Abstract.
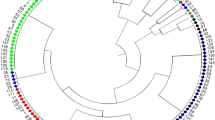
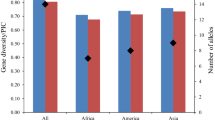
Soybean was domesticated in East Asia, where various kinds of landraces have been established as a result of adaptation to different environments and the diversification of food cultures. Asia is thus an important germplasm pool of soybean. In order to evaluate the genetic structure of the Asian soybean population, we analyzed allelic profiles at 20 simple-sequence repeat (SSR) loci of 131 accessions introduced from 14 Asian countries. The SSR loci produced an average of 11.9 alleles and a mean gene diversity of 0.782 in the accessions tested. Quantification theory III analysis and cluster analysis with the UPGMA method clearly separated the Japanese from the Chinese accessions, suggesting that the Japanese and Chinese populations formed different germplasm pools. The Korean accessions were involved in both germplasm pools, whereas most of the accessions from southeast and south/central Asia were derived from the Chinese pool. Relatively high genetic diversity and the absence of region-specific clusters in the southeast and south/central Asian populations suggest that soybean in these areas has been introduced repeatedly and independently from the diverse Chinese germplasm pool. The present study indicates that the two germplasm pools can be used as exotic genetic resources to enlarge the genetic bases of the respective Asian soybean populations.
Similar content being viewed by others
Author information
Authors and Affiliations
Additional information
Electronic Publication
Rights and permissions
About this article
Cite this article
Abe, .J., Xu, .D., Suzuki, .Y. et al. Soybean germplasm pools in Asia revealed by nuclear SSRs. Theor Appl Genet 106, 445–453 (2003). https://doi.org/10.1007/s00122-002-1073-3
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/s00122-002-1073-3