Abstract
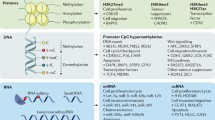
Gastric carcinogenesis occurs when gastric epithelial cells transition through the initial, immortal, premalignant, and malignant stages of transformation. Epigenetic regulations contribute to this multistep process. Due to the critical role of epigenetic modifications , these changes are highly likely to be of clinical use in the future as new biomarkers and therapeutic targets for the early detection and treatment of cancers. Here, we summarize the recent findings on how epigenetic modifications, including DNA methylation, histone modifications, and non-coding RNAs, regulate gastric carcinogenesis, and we discuss potential new strategies for the diagnosis and treatments of gastric cancer. The strategies may be helpful in the further understanding of epigenetic regulation in human diseases.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The latest figures from the World Health Organization (WHO) show that 951,600 new gastric cancer cases and 723,100 gastric cancer-related deaths occurred globally in 2012 [1]. The overall incidence of gastric cancer has declined. However, in China, the morbidity and mortality of gastric cancer rank 2nd and 3rd, respectively, among all malignant tumors [2]. Gastric cancer is classified into the following two subtypes: diffuse and intestinal. Usually, intestinal gastric cancers retain a glandular structure and undergo multiple processes as follows: chronic inflammation, atrophy, intestinal metaplasia, and atypical hyperplasia and eventually into gastric cancer. However, diffuse gastric cancer is relatively rare and is poorly differentiated to the extent that no glandular structure is recognizable. At present, no clear precancerous lesions of diffuse gastric cancer have been defined [3, 4]. Helicobacter pylori (H. pylori) infection plays an important carcinogenic role in both subtypes of gastric cancer. Many trials have demonstrated the possibility of cancer prevention through H. pylori screening and eradication. The malignant transformation of gastric mucosa involves multimolecular events, including gene mutation [5] and epigenetic alteration [6]. This study presents a review of the roles of epigenetic alterations in the malignant transformation of gastric mucosa.
The concept and significance of epigenetics
The concept of epigenetics was first proposed by Waddington [7]. Epigenetics refers to the heritable changes in gene expression that are independent of variations in DNA sequences. The main types of epigenetic processes include DNA methylation, histone modification, and chromatin remodeling as well as the function of non-coding RNA (ncRNA). The basic theory of classical genetics cannot adequately explain the biodiversity within species. For example, identical twins carrying the same DNA sequences may exhibit distinct phenotypes and different susceptibility to diseases. The proposal of epigenetics has compensated for such shortcoming of classical genetic theory. Epigenetics is a component of normal physiological regulation, and abnormal epigenetic regulation may lead to tumorigenesis. Studies have suggested that intestinal-type gastric cancer originates from chronic gastritis, which gradually progresses through stages of chronic atrophic gastritis, intestinal metaplasia, and atypical hyperplasia and ultimately develops into advanced gastric cancer [8, 9]. During the malignant transformation of gastric mucosa, a large number of genes are subjected to epigenetic regulation. The genes show cumulative changes as the disease evolves [10, 11].
Methylation of tumor suppressor genes is an important mechanism responsible for malignant transformation of gastric mucosa
Methylation is a type of chemical modification that occurs in DNA sequences. In mammalian cells, DNA methylation occurs almost exclusively at the fifth carbon atom of the cytosine residues within cytosine–phosphate–guanine (CpG) dinucleotides. CpG dinucleotides tend to form CG-rich clusters called CpG islands. CpG islands are mainly distributed in the core promoter sequence and transcription start site of structural genes. DNA methylation may induce changes in chromatin structure, DNA conformation, DNA stability, and the interactions between DNA and protein, resulting in transcription inhibition [12]. Two adverse phenomena characterize the process of carcinogenesis: locus-specific hypermethylation and global depletion of methyl groups from cancer genomes. Hypermethylation of promoters has been widely shown to contribute to the silencing of tumor suppressor genes during carcinogenesis. Global hypomethylation of the cancer genome was initially shown to cause genome-wide allelic instability, but recently, the involvement of this process in transcriptional gene regulation has become increasingly recognized [13, 14].
Promoter hypermethylation-induced inactivation of tumor suppressor genes is an important mechanism that leads to gastric carcinogenesis [15]. For example, CDH1, the gene encoding epithelial cadherin (E-cadherin), is a tumor suppressor gene located on chromosome 16q22.1. E-cadherin is expressed in normal epithelium and plays a role in calcium-dependent cell adhesion. CDH1 is hypermethylated in 40–80 % of human primary gastric carcinoma. In diffuse gastric cancer, a methylation-induced decrease in E-cadherin expression has been observed in more than 50 % of the undifferentiated early cancers and adjacent non-cancerous gastric epithelial tissues. Therefore, CDH1 methylation-induced loss of E-cadherin expression is an early event in the malignant transformation of gastric mucosa [16, 17]. E-cadherin is also inactivated by mutation and accounts for the hereditary nature of diffuse-type gastric cancer [18]. Runt-related transcription factor 3 (RUNX3) is a key molecule in the transforming growth factor-β (TGF-β) signaling pathway. The expression of RUNX3 is significantly reduced in gastric cancer. The main reason for the decreased RUNX3 expression is DNA hypermethylation in the promoter region. Kim et al. found that RUNX3 CpG island methylation occured in 8.1 % of chronic gastritis cases, 28.1 % of intestinal metaplasia cases, 27.3 % of gastric adenocarcinoma cases, 64 % of primary gastric cancer cases, and 60 % of gastric cancer cell lines [19]. In RUNX3 knockout mice, apoptosis is inhibited. These mice show hypertrophy of gastric mucosa and intestinal metaplasia of gastric epithelial cells, indicating that RUNX3 hypermethylation plays an important role in the malignant transformation of intestinal-type gastric cancer [20, 21]. In addition, chronic gastritis, intestinal metaplasia, gastric adenoma, and gastric cancer show an increasing frequency of p16/cyclin-dependent kinase inhibitor 2A (CDKN2A) methylation [22–24]. This finding indicates that p16/CDKN2A methylation occurs at the initial stage of gastric mucosal malignant transformation and undergoes cumulative change as the disease progresses. Genes related to the malignant transformation of gastric mucosa that undergoes promoter methylation also include the retinoblastoma (RB) gene, von Hippel–Lindau (VHL) tumor suppressor gene, breast cancer 1 (BRCA1) gene, human mutL homolog 1 (hMLH1) gene, X-ray repair cross-complementation group 1 (XRCC1) gene, and ADAM metallopeptidase with thrombospondin type 1 motif 9 (ADAMTS9) gene [25, 26].
In gastric cancer, the distribution characteristics of gene methylation are correlated with biological tumor characteristics and patient prognosis. Patrick Tan and colleagues investigated DNA methylation profiles of 240 primary gastric cancers and gastric cancer cell lines [27]. It has been found that methylomes are widely distributed in gastric cancers. However, these results need to be further verified. In addition, previous data on the methylation of gastric mucosal transformation-related genes are mainly derived from experimental studies of previously established cell lines or small-sized clinical tissue specimen studies. The current knowledge on gene methylation is far from being accurate and comprehensive. Future studies focusing on the following aspects will be more valuable: (1) longitudinal studies: large-scale DNA methylation profiling of clinical tissue specimens—longitudinal, dynamic cohort studies that analyze serial clinical specimens obtained from individual patients at various stages, from inflammation, intestinal metaplasia, and atypical hyperplasia to gastric cancer, are particularly inadequate; a genome-wide longitudinal study of DNA methylation based on such specimens will provide more accurate and comprehensive results; (2) data mining: collection and analysis of the data on gene methylation in normal gastric mucosa, precancerous lesions, and gastric cancer in various populations—exploration of the methylation pattern changes that occur during the malignant transformation of gastric mucosa using large-scale data mining allows a complete understanding of the gene methylation characteristics related to the malignant transformation of gastric mucosa as well as the differentiation of the key methylation changes from the numerous accompanying changes; and (3) non-CpG methylation: non-CpG methylation is an emerging field of research [28, 29]. However, a few data have been obtained for gastric cancer. The research on the distribution, recognition, and regulation of non-CpG methylation in gastric cancer will further deepen our understanding of epigenetic regulation in the transformation of gastric mucosa.
