Abstract
The cytoophidium is a unique type of membraneless compartment comprising of filamentous protein polymers. Inosine monophosphate dehydrogenase (IMPDH) catalyzes the rate-limiting step of de novo GTP biosynthesis and plays critical roles in active cell metabolism. However, the molecular regulation of cytoophidium formation is poorly understood. Here we show that human IMPDH2 polymers bundle up to form cytoophidium-like aggregates in vitro when macromolecular crowders are present. The self-association of IMPDH polymers is suggested to rely on electrostatic interactions. In cells, the increase of molecular crowding with hyperosmotic medium induces cytoophidia, while the decrease of that by the inhibition of RNA synthesis perturbs cytoophidium assembly. In addition to IMPDH, CTPS and PRPS cytoophidium could be also induced by hyperosmolality, suggesting a universal phenomenon of cytoophidium-forming proteins. Finally, our results indicate that the cytoophidium can prolong the half-life of IMPDH, which is proposed to be one of conserved functions of this subcellular compartment.








Similar content being viewed by others
Availability of data and material
All data are available in the text and supplementary information.
References
Liu JL (2016) The cytoophidium and its kind: filamentation and compartmentation of metabolic enzymes. Annu Rev Cell Dev Biol 32:349–372
Liu JL (2010) Intracellular compartmentation of CTP synthase in Drosophila. J Genet Genom 37(5):281–296
Zhang B, Tastan OY, Zhou X, Guo CJ, Liu X, Thind A, Hu HH, Zhao S, Liu JL (2020) The proline synthesis enzyme P5CS forms cytoophidia in Drosophila. J Genet Genom 47(3):131–143
Zhang S, Ding K, Shen QJ, Zhao S, Liu JL (2018) Filamentation of asparagine synthetase in Saccharomyces cerevisiae. PLoS Genet 14(10):e1007737
Chang CC, Lin WC, Pai LM, Lee HS, Wu SC, Ding ST, Liu JL, Sung LY (2015) Cytoophidium assembly reflects upregulation of IMPDH activity. J Cell Sci 128(19):3550–3555
Carcamo WC, Satoh M, Kasahara H, Terada N, Hamazaki T, Chan JY, Yao B, Tamayo S, Covini G, von Muhlen CA et al (2011) Induction of cytoplasmic rods and rings structures by inhibition of the CTP and GTP synthetic pathway in mammalian cells. PLoS ONE 6(12):e29690
Lynch EM, Kollman JM (2020) Coupled structural transitions enable highly cooperative regulation of human CTPS2 filaments. Nat Struct Mol Biol 27(1):42–48
Johnson MC, Kollman JM (2020) Cryo-EM structures demonstrate human IMPDH2 filament assembly tunes allosteric regulation. Elife. https://doi.org/10.7554/eLife.53243
Lynch EM, Hicks DR, Shepherd M, Endrizzi JA, Maker A, Hansen JM, Barry RM, Gitai Z, Baldwin EP, Kollman JM (2017) Human CTP synthase filament structure reveals the active enzyme conformation. Nat Struct Mol Biol 24(6):507–514
Anthony SA, Burrell AL, Johnson MC, Duong-Ly KC, Kuo YM, Simonet JC, Michener P, Andrews A, Kollman JM, Peterson JR (2017) Reconstituted IMPDH polymers accommodate both catalytically active and inactive conformations. Mol Biol Cell 280:2600–2608
Zhou X, Guo CJ, Hu HH, Zhong J, Sun Q, Liu D, Zhou S, Chang CC, Liu JL (2019) Drosophila CTP synthase can form distinct substrate- and product-bound filaments. J Genet Genom 46(11):537–545
Sun Z, Liu JL (2019) Forming cytoophidia prolongs the half-life of CTP synthase. Cell Discov 5:32
Lin WC, Chakraborty A, Huang SC, Wang PY, Hsieh YJ, Chien KY, Lee YH, Chang CC, Tang HY, Lin YT et al (2018) Histidine-dependent protein methylation is required for compartmentalization of CTP synthase. Cell Rep 24(10):2733-2745 e2737
Sun Z, Liu JL (2019) mTOR-S6K1 pathway mediates cytoophidium assembly. J Genet Genom 46(2):65–74
Keppeke GD, Barcelos D, Fernandes M, Comodo AN, Guimaraes DP, Cardili L, Carapeto FCL, Andrade LEC, Landman G (2019) IMP dehydrogenase rod/ring structures in acral melanomas. Pigment Cell Melanoma Res. https://doi.org/10.1111/pcmr.12854
