Abstract.
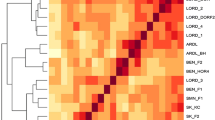
The simple sequence repeat (SSR) or microsatellite marker is currently the preferred molecular marker due to its highly desirable properties. The aim of this study was to develop and characterize more SSR markers because the number of SSR markers currently available in tomato is very limited. Five hundred DNA sequences of tomato were searched for SSRs and analyzed for the design of PCR primers. Of the 158 pairs of SSR primers screened against a set of 19 diverse tomato cultivars, 129 pairs produced the expected DNA fragments in their PCR products, and 65 of them were polymorphic with the polymorphism information content (PIC) ranging from 0.09 to 0.67. Among the polymorphic loci, 2–6 SSR alleles were detected for each locus with an average of 2.7 alleles per locus; 49.2% of these loci had two alleles and 33.8% had three alleles. The vast majority (93.8%) of the microsatellite loci contained di- or tri-nucleotide repeats and only 6.2% had tetra- and penta-nucleotide repeats. It was also found that TA/AT was the most frequent type of repeat, and the polymorphism information content (PIC) was positively correlated with the number of repeats. The set of 19 tomato cultivars were clustered based on the banding patterns generated by the 65 polymorphic SSR loci. Since the markers developed in this study are primarily from expressed sequences, they can be used not only for molecular mapping, cultivar identification and marker-assisted selection, but for identifying gene-trait relations in tomato.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
He, .C., Poysa, .V. & Yu, .K. Development and characterization of simple sequence repeat (SSR) markers and their use in determining relationships among Lycopersicon esculentum cultivars. Theor Appl Genet 106, 363–373 (2003). https://doi.org/10.1007/s00122-002-1076-0
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-002-1076-0