The histone code affects the malignant transformation of gastric mucosa
In eukaryotes, DNA, histones, and non-histone proteins are arranged in a highly ordered pattern to form chromatin. Histones are divided into five classes, including nucleosomal core histones (H2A, H2B, H3, and H4) and linker histones (H1). Each core histone consists of a globular structural domain and an N-terminal tail that is exposed on the surface of the nucleosome. A variety of covalent modifications may occur at the N-terminus of the core histones, including acetylation, methylation, phosphorylation, ubiquitination, and glycosylation. These histone modifications alter the chromatin structure and, therefore, determine the state of gene activation/inactivation; these modifications also regulate physiological processes in the cells. Different histone modifications are orchestrated in both time and space, forming a complex regulatory network known as the “histone code.” Numerous studies have demonstrated histone modification changes in gastric cancer, and such changes are of great clinical significance [30, 31]. Recent studies of gastric cancer-related histone modifications have mainly focused on histone acetylation, methylation, and phosphorylation.
Histone acetylation: histone acetylation is coregulated by histone acetyltransferases (HATs) and histone deacetylases (HDACs). Histone acetylation promotes transcription, whereas histone deacetylation inhibits transcription. Abnormal expression of HDACs and HATs is frequently observed in gastric precancerous lesions and gastric cancer [32, 33]. Current studies have found that histone deacetylation occurs at the promoter region of a number of genes in gastric cancer, including p21(WAF1/CIP1) [34], RIP-associated ICH1/CED3-homologous protein with a death domain (RAIDD) [35], DTW domain containing 1 (DTWD1) [36], p53 upregulated modulator of apoptosis (PUMA) [31], gelsolin and retinoic acid receptor beta [37], deleted in liver cancer-1 (DLC1) [38], and thioredoxin-interacting protein (TXNIP) [39]. In addition, histone deacetylation has been shown to be positively correlated with the downregulated expression of the above genes.
Histone methylation: histone methylation mainly occurs at the lysine (K) and arginine (R) residues of H3 and H4 and is regulated by histone methyltransferases (HMTs) and histone demethylases (HDMs). There are three types of histone methylation: monomethylation, dimethylation, and trimethylation. Different sites and types of histone methylation confer different functions. Methylation of H3K9 and H4K20 inhibits gene expression, whereas methylation of H3K4, H3K36, and H3K79 activates gene expression. H3K27 monomethylation activates gene expression, whereas H3K27 dimethylation and trimethylation inhibit gene expression. For example, H3K27 trimethylation inhibits the expression of Arg kinase-binding protein 2 (ArgBP2) in gastric cancer [40].
Histone phosphorylation: phosphorylation can disrupt the interaction between histones and DNA, and renders the chromatin structure unstable. In addition, phosphorylation may create a surface that binds to protein recognition modules, thereby allowing interaction with specific protein complexes. These two mechanisms enable histone phosphorylation to play a role in chromosome condensation/separation, transcription activation, apoptosis, and DNA damage repair. Studies on the role of histone phosphorylation in the malignant transformation of gastric mucosa are insufficient. Fehri et al. found that H. pylori infection reduces the phosphorylation levels of histone H3S10 and H3T3 in gastric epithelial cells, thus regulating the cell cycle [41]. This finding may represent an important mechanism of H. pylori-induced gastric carcinogenesis. In addition, an increased phosphorylation level of histone H3 was shown to be closely related to the histological type, vascular infiltration, and lymph node metastasis of gastric cancer and is an independent factor associated with a poor prognosis in patients with gastric cancer [42]. The findings indirectly support the hypothesis that histone phosphorylation is involved in the malignant transformation of gastric mucosal cells.
However, relevant studies remain focused on the relationships between the changes in the overall level of certain histone modifications and various pathological states during gastric mucosal carcinogenesis. Studies investigating the mechanisms through which histone modifications affect the malignant transformation of gastric mucosa are currently lacking. To pinpoint the genes or signaling pathways through which histone modifications affect the malignant transformation of gastric mucosa, studies that utilize histone modification-specific antibodies to coprecipitate chromatin or those that employ oligonucleotide microarrays or deep DNA sequencing to identify significantly differentially modified gene loci/chromatin segments and then combine these discoveries with gene function verification would be beneficial.
Non-coding RNAs regulate the malignant transformation of gastric mucosal epithelial cells
Non-coding RNA (ncRNA) is a general term for an RNA molecule that does not encode a protein. ncRNAs include micro RNA (miRNA), piwi-interacting RNA (piRNA), long non-coding RNA (lncRNA), transfer RNA (tRNA), and ribosomal RNA (rRNA). A large number of studies have focused on the roles of miRNAs and lncRNAs in gastric carcinogenesis.
MiRNAs
MiRNAs are a class of evolutionarily conserved, endogenous, non-protein-coding small RNAs. MiRNAs participate in the malignant transformation of gastric mucosal cells by negatively regulating the expression of target genes.
Abnormal expression of miRNA molecules in gastric cancer
Petrocca et al. compared the miRNA expression profiles between tissues with histological signs of chronic gastritis and normal gastric mucosa [43]. It has been found that, in chronic gastritis, the expression of miR-1 and miR-155 is upregulated, whereas the expression of miR-20, miR-26b, miR-202, miR-203, and miR-205 is downregulated. Ueda et al. examined miRNA expression in 160 paired samples of gastric cancer tissues and non-cancerous tissues [44]. The authors found that the expression of 22 miRNAs is upregulated, while the expression of 13 miRNAs is downregulated in gastric cancer tissues compared with the non-cancerous tissues. In addition, 83 % of the patients with gastric cancer could be accurately diagnosed based on miRNA expression profiles in tissue specimens. Microarray analysis has been performed to examine miRNA expression in gastric cancer tissue specimens collected in a large number of countries and geographical regions. The results have shown that the expression of miR-21 [45], miR-27a [46] and miR-196a [47] is significantly elevated in gastric cancer, whereas the expression of lethal-7 (let-7) miRNA [48], miR-101 [49, 50], and miR-29a [51] is markedly reduced. These results clearly demonstrate that miRNAs are involved in gastric carcinogenesis. However, the consistency of previous results is poor. This may be due to sample variation. At present, there is no generally accepted characteristic miRNA expression profile of gastric cancer. Further studies of large and multicenter sample cohort(s) are needed.
Abnormal expression of miRNAs significantly affects the malignant phenotype of gastric cancer cells
The expression of miR-847 is decreased in gastric cancer, which activates the signal transducer and activator of transcription 3 (STAT3)/vascular endothelial growth factor A (VEGF-A) pathway, increases tumor angiogenesis, and promotes the development and progression of gastric cancer [52]. The expression of miR-145 is upregulated in gastric cancer, which inhibits the expression of catenin (cadherin-associated protein), delta 1 (CTNND1), and N-cadherin while promoting the translocation of CTNND1 and E-cadherin from the cytoplasm to the cell membrane. As a result, the proliferation and metastasis of gastric cancer cells are promoted, and the apoptosis of gastric cancer cells is inhibited [53]. The expression of the miR-106b-25 cluster is upregulated in gastric cancer, which inhibits the TGF-β pathway, induces the downregulation of the expression of cyclin-dependent kinase inhibitor 1A (CDKN1A) and BCL2-like 11 apoptosis facilitator (BCL2L11), and promotes the development and progression of gastric cancer [43]. Our study demonstrated that the expression of miR-17-5p is significantly increased in gastric cancer tissues. High miR-17-5p expression inhibits suppressor of cytokine signaling 6 (SOCS6), which promotes the proliferation of gastric cancer cells [54]. The expression of miR-296-5p is abnormally increased in gastric cancer, which inhibits the expression of caudal-related homeobox 1 (CDX1). Furthermore, miR-296-5p/CDX1 affects the phosphorylation level of the extracellular signal-regulated kinases 1 and 2 (ERK1/2) through the mitogen-activated protein kinase (MAPK)/ERK pathway and induces changes in the expression levels of the cell cycle-related protein cyclin D1 and the apoptosis-related proteins B-cell lymphoma 2 (Bcl2) and BCL2-associated X (Bax), thus maintaining the survival of gastric cancer cells and regulating cell proliferation [55]. In addition, miR-150 [56], miR-149 [57], miR-7 [58], miR-199a-5p [59], miR-206 [60, 61], miR-19a/b [62], and miR-218 [63, 64] are all involved in the development and progression of gastric cancer (Fig. 1).