Keppeke GD, Chang CC, Peng M, Chen LY, Lin WC, Pai LM, Andrade LEC, Sung LY, Liu JL (2018) IMP/GTP balance modulates cytoophidium assembly and IMPDH activity. Cell Div 13:5
Duong-Ly KC, Kuo YM, Johnson MC, Cote JM, Kollman JM, Soboloff J, Rall GF, Andrews AJ, Peterson JR (2018) T cell activation triggers reversible inosine-5’-monophosphate dehydrogenase assembly. J Cell Sci 131(17):jcs223289
Calise SJ, Abboud G, Kasahara H, Morel L, Chan EKL (2018) Immune response-dependent assembly of IMP dehydrogenase filaments. Front Immunol 9:2789
Peng M, Chang CC, Liu JL, Sung LY (2021) CTPS and IMPDH form cytoophidia in developmental thymocytes. Exp Cell Res 405(1):112662
Burrell AL, Kollman JM (2022) IMPDH dysregulation in disease: a mini review. Biochem Soc Trans 50(1):71–82
Zech M, Jech R, Boesch S, Skorvanek M, Weber S, Wagner M, Zhao C, Jochim A, Necpal J, Dincer Y et al (2020) Monogenic variants in dystonia: an exome-wide sequencing study. Lancet Neurol 19(11):908–918
Bowne SJ, Sullivan LS, Mortimer SE, Hedstrom L, Zhu J, Spellicy CJ, Gire AI, Hughbanks-Wheaton D, Birch DG, Lewis RA et al (2006) Spectrum and frequency of mutations in IMPDH1 associated with autosomal dominant retinitis pigmentosa and leber congenital amaurosis. Invest Ophthalmol Vis Sci 47(1):34–42
Kofuji S, Hirayama A, Eberhardt AO, Kawaguchi R, Sugiura Y, Sampetrean O, Ikeda Y, Warren M, Sakamoto N, Kitahara S et al (2019) IMP dehydrogenase-2 drives aberrant nucleolar activity and promotes tumorigenesis in glioblastoma. Nat Cell Biol 21(8):1003–1014
Hedstrom L (2009) IMP dehydrogenase: structure, mechanism, and inhibition. Chem Rev 109(7):2903–2928
Chakraborty A, Lin WC, Lin YT, Huang KJ, Wang PY, Chang IY, Wang HI, Ma KT, Wang CY, Huang XR et al (2020) SNAP29 mediates the assembly of histidine-induced CTP synthase filaments in proximity to the cytokeratin network. J Cell Sci 133(9):jcs240200
Thomas EC, Gunter JH, Webster JA, Schieber NL, Oorschot V, Parton RG, Whitehead JP (2012) Different characteristics and nucleotide binding properties of inosine monophosphate dehydrogenase (IMPDH) isoforms. PLoS ONE 7(12):e51096
Ingerson-Mahar M, Briegel A, Werner JN, Jensen GJ, Gitai Z (2010) The metabolic enzyme CTP synthase forms cytoskeletal filaments. Nat Cell Biol 12(8):739–746
Chang CC, Keppeke GD, Sung LY, Liu JL (2018) Interfilament interaction between IMPDH and CTPS cytoophidia. FEBS J 285(20):3753–3768
Labesse G, Alexandre T, Vaupre L, Salard-Arnaud I, Him JL, Raynal B, Bron P, Munier-Lehmann H (2013) MgATP regulates allostery and fiber formation in IMPDHs. Structure 21(6):975–985
Fernandez-Justel D, Nunez R, Martin-Benito J, Jimeno D, Gonzalez-Lopez A, Soriano EM, Revuelta JL, Buey RM (2019) A nucleotide-dependent conformational switch controls the polymerization of human IMP dehydrogenases to modulate their catalytic activity. J Mol Biol 431(5):956–969
Petrovska I, Nuske E, Munder MC, Kulasegaran G, Malinovska L, Kroschwald S, Richter D, Fahmy K, Gibson K, Verbavatz JM et al (2014) Filament formation by metabolic enzymes is a specific adaptation to an advanced state of cellular starvation. Elife. https://doi.org/10.7554/eLife.02409
Dumetz AC, Chockla AM, Kaler EW, Lenhoff AM (2008) Effects of pH on protein-protein interactions and implications for protein phase behavior. Biochem Biophys Acta 1784(4):600–610
Keppeke GD, Chang CC, Antos CL, Peng M, Sung LY, Andrade LEC, Liu JL (2021) IMPDH forms the cytoophidium in zebrafish. Dev Biol 478:89–101
Keppeke GD, Prado MS, Nunes E, Perazzio SF, Rodrigues SH, Ferraz ML, Chan EK, Andrade LE (2016) Differential capacity of therapeutic drugs to induce Rods/Rings structures in vitro and in vivo and generation of anti-Rods/Rings autoantibodies. Clin Immunol 173:149–156
