISSR and RAPD markers. C R Biol 330(11):789–797. https://doi.org/10.1016/j.crvi.2007.08.005
Menz M, Klein R, Mullet J, Obert J, Unruh N, Klein P (2002) A high-density genetic map of Sorghum bicolor (L.) Moench based on 2926 AFLP®, RFLP and SSR markers. Plant Mol Biol 48:483–499
Misganaw A, Feyissa T, Mekonnen T, Desalegne O, Disasa T (2023) Genetic diversity analysis of sorghum genotypes for sustainable genetic resource conservation and its implication for breeding program in Ethiopia. Genet Resour Crop Evol. https://doi.org/10.1007/s10722-023-01539-2
Mohammadi SA, Prasanna BM (2003) Analysis of genetic diversity in crop plants—salient statistical tools and considerations. Crop Sci 43(4):1235–1248. https://doi.org/10.2135/cropsci2003.1235
N’guni D, Geleta M, Hofvander P, Fatih M, Bryngelsson T (2012) Comparative genetic diversity and nutritional quality variation among some important Southern African sorghum accessions [‘Sorghum bicolor’(L.) Moench]. Aust J Crop Sci 6(1):56–64
Narain P (2000) Genetic diversity—conservation and assessment. Curr Sci 79(2):170–175
Nei M (1978) Estimation of average heterozygosity and genetic distance from a small number of individuals. Genetics 89(3):583–590. https://doi.org/10.1093/genetics/89.3.583
Nemera B, Kebede M, Enyew M, Feyissa T (2022) Genetic diversity and population structure of sorghum [Sorghum bicolor (L.) Moench] in Ethiopia as revealed by microsatellite markers. Acta Agric Scand Sect B Soil Plant Sci 72(1):873–884. https://doi.org/10.1080/09064710.2022.2117078
Peakall ROD, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6(1):288–295
Peakall R, Smouse P (2012) 6: genetic analysis in Excel. Population genetic software for teaching and research’’. Bioinformatics (oxford, England) 28:2537–2539
Pecina-Quintero V, Anaya-López JL, Zamarripa-Colmenero A, Montes-García N, Nuñez-Colín C, Solis-Bonilla JL et al (2012) Genetic diversity of sweet sorghum germplasm in Mexico using AFLP and SSR markers. Pesq Agrop Brasileira 47:1095–1102
Petit RJ, El Mousadik A, Pons O (1998) Identifying populations for conservation on the basis of genetic markers. Conserv Biol 12(4):844–855
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155(2):945–959. https://doi.org/10.1093/genetics/155.2.945
Ruiz-Chután JA, Salava J, Janovská D, Žiarovská J, Kalousová M, Fernández E (2019) Assessment of genetic diversity in Sorghum bicolor using RAPD markers. Genetika 51(3):789–803
Sagnard F, Deu M, Dembélé D, Leblois R, Touré L, Diakité M et al (2011) Genetic diversity, structure, gene flow and evolutionary relationships within the Sorghum bicolor wild–weedy–crop complex in a western African region. Theor Appl Genet 123:1231–1246
Saini N, Jain N, Jain S, Jain RK (2004) Assessment of genetic diversity within and among Basmati and non-Basmati rice varieties using AFLP, ISSR and SSR Markers. Euphytica 140(3):133–146. https://doi.org/10.1007/s10681-004-2510-y
Sami R, Yeye M, Usman I, Hassan L, Usman M (2013) Studies on genetic variability in some sweet sorghum (Sorghum bicolor L. Moench.) genotypes. Acad Res J Agric Sci Res 1(1):1–6
Satish L, Shilpha J, Pandian S, Rency AS, Rathinapriya P, Ceasar SA et al (2016) Analysis of genetic variation in sorghum (Sorghum bicolor (L.) Moench.) genotypes with various agronomical traits using SPAR methods. Gene 576(1):581–585. https://doi.org/10.1016/j.gene.2015.10.056
Snowdon JD (1936) The cultivated races of sorghum. The cultivated races of sorghum. https://www.cabdirect.org/cabdirect/abstract/19531700821. Accessed 24 Feb 2023
Sofalian O, Chaparzadeh N, Javanmard A, Hejazi M et al (2008) Study the genetic diversity of wheat landraces from northwest of Iran based on ISSR molecular markers. Int J Agric Biol 10(4):466–468
Tadesse H, Feyissa T (2013) Analysis of genetic diversity of Sorghum bicolor ssp. bicolor (L.) Moench using ISSR markers. Asian J Plant Sci 12(2):61–70. https://doi.org/10.3923/ajps.2013.61.70
Uptmoor R, Wenzel W, Friedt W, Donaldson G, Ayisi K, Ordon F (2003) Comparative analysis on the genetic relatedness of Sorghum bicolor accessions from Southern Africa by RAPDs, AFLPs and SSRs. Theor Appl Genet 106(7):1316–1325. https://doi.org/10.1007/s00122-003-1202-7
Vos P, Hogers R, Bleeker M, Reijans M, van de Lee T, Hornes M et al (1995) AFLP: a new technique for DNA fingerprinting. Nucleic Acids Res 23(21):4407–4414. https://doi.org/10.1093/nar/23.21.4407
Yeh FC, Yang RC, Boyle TB, Ye ZH, Mao IX (1997) The user-friendly shareware for population genetic analysis. Molecular Biology and Biotechnology Center, University of Alberta, Edmonton, Canada
Acknowledgements
Authors greatly acknowledged to ICRISAT for providing us sorghum accessions from the world collection, and to Ouafae PAKHROU, Farid RACHIDI and Mohamed ALAMI for their help in the statistical analysis. This work was supported by The Moroccan Ministry of Higher Education and Scientific Research, in the framework of the PROTARS II-P52/03.
Funding
No funding.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by ML. The first draft of the manuscript was written by ML. Revised version was edited by RK and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing interest.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Medraoui, L., Rabeh, K., Ater, M. et al. Genetic diversity analysis of sorghum (Sorghum bicolor L. Moench) landraces from northwestern Morocco using ISSR and AFLP markers. Genet Resour Crop Evol 71, 835–850 (2024). https://doi.org/10.1007/s10722-023-01665-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-023-01665-x