NcRNA-oriented network in the malignant transformation process of gastric cancer. This figure provides insight into the roles of ncRNA and its related protein in the malignant transformation of gastric mucosal cells. It can be seen that a complex network composed of ncRNA and its upstream and downstream components affects the malignant transformation of gastric mucosal cells
LncRNA
LncRNA refers to a class of RNA molecules greater than 200 nucleotides (nt) in length that are transcribed mainly by RNA polymerase II, that lack apparent open reading frames (ORFs) and that do not encode proteins. However, lncRNAs participate in the regulation of a variety of intracellular signaling processes (including tumorigenesis) through the modification of chromatins, activation of transcription, and interference with transcription. Recent studies have shown that several lncRNAs are abnormally expressed in gastric cancer. Moreover, abnormal expression of lncRNAs plays an important role in the development, progression, invasion, and metastasis of gastric cancer.
HOX transcript antisense intergenic RNA (HOTAIR) is located on chromosome 12q13.13. HOTAIR is a 2158-nt lncRNA that possesses a trans-regulatory function. The 5′ end of HOTAIR binds to the initiation complex known as polycomb repressive complex 2 (PRC2). Binding of HOTAIR to PRC2 induces the phosphorylation of EZH2, a subunit of PRC2, at threonine 345 and the subsequent trimethylation of chromosome-bound histone H3K27, thereby inhibiting the expression of the target genes. In addition, the 3′ end of HOTAIR binds to the lysine-specific demethylase 1 (LSD1)/REST corepressor 1 (CoREST)/repressor element 1 (RE1) silencing transcription factor (REST) complex, which mediates histone H3K4me2 demethylation and, thereby, regulates the transcriptional activity of target genes. HOTAIR expression is significantly increased in gastric cancer tissues compared with paracancerous tissues. In diffuse gastric cancer, the high HOTAIR expression group exhibits drastically increased invasion and lymph node metastasis and a decreased overall survival rate in comparison with the low HOTAIR expression group. In addition, studies have shown that inhibition of HOTAIR expression in gastric cancer cells decreases the expression of matrix metalloproteinases 1 and 3, reduces the invasive capability of cancer cells, and reverses the epithelial–mesenchymal transition (EMT) in gastric cancer cells. These findings demonstrate that HOTAIR plays an important role in the development and progression of gastric cancer [65–67]. The H19 gene (full length: 2.5 kb) is located on the human chromosome 11p15.5 region and contains a total of 5 exons and 4 introns. The processed, mature H19 has a length of 2.3 kb. Due to the lack of obvious ORFs, H19 is defined as an lncRNA. H19 expression is significantly elevated in gastric cancer tissues compared with paracancerous tissues [68]. Overexpression of H19 enhances the proliferative capacity of the cells, whereas small interfering RNA (siRNA)-mediated interference of H19 expression enhances apoptosis. The effects of overexpression and downregulation of H19 are related to the inactivation and activation of the TP53 gene. Recent studies have shown that transcription of the H19 gene also produces a mature miRNA, namely, miR-675. H19 is capable of regulating the progression of gastric cancer through the H19/miR-675/runt-related transcription factor 1 (RUNX1) signaling axis [69, 70]. In addition, tumor suppressor candidate 7 (TUSC7) [71], maternally expressed 3 (MEG3) [72], BM742401 [73], colon cancer-associated transcript 1 (CCAT1) [74], and multidrug resistant (MDR)-related and upregulated lncRNA (MRUL) [75] have been found to be differentially expressed between gastric cancer cells and normal gastric mucosal cells and affect the malignant phenotype of gastric cancer cells.
Current studies in the field of ncRNAs have mainly focused on the effects of such molecules on the malignant phenotypes of gastric cancer cells, including growth, proliferation, metastasis, and drug resistance. The conclusion that ncRNAs participate in the malignant transformation of gastric mucosal cells is based on the findings that ncRNAs are differentially expressed between normal gastric mucosal cells and gastric cancer cells and that the differential expression of ncRNAs induces functional changes in certain malignant phenotype of gastric cancer cells. There are virtually no functional studies that directly address the malignant transformation of normal gastric mucosa. In addition, the intrinsic link between various ncRNA molecules remains unclear. We simulated the interactions between a number of gastric carcinogenesis-related molecules that have been identified by our study or reported in the literature. However, further biological experiments are required to discover the ncRNA-regulated network.
H. pylori infection promotes gastric cancer mainly through epigenetic regulation
H. pylori infection is the most important risk factor for gastric cancer. The epigenetic changes induced by H. pylori compose one of the principal molecular mechanisms of gastric carcinogenesis.
H. pylori infection and gene methylation
Numerous studies have demonstrated that H. pylori infection is closely related to abnormal CpG island methylation. Maekita et al. found that methylation levels of all the detected regions were much higher in H. pylori-positive samples than in H. pylori-negative samples among healthy volunteers [76]. Nakajima et al. analyzed the promoter methylation of CpG islands of 48 genes that may be methylated in gastric cancer cell lines. The results showed that 26 genes were consistently methylated in individuals with current or past infection by H. pylori [77]. Shin et al. identified quite distinct methylation profiles according to the presence or absence of current H. pylori infection in non-cancerous gastric mucosae from patients with gastric cancer [78]. Cheng concluded that FOXD3-mediated transcriptional control of tumor suppressors is deregulated by H. pylori infection-induced hypermethylation. This in turn could affect the suppression of gastric tumors [79]. These findings indicate that H. pylori infection potently induces CpG island methylation and may be responsible for the initiation of gastric carcinogenesis.
H. pylori-mediated chronic inflammation is one of the important causes of DNA methylation. A number of studies have suggested that methylation levels in the gastric mucosa after H. pylori infection decrease after H. pylori eradication [80, 81]. These data support the idea that H. pylori-mediated inflammation induces methylation. However, how the inflammation triggers DNA methylation is not yet known.
H. pylori infection and histone modification
A relatively few studies have investigated whether H. pylori affects histone modifications. The infection of gastric epithelial cells with H. pylori leads to hyperacetylation of histone H4 [82], which induces the binding of histone H1 to ATP [83] and causes histone H3 dephosphorylation and deacetylation [41, 84]. These changes result in the abnormal expression of oncogenes and tumor suppressor genes [85, 86], which contributes to malignant transformation of gastric epithelial cells.
H. pylori infection is a major cause of abnormal miRNA expression
MiRNAs play an important role in H. pylori infection-induced malignant transformation of gastric mucosa. Zhang et al. demonstrated for the first time that H. pylori infection is able to induce changes in miRNA expression profiles [87]. They found that miR-21 expression is significantly increased in H. pylori-positive gastric tissues, indicating that the increased expression of miR-21 may be related to H. pylori infection. In AGS human gastric carcinoma cells, H. pylori infection promotes the secretion of nuclear factor kappa B (NF-κB) and interleukin 6 (IL-6) and activates activator protein 1 (AP-1) and STAT3, resulting in significantly upregulated miR-21 expression and drastically enhanced cell proliferative and invasive capabilities. Using miRNA microarrays, Matsushima et al. identified 55 miRNAs that were differentially expressed between H. pylori-positive and H. pylori-negative endoscopic biopsy specimens [88]. Among the 55 miRNAs, the expression of 30 miRNAs was significantly reduced. A portion of the miRNAs (including miR-223, miR-375, and miR-200c) was found to be significantly correlated with gastric mucosal inflammatory activity, chronic inflammation, and H. pylori infection severity scores. Correlation analysis showed that 8 miRNAs can be used to accurately predict whether H. pylori infection is present. Infection of the cells with an H. pylori strain containing the wild-type CagA (cytotoxin-associated gene A) structural domain induced changes in the expression of certain miRNAs (e.g., let-7, miR-125a, and miR-500), whereas H. pylori strains with mutant CagA showed no such effect. Studies conducted by Saito et al. showed that miR-17 and miR-20a are also involved in the gastric cancer-promoting signaling pathways mediated by CagA [89, 90]. CagA activates c-Myc through the activation of the Erk pathway, which further stimulates the expression of miR-17 and miR-20a. MiR-20a is capable of suppressing p21 expression. In addition, miR-146a [91], miR-155 [92], and miR-218 [93] are also involved in H. pylori infection-related malignant transformation of gastric mucosa. In a study conducted by Matsushima et al., patients who tested positive for H. pylori infection were successfully cured with an anti-H. pylori regimen and were reexamined 4 weeks after eradication of H. pylori infection [88]. Cure of the H. pylori infection not only restored the levels of 14 miRNAs whose expression was downregulated during H. pylori infection but also significantly reduced the levels of a portion of the miRNAs whose expression was upregulated by H. pylori infection. This phenomenon indicates that downregulation/inhibition of the expression of cancer-promoting miRNAs using methods, such as oligonucleotides and miRNA sponges combined with the introduction of exogenous cancer-suppressing miRNAs, may reduce or even partially block the promoting effect of H. pylori on gastric cancer.