Ji Y, Gu J, Makhov AM, Griffith JD, Mitchell BS (2006) Regulation of the interaction of inosine monophosphate dehydrogenase with mycophenolic Acid by GTP. J Biol Chem 281(1):206–212
Keppeke GD, Calise SJ, Chan EK, Andrade LE (2015) Assembly of IMPDH2-based, CTPS-based, and mixed rod/ring structures is dependent on cell type and conditions of induction. J Genet Genom 42(6):287–299
Ralston GB (1990) Effects of crowding in protein solutions. J Chem Educ 67(10):857–860
Jalihal AP, Pitchiaya S, Xiao L, Bawa P, Jiang X, Bedi K, Parolia A, Cieslik M, Ljungman M, Chinnaiyan AM et al (2020) Multivalent proteins rapidly and reversibly phase-separate upon osmotic cell volume change. Mol Cell 79(6):978–990
Noree C, Begovich K, Samilo D, Broyer R, Monfort E, Wilhelm JE (2019) A quantitative screen for metabolic enzyme structures reveals patterns of assembly across the yeast metabolic network. Mol Biol Cell 30(21):2721–2736
Gou KM, Chang CC, Shen QJ, Sung LY, Liu JL (2014) CTP synthase forms cytoophidia in the cytoplasm and nucleus. Exp Cell Res 323(1):242–253
Begovich K, Yelon D, Wilhelm JE (2020) PRPS polymerization influences lens fiber organization in zebrafish. Dev Dyn 249(8):1018–1031
Alfieri RR, Petronini PG (2007) Hyperosmotic stress response: comparison with other cellular stresses. Pflug Arch 454(2):173–185
Delarue M, Brittingham GP, Pfeffer S, Surovtsev IV, Pinglay S, Kennedy KJ, Schaffer M, Gutierrez JI, Sang D, Poterewicz G et al (2018) mTORC1 controls phase separation and the biophysical properties of the cytoplasm by tuning crowding. Cell 174(2):338-349 e320
Moriconi WJ, Slavik M, Taylor S (1986) 3-Deazauridine (NSC 126849): an interesting modulator of biochemical response. Invest New Drugs 4(1):67–84
Chiti F, Dobson CM (2017) Protein misfolding, amyloid formation, and human disease: a summary of progress over the last decade. Annu Rev Biochem 86:27–68
Kopito RR (2000) Aggresomes, inclusion bodies and protein aggregation. Trends Cell Biol 10(12):524–530
Lee JE, Cathey PI, Wu H, Parker R, Voeltz GK (2020) Endoplasmic reticulum contact sites regulate the dynamics of membraneless organelles. Science. https://doi.org/10.1126/science.aay7108
Hansen JM, Horowitz A, Lynch EM, Farrell DP, Quispe J, DiMaio F, Kollman JM (2021) Cryo-EM structures of CTP synthase filaments reveal mechanism of pH-sensitive assembly during budding yeast starvation. Elife 10:e73368
Zhou HX, Rivas G, Minton AP (2008) Macromolecular crowding and confinement: biochemical, biophysical, and potential physiological consequences. Annu Rev Biophys 37:375–397
Zimmerman SB, Trach SO (1991) Estimation of macromolecule concentrations and excluded volume effects for the cytoplasm of Escherichia coli. J Mol Biol 222(3):599–620
Andreadis C, Hulme L, Wensley K, Liu JL (2019) The TOR pathway modulates cytoophidium formation in Schizosaccharomyces pombe. J Biol Chem 294(40):14686–14703
Garcia-Mata R, Gao YS, Sztul E (2002) Hassles with taking out the garbage: aggravating aggresomes. Traffic 3(6):388–396
Ries M, Sastre M (2016) Mechanisms of abeta clearance and degradation by glial cells. Front Aging Neurosci 8:160
Juda P, Smigova J, Kovacik L, Bartova E, Raska I (2014) Ultrastructure of cytoplasmic and nuclear inosine-5’-monophosphate dehydrogenase 2 “rods and rings” inclusions. J Histochem Cytochem 62(10):739–750
Plana-Bonamaiso A, Lopez-Begines S, Fernandez-Justel D, Junza A, Soler-Tapia A, Andilla J, Loza-Alvarez P, Rosa JL, Miralles E, Casals I et al (2020) Post-translational regulation of retinal IMPDH1 in vivo to adjust GTP synthesis to illumination conditions. Elife 9:e56418
