Abstract
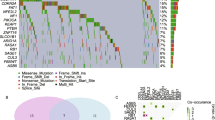
The purpose of this study was to identify key genetic pathways involved in non-small cell lung cancer (NSCLC) and understand their role in tumor progression. We performed a genome wide scanning using paired tumors and corresponding 16 mucosal biopsies from four follow-up lung cancer patients on Affymetrix 250K-NSpI array platform. We found that a single gene SH3GL2 located on human chromosome 9p22 was most frequently deleted in all the tumors and corresponding mucosal biopsies. We further validated the alteration pattern of SH3GL2 in a substantial number of primary NSCLC tumors at DNA and protein level. We also overexpressed wild-type SH3GL2 in three NSCLC cell lines to understand its role in NSCLC progression. Validation in 116 primary NSCLC tumors confirmed frequent loss of heterozygosity of SH3GL2 in overall 51 % (49/97) of the informative cases. We found significantly low (p = 0.0015) SH3GL2 protein expression in 71 % (43/60) primary tumors. Forced overexpression of wild-type (wt) SH3GL2 in three NSCLC cell lines resulted in a marked reduction of active epidermal growth factor receptor (EGFR) expression and an increase in EGFR internalization and degradation. Significantly decreased in vitro (p = 0.0015–0.030) and in vivo (p = 0.016) cellular growth, invasion (p = 0.029–0.049), and colony formation (p = 0.023–0.039) were also evident in the wt-SH3GL2-transfected cells accompanied by markedly low expression of activated AKT(Ser473), STAT3 (Tyr705), and PI3K. Downregulation of SH3GL2 interactor USP9X and activated ß-catenin was also evident in the SH3GL2-transfected cells. Our results indicate that SH3GL2 is frequently deleted in NSCLC and regulates cellular growth and invasion by modulating EGFR function.






Similar content being viewed by others
References
Sato M, Shames DS, Gazdar AF, Minna JD (2007) A translational view of the molecular pathogenesis of lung cancer. J Thoracic Oncol 2:327–343
http://www.cancer.gov/cancertopics/types/lung. Accessed 7 March 2012
Andrews J, Yeh P, Pao W, Horn L (2011) Molecular predictors of response to chemotherapy in non-small cell lung cancer. Cancer J 17:104–113
Sun S, Schiller JH, Gazdar AF (2007) Lung cancer in never smokers—a different disease. Nature Rev Cancer 7:778–790
Jonsson S, Varella-Garcia M, Miller YE, Wolf HJ, Byers T, Braudrick S, Kiatsimkul P, Lewis M, Kennedy TC, Keith RL et al (2008) Chromosomal aneusomy in bronchial high-grade lesions is associated with invasive lung cancer. Am J Respir Crit Care Med 177:342–347
Tonon G, Wong KK, Maulik G, Brennan C, Feng B, Zhang Y, Khatry DB, Protopopov A, You MJ, Aguirre AJ et al (2005) High-resolution genomic profiles of human lung cancer. Proc Natl Acad Sci U S A 102:9625–9630
Varella-Garcia M, Chen L, Powell RL, Hirsch FR, Kennedy TC, Keith R, Miller YE, Mitchell JD, Franklin WA (2007) Spectral karyotyping detects chromosome damage in bronchial cells of smokers and patients with cancer. Am J Respir Crit Care Med 6:505–512
Massion PP, Zou Y, Uner H, Kiatsimkul P, Wolf HJ, Baron AE, Byers T, Jonsson S, Lam S, Hirsch FR et al (2009) Recurrent genomic gains in preinvasive lesions as a biomarker of risk for lung cancer. PLoS One 4:e5611
Weir BA, Woo MS, Getz G, Perner S, Ding L, Beroukhim R, Lin WM, Province MA, Kraja A, Johnson LA et al (2007) Characterizing the cancer genome in lung adenocarcinoma. Nature 450:893–898
Lockwood WW, Chari R, Coe BP, Thu KL, Garnis C, Malloff CA, Campbell J, Williams AC, Hwang D, Zhu CQ et al (2010) Integrative genomic analyses identify BRF2 as a novel lineage-specific oncogene in lung squamous cell carcinoma. PLoS Med 7:e1000315
Thomas RK, Weir B, Meyerson M (2006) Genomic approaches to lung cancer. Clin Cancer Res 12:4384s–4391s
Dasgupta S, Yung RC, Westra WH, Rini DA, Brandes J, Sidransky D (2009) Following mitochondrial footprints through a long mucosal path to lung cancer. PLoS One 4:e6533