The effects of various types of epigenetic regulations (such as DNA methylation, histone modification, and ncRNA) on the malignant transformation of gastric mucosa are not independent. Instead, the epigenetic effects interact synergistically to promote gastric carcinogenesis. In addition, different epigenetic changes may coordinate in the regulation of the expression of one carcinogenesis-related gene (Fig. 2). For example, ubiquitin-like containing PHD and RING finger domains 1 (UHRF1) is known to maintain DNA methylation via the recruitment of DNA methyltransferase 1 (DNMT1) [94]. We identified and verified miR-146a/b as direct upstream regulators of UHRF1 [95]. Duursma et al. found that miR-148 targets human DNMT3b [96]. MiR-146a/b and miR-148 can regulate RUNX3 expression via the effects of UHRF1 and DNMT1 on promoter methylation. In addition, increased H3K9 dimethylation and reduced H3 acetylation synergistically inhibit the transcription of RUNX3 [97, 98]. Moreover, miR-130b [99], miR-301a [100], miR-106a [101], miR-103a [102], miR-495 [103], and miR-532-5p [104] directly inhibit RUNX3 translation at the post-transcriptional level. Decreased RUNX3 expression directly downregulates miR-30a expression, which enhances the expression of the miR-30a target gene vimentin and promotes EMT in gastric cancer cells [105]. RUNX3 regulates gastric cancer cell proliferation via the TGF-β [106] and Wnt [107, 108] pathways and affects angiogenesis in gastric cancer by regulating the expression of vascular endothelial growth factor (VEGF) [109]. In contrast, different epigenetic changes may regulate the expression of different carcinogenesis-related genes, whereby they cooperate to promote the malignant transformation of gastric mucosa.
Epigenetic regulation of RUNX3 in the malignant transformation of gastric mucosal cells. MiR-146a/b and miR-148 directly inhibited UHRF1 and DNMT3b, respectively. Downregulation of miR-146a/b and miR-148 led to the increase in UHRF1 and DNMT3b, and this effect in turn inactivated RUNX3 via promoter methylation in gastric cancer. In addition, increased H3K9 dimethylation and reduced H3 acetylation, as well as the increased miR-130b, miR-301a, miR-106a, miR-103a, miR-495, and miR-532-5p, synergistically inhibited the expression of RUNX3
Conclusions
The exploitation of characteristic epigenetic alterations during the malignant transformation of gastric mucosa allows for the prevention, diagnosis, treatment, and prognostic evaluation of gastric cancer from a new perspective independent of protein expression. For example, it has been reported that promoter methylation of death-associated protein kinase (DAPK) [110], E-cadherin [111], and p16 [112] genes may serve as a criterion for sensitive and specific diagnosis of gastric cancer. We have found that the methylation status of the ring finger protein 180 (RNF180) gene may be used to predict the malignant potential of intestinal metaplasia and atypical hyperplasia of gastric mucosa and diagnose early gastric cancer (unpublished data). Currently, we have established a fluorescence-based quantitative technique that enables the analysis of RNF180 gene methylation and meets the registration requirements for diagnostic reagents. We have initiated a clinical trial application to test this diagnostic kit.
The reversibility of epigenetic alterations (such as DNA methylation and histone modification) has recently become a hot topic in drug development. Studies have found that inhibition of deacetylase may suppress the malignant phenotype of gastric cancer cells [39, 113–116] and increase the sensitivity of gastric cancer cells to chemotherapy [115, 117, 118]. Histone deacetylase inhibitors, such as Vorinostat (Zolinza, suberoylanilide hydroxamic acid, SAHA) developed by Merck & Co., Inc. (USA) and Chidamide developed in China, have been successfully used in the clinical treatment of cancer [119–121]. Currently, a number of DNA methyltransferase inhibitors (DNMTi) and histone deacetylase inhibitors are undergoing clinical trials to assess their safety and efficacy for the treatment of tumors [122, 123].
However, unfavorable epigenetic changes induced by epigenetic drugs may lead to severe side effects. The discovery of ways by which the specificity against tumor cells can be enhanced and the side effects can be reduced is still a hot topic. The epigenetic modifications and their combinations that are involved in the malignant transformation of gastric mucosa are highly complex and diverse. There are many outstanding issues requiring clarification. In-depth studies and further elucidation of the epigenetic networks that regulate the malignant transformation of gastric mucosa will provide a wealth of pathways and targets for understanding gastric cancer development and progression, conducting molecular typing, establishing new therapies, and developing new drugs.
References
Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A (2015) Global cancer statistics, 2012. CA Cancer J Clin 65(2):87–108. doi:10.3322/caac.21262
Chen W (2015) Cancer statistics: updated cancer burden in China. Chin J Cancer Res Chung-kuo Yen Cheng Yen Chiu 27(1):1. doi:10.3978/j.issn.1000-9604.2015.02.07
Ishige T, Nishimura M, Satoh M, Fujimoto M, Fukuyo M, Semba T, Kado S, Tsuchida S, Sawai S, Matsushita K, Togawa A, Matsubara H, Kaneda A, Nomura F (2016) Combined secretomics and transcriptomics revealed cancer-derived GDF15 is involved in diffuse-type gastric cancer progression and fibroblast activation. Sci Rep 6:21681. doi:10.1038/srep21681
Ushiku T, Ishikawa S, Kakiuchi M, Tanaka A, Katoh H, Aburatani H, Lauwers GY, Fukayama M (2016) RHOA mutation in diffuse-type gastric cancer: a comparative clinicopathology analysis of 87 cases. Gastric Cancer 19(2):403–411. doi:10.1007/s10120-015-0493-0
Wang K, Kan J, Yuen ST, Shi ST, Chu KM, Law S, Chan TL, Kan Z, Chan AS, Tsui WY, Lee SP, Ho SL, Chan AK, Cheng GH, Roberts PC, Rejto PA, Gibson NW, Pocalyko DJ, Mao M, Xu J, Leung SY (2011) Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat Genet 43(12):1219–1223. doi:10.1038/ng.982
Liang Q, Yao X, Tang S, Zhang J, Yau TO, Li X, Tang CM, Kang W, Lung RW, Li JW, Chan TF, Xing R, Lu Y, Lo KW, Wong N, To KF, Yu C, Chan FK, Sung JJ, Yu J (2014) Integrative identification of Epstein-Barr virus-associated mutations and epigenetic alterations in gastric cancer. Gastroenterology 147(6):1350–1362. doi:10.1053/j.gastro.2014.08.036 (e1354)
