Abstract
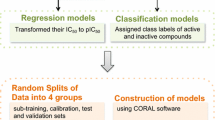
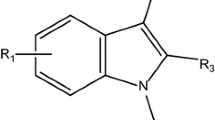
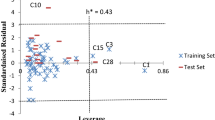
A series of indole-based C-3 pyridone compounds active against Hepatitis C virus as HCV NS5B polymerase inhibitors have been explored under the framework of quantitative structure–activity relationships (QSARs) and pharmacophore modeling in the present article. QSAR models were developed by considering various kinds of theoretical molecular descriptors including constitutional and geometrical, topological, functional group, and atom-centered fragments indices computed solely from the structure of indole-based C-3 pyridone compounds utilizing stepwise forward−backward variable selections incorporated in multiple linear regression (MLR) methods. MLR shows that topological indices can contribute the maximum impact on biological activity obtained in terms of model quality parameters, such as R 2 = 0.946, \( Q_{\text{Loo}}^{2} \) = 0.883, and \( R_{\text{pred}}^{2} \) = 0.642, respectively. Constitutional and geometrical-based model can produce R 2 = 0.849, \( Q_{\text{Loo}}^{2} \) = 0.776, and \( R_{\text{pred}}^{2} \) = 0.623, respectively; whereas, model utilizing functional group and atom-centered fragments indices can contribute R 2 = 0.885, \( Q_{\text{Loo}}^{2} \) = 0.812, and \( R_{\text{pred}}^{2} \) = 0.612, respectively. Prediction of essential structural features is strongly achieved by introducing pharmacophore modeling of these highly active congeners which showed that more number of hydrogen bonding interactions between the substituents associated to the various points of diversity of the indole nucleus including N-1, C-2, and C-3 aromatic groups with the HCV NS5B polymerase target may enhance the biological activities in this congeneric ligands.




Similar content being viewed by others
References
Basak SC (1987) Use of molecular complexity indices in predictive pharmacology and toxicology: a QSAR approach. Med Sci Res 15:605–609
Basak SC (2013) Mathematical descriptors in the prediction of property, bioactivity, and toxicity of chemicals from their structure: a chemical-cum-biochemical approach. Curr Comput Aided Drug Des 9(4):449–462
Basak SC, Grunwald GD, Niemi GJ (1997) In: Balaban AT (ed) From chemical topology to three-dimensional geometry. Plenum Press, New York, pp 73–116
Beaulieu PL (2007) Non-nucleoside inhibitors of the hcv ns5b polymerase: progress in the discovery and development of novel agents for the treatment of hcv infections. Curr Opin Invest Drugs 8:614–634
Chen KX, Vibulbhan B, Yang W, Sannigrahi M, Velazquez F, Chan T-Y, Venkatraman AGN, Zeng Q, Bennet F, Jiang Y, Lesburg CA, Duca J, Pinto P, Gavalas S, Huang Y, Wu W, Selyutin O, Agrawal S, Feld B, Huang H-C, Li C, Cheng K-C, Shih N-Y, Kozlowski JA, Rosenblum SB, Njoroge FG (2012) Structure–activity relationship (SAR) Development and discovery of potent indole-based inhibitors of the Hepatitis C Virus (HCV) NS5B Polymerase. J Med Chem 55:754–765
Choo QL, Kuo G, Weiner AJ, Overby LR, Bradley DW, Houghton M (1989) Isolation of a cDNA clone derived from a blood-borne non-A, non-B viral hepatitis genome. Science 244:359–362
Estrada E (1999) In: Devillers J, Balaban AT (eds) Topological indices and related descriptors in QSAR and QSPR. Gordon and Breach, Amsterdam, pp 403–453
Fukui K (1982) Role of frontier orbitals in chemical reactions. Science 218:747–754
Katritzky AR, Petrukhin R, Tatham D, Basak S, Benfenati E, Karelson M, Maran U (2001) Interpretation of quantitative structure–property–activity relationships. J Chem Inf Comput Sci 41:679–685