Dasgupta S, Koch R, Westra WH, Califano JA, Ha PK, Sidransky D, Koch WM (2010) Mitochondrial DNA mutation in margins and tumors of recurrent HNSCC patients. Can Prev Res 3:1205–1211
Dasgupta S, Soudry E, Mukhopadhyay N, Shao C, Yee J, Lam S et al (2011) Mitochondrial DNA mutations in respiratory complex-I in never-smoker lung cancer patients contribute to lung cancer progression and associated with EGFR gene mutation. J Cell Physiol 227(6):2451–2460
Dasgupta S, Bhattacharya-Chatterjee M, O’Malley BW Jr, Chatterjee SK (2004) Reversal of immune suppression following vaccination with recombinant vaccinia virus expressing IL-2 in an orthotopic murine model of head and neck squamous cell carcinoma. Cancer Therapy 2:375–388
Dasgupta S, Hoque MO, Upadhyay S, Sidransky D (2008) Mitochondrial cytochrome B gene mutation promotes tumor growth in bladder cancer. Cancer Res 68:700–706
Szymkiewicz I, Kowanetz K, Soubeyran P, Dinarina A, Lipkowitz S, Dikic I (2002) CIN85 participates in Cb-1-b down-regulation of receptor tyrosine kinases. J Biochem 277:39666–39672
Dasgupta S, Hoque MO, Upadhyay S, Sidransky D (2009) Forced cytochrome B gene mutation expression induces mitochondrial proliferation and prevents apoptosis in human uroepithelial SV-HUC-1 cells. Int J Cancer 125:2829–2835
Soubyran S, Kowanetz K, Szymkiewicz I, Langdon WY, Dikic I (2002) CbI-CIN85-endophlin complex mediates ligand-induced downregulation of EGF receptors. Nature 416:183–187
Ettenberg SA, Keane MM, Nau MM, Frankel M, Wang LM, Pierce JH, Lipkowitz S (2009) cbl-b inhibits epidermal growth factor receptor signaling. Oncogene 18:1855–1866
Levkowitz G, Waterman H, Ettenberg SA, Katz M, Tsygankov AY, Alroy I, Lavi S, Iwai K, Reiss Y, Ciechanover A et al (1999) Ubiquitin ligase activity and tyrosine phosphorylation underlie suppression of growth factor signaling by c-Cbl/Sli-1. Mol Cell 4:1029–1040
Kjaerulff O, Brodin L, Jung A (2011) The structure and function of endophilin proteins. Cell Biochem Biophys 60:137–154
Hoshida Y, Villanueva A, Kobayashi M, Peix J, Chiang DY, Camargo A, Gupta S, Moore J, Wrobel MJ, Lerner J et al (2008) Gene expression in fixed tissues and outcome in hepatocellular carcinoma. N Engl J Med 359:1995–2004
Sinha S, Chunder N, Mukherjee N, Alam N, Roy A, Roychoudhury S, Panda CK (2008) Frequent deletion and methylation in SH3GL2 and CDKN2A loci are associated with early- and late-onset breast carcinoma. Ann Surg Oncol 5:1070–1080
Ghosh A, Ghosh S, Maiti GP, Sabbir MG, Alam N, Sikdar N, Roy B, Roychoudhury S, Panda CK (2009) SH3GL2 and CDKN2A/2B loci are independently altered in early dysplastic lesions of head and neck: correlation with HPV infection and tobacco habit. J Pathol 217:408–419
Osterberg L, Levan K, Partheen K, Delle U, Olsson B, Sundfeldt K, Horvath G (2009) Potential predictive markers of chemotherapy resistance in stage III ovarian serous carcinomas. BMC Cancer 9:368
Hu T, Li C (2010) Convergence between Wnt-β-catenin and EGFR signaling in cancer. Mol Cancer 9:236
Vucic D, Dixit VM, Wertz IE (2011) Ubiquitination in apoptosis: a post translational modification at the edge of life and death. Nat Rev Mol Cell Biol 12:439–452
Acknowledgments
This work was supported by EDRN-UO1 CA 084986 (DS); US–Egypt Joint Science and Technology fund (58-3148-169, SD), AD Williams (646299, SD) and Elsa U. Pardee Foundation (548424, SD), NIH-CA80127, and NIH-CA80354 (PY). PBF holds the Thelma Newmeyer Corman Chair in Cancer Research in the VCU Massey Cancer Center.
Conflict of interest
No potential conflict of interests exists.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM 1
(PDF 338 KB)
Rights and permissions
About this article
Cite this article
Dasgupta, S., Jang, J.S., Shao, C. et al. SH3GL2 is frequently deleted in non-small cell lung cancer and downregulates tumor growth by modulating EGFR signaling. J Mol Med 91, 381–393 (2013). https://doi.org/10.1007/s00109-012-0955-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-012-0955-3