Waddington CH (1939) Preliminary notes on the development of the wings in normal and mutant strains of Drosophila. Proc Natl Acad Sci USA 25(7):299–307
Dobrilla G, Benvenuti S, Amplatz S, Zancanella L (1994) Chronic gastritis, intestinal metaplasia, dysplasia and Helicobacter pylori in gastric cancer: putting the pieces together. Ital J Gastroenterol 26(9):449–458
Correa P (1995) Helicobacter pylori and gastric carcinogenesis. Am J Surg Pathol 19(Suppl 1):S37–S43
Gigek CO, Chen ES, Calcagno DQ, Wisnieski F, Burbano RR, Smith MA (2012) Epigenetic mechanisms in gastric cancer. Epigenomics 4(3):279–294. doi:10.2217/epi.12.22
Calcagno DQ, de Arruda Cardoso Smith M, Burbano RR (2015) Cancer type-specific epigenetic changes: gastric cancer. Methods Mol Biol 1238:79–101. doi:10.1007/978-1-4939-1804-1_5
Bird AP (1986) CpG-rich islands and the function of DNA methylation. Nature 321(6067):209–213. doi:10.1038/321209a0
Stefanska B, Huang J, Bhattacharyya B, Suderman M, Hallett M, Han ZG, Szyf M (2011) Definition of the landscape of promoter DNA hypomethylation in liver cancer. Cancer Res 71(17):5891–5903. doi:10.1158/0008-5472.CAN-10-3823
Wojdacz TK, Windelov JA, Thestrup BB, Damsgaard TE, Overgaard J, Hansen L (2014) Identification and characterization of locus-specific methylation patterns within novel loci undergoing hypermethylation during breast cancer pathogenesis. Breast Cancer Res 16(1):R17. doi:10.1186/bcr3612
Aran D, Hellman A (2013) DNA methylation of transcriptional enhancers and cancer predisposition. Cell 154(1):11–13. doi:10.1016/j.cell.2013.06.018
Grady WM, Willis J, Guilford PJ, Dunbier AK, Toro TT, Lynch H, Wiesner G, Ferguson K, Eng C, Park JG, Kim SJ, Markowitz S (2000) Methylation of the CDH1 promoter as the second genetic hit in hereditary diffuse gastric cancer. Nat Genet 26(1):16–17. doi:10.1038/79120
Chan AO, Lam SK, Wong BC, Wong WM, Yuen MF, Yeung YH, Hui WM, Rashid A, Kwong YL (2003) Promoter methylation of E-cadherin gene in gastric mucosa associated with Helicobacter pylori infection and in gastric cancer. Gut 52(4):502–506
Senekowitsch-Schmidtke R, Schuhmacher C, Becker KF, Nikula TK, Seidl C, Becker I, Miederer M, Apostolidis C, Adam C, Huber R, Kremmer E, Fischer K, Schwaiger M (2001) Highly specific tumor binding of a 213Bi-labeled monoclonal antibody against mutant E-cadherin suggests its usefulness for locoregional alpha-radioimmunotherapy of diffuse-type gastric cancer. Cancer Res 61(7):2804–2808
Kim TY, Lee HJ, Hwang KS, Lee M, Kim JW, Bang YJ, Kang GH (2004) Methylation of RUNX3 in various types of human cancers and premalignant stages of gastric carcinoma. Lab Invest 84(4):479–484. doi:10.1038/labinvest.3700060
Sato F, Meltzer SJ (2006) CpG island hypermethylation in progression of esophageal and gastric cancer. Cancer 106(3):483–493. doi:10.1002/cncr.21657
Lu XX, Yu JL, Ying LS, Han J, Wang S, Yu QM, Wang XB, Fang XH, Ling ZQ (2012) Stepwise cumulation of RUNX3 methylation mediated by Helicobacter pylori infection contributes to gastric carcinoma progression. Cancer 118(22):5507–5517. doi:10.1002/cncr.27604
Lu ZM, Zhou J, Wang X, Guan Z, Bai H, Liu ZJ, Su N, Pan K, Ji J, Deng D (2012) Nucleosomes correlate with in vivo progression pattern of de novo methylation of p16 CpG islands in human gastric carcinogenesis. PLoS One 7(4):e35928. doi:10.1371/journal.pone.0035928
Kang GH, Lee S, Kim JS, Jung HY (2003) Profile of aberrant CpG island methylation along the multistep pathway of gastric carcinogenesis. Lab Invest 83(5):635–641
Kang GH, Lee S, Kim JS, Jung HY (2003) Profile of aberrant CpG island methylation along multistep gastric carcinogenesis. Lab Invest 83(4):519–526
Kim H, Kim YH, Kim SE, Kim NG, Noh SH, Kim H (2003) Concerted promoter hypermethylation of hMLH1, p16INK4A, and E-cadherin in gastric carcinomas with microsatellite instability. J Pathol 200(1):23–31. doi:10.1002/path.1325
Tahara T, Arisawa T (2015) DNA methylation as a molecular biomarker in gastric cancer. Epigenomics 7(3):475–486. doi:10.2217/epi.15.4
Zouridis H, Deng N, Ivanova T, Zhu Y, Wong B, Huang D, Wu YH, Wu Y, Tan IB, Liem N, Gopalakrishnan V, Luo Q, Wu J, Lee M, Yong WP, Goh LK, Teh BT, Rozen S, Tan P (2012) Methylation subtypes and large-scale epigenetic alterations in gastric cancer. Sci Transl Med 4(156):156ra140. doi:10.1126/scitranslmed.3004504
Sharma G, Sowpati DT, Singh P, Khan MZ, Ganji R, Upadhyay S, Banerjee S, Nandicoori VK, Khosla S (2016) Genome-wide non-CpG methylation of the host genome during M. tuberculosis infection. Sci Rep 6:25006. doi:10.1038/srep25006
Pietrzak M, Rempala GA, Nelson PT, Hetman M (2016) Non-random distribution of methyl-CpG sites and non-CpG methylation in the human rDNA promoter identified by next generation bisulfite sequencing. Gene 585(1):35–43. doi:10.1016/j.gene.2016.03.028
Berger SL (2001) Molecular biology. The histone modification circus. Science 292(5514):64–65
Yang Y, Yin X, Yang H, Xu Y (2015) Histone demethylase LSD2 acts as an E3 ubiquitin ligase and inhibits cancer cell growth through promoting proteasomal degradation of OGT. Mol Cell 58(1):47–59. doi:10.1016/j.molcel.2015.01.038
Song J, Noh JH, Lee JH, Eun JW, Ahn YM, Kim SY, Lee SH, Park WS, Yoo NJ, Lee JY, Nam SW (2005) Increased expression of histone deacetylase 2 is found in human gastric cancer. APMIS 113(4):264–268. doi:10.1111/j.1600-0463.2005.apm_04.x
Sudo T, Mimori K, Nishida N, Kogo R, Iwaya T, Tanaka F, Shibata K, Fujita H, Shirouzu K, Mori M (2011) Histone deacetylase 1 expression in gastric cancer. Oncol Rep 26(4):777–782. doi:10.3892/or.2011.1361
Mitani Y, Oue N, Hamai Y, Aung PP, Matsumura S, Nakayama H, Kamata N, Yasui W (2005) Histone H3 acetylation is associated with reduced p21(WAF1/CIP1) expression by gastric carcinoma. J Pathol 205(1):65–73. doi:10.1002/path.1684
Shen Q, Tang W, Sun J, Feng L, Jin H, Wang X (2014) Regulation of CRADD-caspase 2 cascade by histone deacetylase 1 in gastric cancer. Am J Transl Res 6(5):538–547
Ma Y, Yue Y, Pan M, Sun J, Chu J, Lin X, Xu W, Feng L, Chen Y, Chen D, Shin VY, Wang X, Jin H (2015) Histone deacetylase 3 inhibits new tumor suppressor gene DTWD1 in gastric cancer. Am J Cancer Res 5(2):663–673