Koes DR, Camacho CJ (2011) Pharmer: efficient and exact pharmacophore search. J Chem Inf Model 51:1307–1314
Langer T, Krovat EM (2003) Chemical feature-based pharmacophores and virtual library screening for discovery of new leads. Curr Opin Drug Discov Dev 6:370–376
Leach AR, Gillet VJ, Lewis RA, Taylor R (2009) Three dimensional pharmacophore methods in drug discovery. J Med Chem 53:539–558
Liu Y, He Y, Molla A (2009) Hepatitis C virus polymerase as a target for antiviral drug intervention: non-nucleoside inhibitors. In: LaFemina RL (ed) Antiviral research: strategies in antiviral drug discovery. ASM Press, Washington, DC, pp 137–151
Mason JS, Good AC, Martin EJ (2001) 3-D pharmacophores in drug discovery. Curr Pharm Des 7:567–597
Miller AJ (1990) Subset selections in regression. Chapman and Hall, New York
Mills N (2006) ChemDraw Ultra 10.0. J Am Chem Soc 128:13649–13650
Minitab® Statistical Software: Minitab 2010. www.minitab.com
Nandi S, Bagchi MC (2011) In silico design of potent EGFR kinase inhibitors using combinatorial libraries. Mol Simul 37(3):196–209
Pompe M, Novič M (1999) Prediction of gas-chromatographic retention indices using topological descriptors. J Chem Inf Comput Sci 39:59–67
Randic M (1975) On characterization of molecular branching. J Am Chem Soc 79:6609–6615
Randic M (1984) On molecular identification numbers. J Chem Inf Comput Sci 24:164–175
Randic M (2001) Novel shape descriptors for molecular graphs. J Chem Inf Comput Sci 41:607–613
Rao CR (1973) Linear statistical inference and its applications, 2nd edn. Wiley, New York
Roy PP, Roy K (2009) Comparative chemometric modeling of cytochrome 3A4 inhibitory activity of structurally diverse compounds using stepwise MLR, FA-MLR, PLS, GFA PLS and ANN techniques. Eur J Med Chem 44:2913–2922
Schuster D, Langer T (2005) The identification of ligand features essential for PXR activation by pharmacophore modeling. J Chem Info Model 45:31
Shoichet BK (2004) Virtual screening of chemical libraries. Nature 432:862
Sustmann R (1974) Orbital energy control of cycloaddition reactivity. Pure Appl Chem 40:569–593
SYSTAT, Version 7.0, SPSS Inc., 444 North Michigan Avenue, Chicago, IL 60611
Todeschini R, Consonni V (2009) Molecular descriptors for chemoinformatics, revised and enlarged edition, 2nd edn. Wiley, Weinheim
Todeschini R, Consonni V. Dragon software (version 5.4-2006). Milano, Italy
Ward JH (1963) Hierarchical grouping to optimize an objective function. J Am Stat Assoc 58:236–244
Wolber G, Langer T (2005) LigandScout: 3-D pharmacophores derived from protein-bound ligands and their use as virtual screening filters. J Chem Inf Model 45:160–169
Acknowledgments
Authors are sincerely thankful to their corresponding Institute for providing necessary research facilities. Ankita shows deep sense of gratitude to her supervisor Dr. Nandi for his excellent guidance. MCB acknowledges the Council of Scientific and Industrial Research (CSIR), New Delhi, India for the Grant of a CSIR Emeritus Scientist award to him. SN is thankful to National Institute of Chemistry Slovenia for availing DRAGON Professional version 5.4-2006 software for the calculation of theoretical molecular descriptors and LIGANDSCOUT 2.02 software for pharmacophore modeling used in the present work.
Conflict of interest
None.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Batra, A., Nandi, S. & Bagchi, M.C. QSAR and pharmacophore modeling of indole-based C-3 pyridone compounds as HCV NS5B polymerase inhibitors utilizing computed molecular descriptors. Med Chem Res 24, 2432–2440 (2015). https://doi.org/10.1007/s00044-014-1304-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-014-1304-6