Nuske E, Marini G, Richter D, Leng W, Bogdanova A, Franzmann TM, Pigino G, Alberti S (2020) Filament formation by the translation factor eIF2B regulates protein synthesis in starved cells. Biol Open 9(7):bio046391
Kozhevnikova EN, van der Knaap JA, Pindyurin AV, Ozgur Z, van Ijcken WF, Moshkin YM, Verrijzer CP (2012) Metabolic enzyme IMPDH is also a transcription factor regulated by cellular state. Mol Cell 47(1):133–139
McLean JE, Hamaguchi N, Belenky P, Mortimer SE, Stanton M, Hedstrom L (2004) Inosine 5’-monophosphate dehydrogenase binds nucleic acids in vitro and in vivo. Biochem J 379(Pt 2):243–251
Mortimer SE, Xu D, McGrew D, Hamaguchi N, Lim HC, Bowne SJ, Daiger SP, Hedstrom L (2008) IMP dehydrogenase type 1 associates with polyribosomes translating rhodopsin mRNA. J Biol Chem 283(52):36354–36360
Wang S, Chao F, Zhang C, Han D, Xu G, Chen G (2021) Circular RNA circPFKP promotes cell proliferation by activating IMPDH2 in prostate cancer. Cancer Lett 524:109–120
Bianchi-Smiraglia A, Wolff DW, Marston DJ, Deng Z, Han Z, Moparthy S, Wombacher RM, Mussell AL, Shen S, Chen J et al (2021) Regulation of local GTP availability controls RAC1 activity and cell invasion. Nat Commun 12(1):6091
Ruan H, Song Z, Cao Q, Ni D, Xu T, Wang K, Bao L, Tong J, Xiao H, Xiao W et al (2020) IMPDH1/YB-1 positive feedback loop assembles cytoophidia and represents a therapeutic target in metastatic tumors. Mol Ther 28(5):1299–1313
Acknowledgements
We thank the Molecular and Cell Biology Core Facility (MCBCF) at the School of Life Science and Technology, ShanghaiTech University for providing technical support.
Funding
This work was supported by grants from Ministry of Science and Technology of China (No. 2021YFA0804700), National Natural Science Foundation of China (No. 31771490), Shanghai Science and Technology Commission (No. 20JC1410500) and the UK Medical Research Council (grant nos. MC_UU_12021/3 and MC_U137788471) for grants to J.L.L.
Author information
Authors and Affiliations
Contributions
CCC conceived and designed the studies, analyzed the data. CCC, MP, JZ, ZZ and GDK performed the experiments. JLL and LYS supervised the project. CCC and JLL wrote and edited the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors have no relevant financial or non-financial interests to disclose.
Consent for publication
Not applicable.
Ethics approval and consent to participate
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.

18_2022_4448_MOESM1_ESM.jpg
Supplementary file1 Figure S1. Hyperosmotic medium does not induce the aggregation of PAICS, GMPS and HPRT. Immunofluorescence of wild-type HEK 293T cells treated with 300 mM sucrose for 3 hours. No detectable aggregates of PAICS, GMPS or HPRT were observed in the cells. Scale bars = 20 μm. (JPG 994 kb)
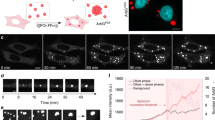
Supplementary file2 Movie S1. Live-cell imaging showing the dynamics of IMPDH amorphous clumps. OFP-IMPDH2 expressing HeLa cells were treated with MPA. Time intervals of each frame is 40 seconds. (AVI 59 kb)
Supplementary file3 Movie S2. Live-cell imaging showing an IMPDH amorphous clump transforms in association with the ER. GFP-Sec61β (green) and OFP-IMPDH2 (red) expressing HeLa cells were treated with MPA. Time intervals of each frame is 5 seconds. (AVI 688 kb)
Supplementary file4 Movie S3. Live-cell imaging showing the transformation of IMPDH amorphous clumps to filaments. OFP-IMPDH2 expressing HeLa cells were treated with MPA. Time intervals of each frame is 60 seconds. (AVI 100 kb)
Rights and permissions
About this article
Cite this article
Chang, CC., Peng, M., Zhong, J. et al. Molecular crowding facilitates bundling of IMPDH polymers and cytoophidium formation. Cell. Mol. Life Sci. 79, 420 (2022). https://doi.org/10.1007/s00018-022-04448-2
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00018-022-04448-2