Kim JH, Choi YK, Kwon HJ, Yang HK, Choi JH, Kim DY (2004) Downregulation of gelsolin and retinoic acid receptor beta expression in gastric cancer tissues through histone deacetylase 1. J Gastroenterol Hepatol 19(2):218–224
Kim TY, Kim IS, Jong HS, Lee JW, Kim TY, Jung M, Bang YJ (2008) Transcriptional induction of DLC-1 gene through Sp1 sites by histone deacetylase inhibitors in gastric cancer cells. Exp Mol Med 40(6):639–646. doi:10.3858/emm.2008.40.6.639
Lee JH, Jeong EG, Choi MC, Kim SH, Park JH, Song SH, Park J, Bang YJ, Kim TY (2010) Inhibition of histone deacetylase 10 induces thioredoxin-interacting protein and causes accumulation of reactive oxygen species in SNU-620 human gastric cancer cells. Mol Cells 30(2):107–112. doi:10.1007/s10059-010-0094-z
Tong Y, Li Y, Gu H, Wang C, Liu F, Shao Y, Li J, Cao L, Li F (2015) MORC2 downregulates ArgBP2 via histone methylation in gastric cancer cells. Biochem Biophys Res Commun. doi:10.1016/j.bbrc.2015.10.059
Fehri LF, Rechner C, Janssen S, Mak TN, Holland C, Bartfeld S, Bruggemann H, Meyer TF (2009) Helicobacter pylori-induced modification of the histone H3 phosphorylation status in gastric epithelial cells reflects its impact on cell cycle regulation. Epigenetics 4(8):577–586
Takahashi H, Murai Y, Tsuneyama K, Nomoto K, Okada E, Fujita H, Takano Y (2006) Overexpression of phosphorylated histone H3 is an indicator of poor prognosis in gastric adenocarcinoma patients. Appl Immunohistochem Mol Morphol 14(3):296–302
Petrocca F, Visone R, Onelli MR, Shah MH, Nicoloso MS, de Martino I, Iliopoulos D, Pilozzi E, Liu CG, Negrini M, Cavazzini L, Volinia S, Alder H, Ruco LP, Baldassarre G, Croce CM, Vecchione A (2008) E2F1-regulated microRNAs impair TGFbeta-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13(3):272–286. doi:10.1016/j.ccr.2008.02.013
Ueda T, Volinia S, Okumura H, Shimizu M, Taccioli C, Rossi S, Alder H, Liu CG, Oue N, Yasui W, Yoshida K, Sasaki H, Nomura S, Seto Y, Kaminishi M, Calin GA, Croce CM (2010) Relation between microRNA expression and progression and prognosis of gastric cancer: a microRNA expression analysis. Lancet Oncol 11(2):136–146. doi:10.1016/S1470-2045(09)70343-2
Zhang Z, Li Z, Gao C, Chen P, Chen J, Liu W, Xiao S, Lu H (2008) miR-21 plays a pivotal role in gastric cancer pathogenesis and progression. Lab Invest 88(12):1358–1366. doi:10.1038/labinvest.2008.94
Liu T, Tang H, Lang Y, Liu M, Li X (2009) MicroRNA-27a functions as an oncogene in gastric adenocarcinoma by targeting prohibitin. Cancer Lett 273(2):233–242. doi:10.1016/j.canlet.2008.08.003
Sun M, Liu XH, Li JH, Yang JS, Zhang EB, Yin DD, Liu ZL, Zhou J, Ding Y, Li SQ, Wang ZX, Cao XF, De W (2012) MiR-196a is upregulated in gastric cancer and promotes cell proliferation by downregulating p27(kip1). Mol Cancer Ther 11(4):842–852. doi:10.1158/1535-7163.MCT-11-1015
Motoyama K, Inoue H, Nakamura Y, Uetake H, Sugihara K, Mori M (2008) Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin Cancer Res 14(8):2334–2340. doi:10.1158/1078-0432.CCR-07-4667
Zhou X, Xia Y, Li L, Zhang G (2015) MiR-101 inhibits cell growth and tumorigenesis of Helicobacter pylori related gastric cancer by repression of SOCS2. Cancer Biol Ther 16(1):160–169. doi:10.4161/15384047.2014.987523
He XP, Shao Y, Li XL, Xu W, Chen GS, Sun HH, Xu HC, Xu X, Tang D, Zheng XF, Xue YP, Huang GC, Sun WH (2012) Downregulation of miR-101 in gastric cancer correlates with cyclooxygenase-2 overexpression and tumor growth. FEBS J 279(22):4201–4212. doi:10.1111/febs.12013
Cui Y, Su WY, Xing J, Wang YC, Wang P, Chen XY, Shen ZY, Cao H, Lu YY, Fang JY (2011) MiR-29a inhibits cell proliferation and induces cell cycle arrest through the downregulation of p42.3 in human gastric cancer. PLoS One 6(10):e25872. doi:10.1371/journal.pone.0025872
Zhang X, Tang J, Zhi X, Xie K, Wang W, Li Z, Zhu Y, Yang L, Xu H, Xu Z (2015) miR-874 functions as a tumor suppressor by inhibiting angiogenesis through STAT3/VEGF-A pathway in gastric cancer. Oncotarget 6(3):1605–1617. doi:10.18632/oncotarget.2748
Xing AY, Wang YW, Su ZX, Shi DB, Wang B, Gao P (2015) Catenin-delta1, negatively regulated by miR-145, promotes tumour aggressiveness in gastric cancer. J Pathol 236(1):53–64. doi:10.1002/path.4495
Wu Q, Luo G, Yang Z, Zhu F, An Y, Shi Y, Fan D (2014) miR-17-5p promotes proliferation by targeting SOCS6 in gastric cancer cells. FEBS Lett 588(12):2055–2062. doi:10.1016/j.febslet.2014.04.036
Li T, Lu YY, Zhao XD, Guo HQ, Liu CH, Li H, Zhou L, Han YN, Wu KC, Nie YZ, Shi YQ, Fan DM (2014) MicroRNA-296-5p increases proliferation in gastric cancer through repression of Caudal-related homeobox 1. Oncogene 33(6):783–793. doi:10.1038/onc.2012.637
Wu Q, Jin H, Yang Z, Luo G, Lu Y, Li K, Ren G, Su T, Pan Y, Feng B, Xue Z, Wang X, Fan D (2010) MiR-150 promotes gastric cancer proliferation by negatively regulating the pro-apoptotic gene EGR2. Biochem Biophys Res Commun 392(3):340–345. doi:10.1016/j.bbrc.2009.12.182
Wang Y, Zheng X, Zhang Z, Zhou J, Zhao G, Yang J, Xia L, Wang R, Cai X, Hu H, Zhu C, Nie Y, Wu K, Zhang D, Fan D (2012) MicroRNA-149 inhibits proliferation and cell cycle progression through the targeting of ZBTB2 in human gastric cancer. PLoS One 7(10):e41693. doi:10.1371/journal.pone.0041693
Zhao X, Dou W, He L, Liang S, Tie J, Liu C, Li T, Lu Y, Mo P, Shi Y, Wu K, Nie Y, Fan D (2013) MicroRNA-7 functions as an anti-metastatic microRNA in gastric cancer by targeting insulin-like growth factor-1 receptor. Oncogene 32(11):1363–1372. doi:10.1038/onc.2012.156
Zhao X, He L, Li T, Lu Y, Miao Y, Liang S, Guo H, Bai M, Xie H, Luo G, Zhou L, Shen G, Guo C, Bai F, Sun S, Wu K, Nie Y, Fan D (2014) SRF expedites metastasis and modulates the epithelial to mesenchymal transition by regulating miR-199a-5p expression in human gastric cancer. Cell Death Differ 21(12):1900–1913. doi:10.1038/cdd.2014.109
Zhang L, Xia L, Zhao L, Chen Z, Shang X, Xin J, Liu M, Guo X, Wu K, Pan Y, Fan D (2015) Activation of PAX3-MET pathways due to miR-206 loss promotes gastric cancer metastasis. Carcinogenesis 36(3):390–399. doi:10.1093/carcin/bgv009
Zhang L, Liu X, Jin H, Guo X, Xia L, Chen Z, Bai M, Liu J, Shang X, Wu K, Pan Y, Fan D (2013) miR-206 inhibits gastric cancer proliferation in part by repressing cyclinD2. Cancer Lett 332(1):94–101. doi:10.1016/j.canlet.2013.01.023
Wu Q, Yang Z, Wang F, Hu S, Yang L, Shi Y, Fan D (2013) MiR-19b/20a/92a regulates the self-renewal and proliferation of gastric cancer stem cells. J Cell Sci 126(Pt 18):4220–4229. doi:10.1242/jcs.127944
Tie J, Pan Y, Zhao L, Wu K, Liu J, Sun S, Guo X, Wang B, Gang Y, Zhang Y, Li Q, Qiao T, Zhao Q, Nie Y, Fan D (2010) MiR-218 inhibits invasion and metastasis of gastric cancer by targeting the Robo1 receptor. PLoS Genet 6(3):e1000879. doi:10.1371/journal.pgen.1000879
Wang SM, Tie J, Wang WL, Hu SJ, Yin JP, Yi XF, Tian ZH, Zhang XY, Li MB, Li ZS, Nie YZ, Wu KC, Fan DM (2015) POU2F2-oriented network promotes human gastric cancer metastasis. Gut. doi:10.1136/gutjnl-2014-308932
Endo H, Shiroki T, Nakagawa T, Yokoyama M, Tamai K, Yamanami H, Fujiya T, Sato I, Yamaguchi K, Tanaka N, Iijima K, Shimosegawa T, Sugamura K, Satoh K (2013) Enhanced expression of long non-coding RNA HOTAIR is associated with the development of gastric cancer. PLoS One 8(10):e77070. doi:10.1371/journal.pone.0077070
Du M, Wang W, Jin H, Wang Q, Ge Y, Lu J, Ma G, Chu H, Tong N, Zhu H, Wang M, Qiang F, Zhang Z (2015) The association analysis of lncRNA HOTAIR genetic variants and gastric cancer risk in a Chinese population. Oncotarget 6(31):31255–31262. doi:10.18632/oncotarget.5158
Xu ZY, Yu QM, Du YA, Yang LT, Dong RZ, Huang L, Yu PF, Cheng XD (2013) Knockdown of long non-coding RNA HOTAIR suppresses tumor invasion and reverses epithelial-mesenchymal transition in gastric cancer. Int J Biol Sci 9(6):587–597. doi:10.7150/ijbs.6339
Yang F, Bi J, Xue X, Zheng L, Zhi K, Hua J, Fang G (2012) Up-regulated long non-coding RNA H19 contributes to proliferation of gastric cancer cells. FEBS J 279(17):3159–3165. doi:10.1111/j.1742-4658.2012.08694.x
Zhuang M, Gao W, Xu J, Wang P, Shu Y (2014) The long non-coding RNA H19-derived miR-675 modulates human gastric cancer cell proliferation by targeting tumor suppressor RUNX1. Biochem Biophys Res Commun 448(3):315–322. doi:10.1016/j.bbrc.2013.12.126
Li H, Yu B, Li J, Su L, Yan M, Zhu Z, Liu B (2014) Overexpression of lncRNA H19 enhances carcinogenesis and metastasis of gastric cancer. Oncotarget 5(8):2318–2329. doi:10.18632/oncotarget.1913
Qi P, Xu MD, Shen XH, Ni SJ, Huang D, Tan C, Weng WW, Sheng WQ, Zhou XY, Du X (2015) Reciprocal repression between TUSC7 and miR-23b in gastric cancer. Int J Cancer 137(6):1269–1278. doi:10.1002/ijc.29516
Sun M, Xia R, Jin F, Xu T, Liu Z, De W, Liu X (2014) Downregulated long noncoding RNA MEG3 is associated with poor prognosis and promotes cell proliferation in gastric cancer. Tumour Biol 35(2):1065–1073. doi:10.1007/s13277-013-1142-z
Park SM, Park SJ, Kim HJ, Kwon OH, Kang TW, Sohn HA, Kim SK, Moo Noh S, Song KS, Jang SJ, Sung Kim Y, Kim SY (2013) A known expressed sequence tag, BM742401, is a potent lincRNA inhibiting cancer metastasis. Exp Mol Med 45:e31. doi:10.1038/emm.2013.59
Mizrahi I, Mazeh H, Grinbaum R, Beglaibter N, Wilschanski M, Pavlov V, Adileh M, Stojadinovic A, Avital I, Gure AO, Halle D, Nissan A (2015) Colon cancer associated transcript-1 (CCAT1) expression in adenocarcinoma of the stomach. J Cancer 6(2):105–110. doi:10.7150/jca.10568
Wang Y, Zhang D, Wu K, Zhao Q, Nie Y, Fan D (2014) Long noncoding RNA MRUL promotes ABCB1 expression in multidrug-resistant gastric cancer cell sublines. Mol Cell Biol 34(17):3182–3193. doi:10.1128/MCB.01580-13
Maekita T, Nakazawa K, Mihara M, Nakajima T, Yanaoka K, Iguchi M, Arii K, Kaneda A, Tsukamoto T, Tatematsu M, Tamura G, Saito D, Sugimura T, Ichinose M, Ushijima T (2006) High levels of aberrant DNA methylation in Helicobacter pylori-infected gastric mucosae and its possible association with gastric cancer risk. Clin Cancer Res 12(3 Pt 1):989–995. doi:10.1158/1078-0432.CCR-05-2096
Nakajima T, Yamashita S, Maekita T, Niwa T, Nakazawa K, Ushijima T (2009) The presence of a methylation fingerprint of Helicobacter pylori infection in human gastric mucosae. Int J Cancer 124(4):905–910. doi:10.1002/ijc.24018
Shin CM, Kim N, Jung Y, Park JH, Kang GH, Park WY, Kim JS, Jung HC, Song IS (2011) Genome-wide DNA methylation profiles in noncancerous gastric mucosae with regard to Helicobacter pylori infection and the presence of gastric cancer. Helicobacter 16(3):179–188. doi:10.1111/j.1523-5378.2011.00838.x
Cheng AS, Li MS, Kang W, Cheng VY, Chou JL, Lau SS, Go MY, Lee CC, Ling TK, Ng EK, Yu J, Huang TH, To KF, Chan MW, Sung JJ, Chan FK (2013) Helicobacter pylori causes epigenetic dysregulation of FOXD3 to promote gastric carcinogenesis. Gastroenterology 144(1):122–133. doi:10.1053/j.gastro.2012.10.002 (e129)
Touati E (2010) When bacteria become mutagenic and carcinogenic: lessons from H. pylori. Mutat Res 703(1):66–70. doi:10.1016/j.mrgentox.2010.07.014
Nakajima T, Enomoto S, Yamashita S, Ando T, Nakanishi Y, Nakazawa K, Oda I, Gotoda T, Ushijima T (2010) Persistence of a component of DNA methylation in gastric mucosae after Helicobacter pylori eradication. J Gastroenterol 45(1):37–44. doi:10.1007/s00535-009-0142-7
Xia G, Schneider-Stock R, Diestel A, Habold C, Krueger S, Roessner A, Naumann M, Lendeckel U (2008) Helicobacter pylori regulates p21(WAF1) by histone H4 acetylation. Biochem Biophys Res Commun 369(2):526–531. doi:10.1016/j.bbrc.2008.02.073
Turkina MV, Olofsson A, Magnusson KE, Arnqvist A, Vikstrom E (2015) Helicobacter pylori vesicles carrying CagA localize in the vicinity of cell-cell contacts and induce histone H1 binding to ATP in epithelial cells. FEMS Microbiol Lett 362(11). doi:10.1093/femsle/fnv076
Ding SZ, Fischer W, Kaparakis-Liaskos M, Liechti G, Merrell DS, Grant PA, Ferrero RL, Crowe SE, Haas R, Hatakeyama M, Goldberg JB (2010) Helicobacter pylori-induced histone modification, associated gene expression in gastric epithelial cells, and its implication in pathogenesis. PLoS One 5(4):e9875. doi:10.1371/journal.pone.0009875
Byun SW, Chang YJ, Chung IS, Moss SF, Kim SS (2012) Helicobacter pylori decreases p27 expression through the delta opioid receptor-mediated inhibition of histone acetylation within the p27 promoter. Cancer Lett 326(1):96–104. doi:10.1016/j.canlet.2012.07.032
Liang X, Zeng J, Wang L, Shen L, Li S, Ma L, Ci X, Yu J, Jia M, Sun Y, Liu Z, Liu S, Li W, Yu H, Chen C, Jia J (2014) Histone demethylase RBP2 induced by Helicobacter pylori CagA participates in the malignant transformation of gastric epithelial cells. Oncotarget 5(14):5798–5807. doi:10.18632/oncotarget.2185
Zhang Z, Li Z, Gao C, Chen P, Chen J, Liu W, Xiao S, Lu H (2008) miR-21 plays a pivotal role in gastric cancer pathogenesis and progression. Lab Invest. doi:10.1038/labinvest.2008.94
Matsushima K, Isomoto H, Inoue N, Nakayama T, Hayashi T, Nakayama M, Nakao K, Hirayama T, Kohno S (2011) MicroRNA signatures in Helicobacter pylori-infected gastric mucosa. Int J Cancer 128(2):361–370. doi:10.1002/ijc.25348
Saf C, Gulcan EM, Ozkan F, Cobanoglu Saf SP, Vitrinel A (2015) Assessment of p21, p53 expression, and Ki-67 proliferative activities in the gastric mucosa of children with Helicobacter pylori gastritis. Eur J Gastroenterol Hepatol 27(2):155–161. doi:10.1097/MEG.0000000000000246
Saito Y, Murata-Kamiya N, Hirayama T, Ohba Y, Hatakeyama M (2010) Conversion of Helicobacter pylori CagA from senescence inducer to oncogenic driver through polarity-dependent regulation of p21. J Exp Med 207(10):2157–2174. doi:10.1084/jem.20100602
Liu Z, Xiao B, Tang B, Li B, Li N, Zhu E, Guo G, Gu J, Zhuang Y, Liu X, Ding H, Zhao X, Guo H, Mao X, Zou Q (2010) Up-regulated microRNA-146a negatively modulate Helicobacter pylori-induced inflammatory response in human gastric epithelial cells. Microbes Infect/Inst Pasteur 12(11):854–863. doi:10.1016/j.micinf.2010.06.002
Crone SG, Jacobsen A, Federspiel B, Bardram L, Krogh A, Lund AH, Friis-Hansen L (2012) microRNA-146a inhibits G protein-coupled receptor-mediated activation of NF-kappaB by targeting CARD10 and COPS8 in gastric cancer. Mol Cancer 11:71. doi:10.1186/1476-4598-11-71
Gao C, Zhang Z, Liu W, Xiao S, Gu W, Lu H (2010) Reduced microRNA-218 expression is associated with high nuclear factor kappa B activation in gastric cancer. Cancer 116(1):41–49. doi:10.1002/cncr.24743
Bashtrykov P, Jankevicius G, Jurkowska RZ, Ragozin S, Jeltsch A (2014) The UHRF1 protein stimulates the activity and specificity of the maintenance DNA methyltransferase DNMT1 by an allosteric mechanism. J Biol Chem 289(7):4106–4115. doi:10.1074/jbc.M113.528893
Zhou L, Zhao X, Han Y, Lu Y, Shang Y, Liu C, Li T, Jin Z, Fan D, Wu K (2013) Regulation of UHRF1 by miR-146a/b modulates gastric cancer invasion and metastasis. FASEB J 27(12):4929–4939. doi:10.1096/fj.13-233387
Duursma AM, Kedde M, Schrier M, le Sage C, Agami R (2008) miR-148 targets human DNMT3b protein coding region. RNA 14(5):872–877. doi:10.1261/rna.972008
Fujii S, Ito K, Ito Y, Ochiai A (2008) Enhancer of zeste homologue 2 (EZH2) down-regulates RUNX3 by increasing histone H3 methylation. J Biol Chem 283(25):17324–17332. doi:10.1074/jbc.M800224200
Lee SH, Kim J, Kim WH, Lee YM (2009) Hypoxic silencing of tumor suppressor RUNX3 by histone modification in gastric cancer cells. Oncogene 28(2):184–194. doi:10.1038/onc.2008.377
Lai KW, Koh KX, Loh M, Tada K, Subramaniam MM, Lim XY, Vaithilingam A, Salto-Tellez M, Iacopetta B, Ito Y, Soong R, Singapore Gastric Cancer C (2010) MicroRNA-130b regulates the tumour suppressor RUNX3 in gastric cancer. Eur J Cancer 46(8):1456–1463. doi:10.1016/j.ejca.2010.01.036
Wang M, Li C, Yu B, Su L, Li J, Ju J, Yu Y, Gu Q, Zhu Z, Liu B (2013) Overexpressed miR-301a promotes cell proliferation and invasion by targeting RUNX3 in gastric cancer. J Gastroenterol 48(9):1023–1033. doi:10.1007/s00535-012-0733-6
Zhang Y, Lu Q, Cai X (2013) MicroRNA-106a induces multidrug resistance in gastric cancer by targeting RUNX3. FEBS Lett 587(18):3069–3075. doi:10.1016/j.febslet.2013.06.058
Jiang H, Yu WW, Wang LL, Peng Y (2015) miR-130a acts as a potential diagnostic biomarker and promotes gastric cancer migration, invasion and proliferation by targeting RUNX3. Oncol Rep 34(3):1153–1161. doi:10.3892/or.2015.4099
Lee SH, Jung YD, Choi YS, Lee YM (2015) Targeting of RUNX3 by miR-130a and miR-495 cooperatively increases cell proliferation and tumor angiogenesis in gastric cancer cells. Oncotarget 6(32):33269–33278. doi:10.18632/oncotarget.5037
Xu X, Zhang Y, Liu Z, Zhang X, Jia J (2016) miRNA-532-5p functions as an oncogenic microRNA in human gastric cancer by directly targeting RUNX3. J Cell Mol Med 20(1):95–103. doi:10.1111/jcmm.12706
Liu Z, Chen L, Zhang X, Xu X, Xing H, Zhang Y, Li W, Yu H, Zeng J, Jia J (2014) RUNX3 regulates vimentin expression via miR-30a during epithelial-mesenchymal transition in gastric cancer cells. J Cell Mol Med 18(4):610–623. doi:10.1111/jcmm.12209
Yano T, Ito K, Fukamachi H, Chi XZ, Wee HJ, Inoue K, Ida H, Bouillet P, Strasser A, Bae SC, Ito Y (2006) The RUNX3 tumor suppressor upregulates Bim in gastric epithelial cells undergoing transforming growth factor beta-induced apoptosis. Mol Cell Biol 26(12):4474–4488. doi:10.1128/MCB.01926-05
Ito K (2011) RUNX3 in oncogenic and anti-oncogenic signaling in gastrointestinal cancers. J Cell Biochem 112(5):1243–1249. doi:10.1002/jcb.23047
Ito K, Lim AC, Salto-Tellez M, Motoda L, Osato M, Chuang LS, Lee CW, Voon DC, Koo JK, Wang H, Fukamachi H, Ito Y (2008) RUNX3 attenuates beta-catenin/T cell factors in intestinal tumorigenesis. Cancer Cell 14(3):226–237. doi:10.1016/j.ccr.2008.08.004
Peng Z, Wei D, Wang L, Tang H, Zhang J, Le X, Jia Z, Li Q, Xie K (2006) RUNX3 inhibits the expression of vascular endothelial growth factor and reduces the angiogenesis, growth, and metastasis of human gastric cancer. Clin Cancer Res 12(21):6386–6394. doi:10.1158/1078-0432.CCR-05-2359
Kato K, Iida S, Uetake H, Takagi Y, Yamashita T, Inokuchi M, Yamada H, Kojima K, Sugihara K (2008) Methylated TMS1 and DAPK genes predict prognosis and response to chemotherapy in gastric cancer. Int J Cancer 122(3):603–608. doi:10.1002/ijc.23143
Xing X, Tang YB, Yuan G, Wang Y, Wang J, Yang Y, Chen M (2013) The prognostic value of E-cadherin in gastric cancer: a meta-analysis. Int J Cancer 132(11):2589–2596. doi:10.1002/ijc.27947
Peng D, Zhang H, Sun G (2014) The relationship between P16 gene promoter methylation and gastric cancer: a meta-analysis based on Chinese patients. J Cancer Res Ther 10(Suppl):292–295. doi:10.4103/0973-1482.151535
Lee KH, Choi EY, Kim MK, Kim KO, Jang BI, Kim SW, Kim SW, Song SK, Kim JR (2010) Inhibition of histone deacetylase activity down-regulates urokinase plasminogen activator and matrix metalloproteinase-9 expression in gastric cancer. Mol Cell Biochem 343(1–2):163–171. doi:10.1007/s11010-010-0510-x
Lin L, Jiang H, Huang M, Hou X, Sun X, Jiang X, Dong X, Sun X, Zhou B, Qiao H (2015) Depletion of histone deacetylase 1 inhibits metastatic abilities of gastric cancer cells by regulating the miR-34a/CD44 pathway. Oncol Rep 34(2):663–672. doi:10.3892/or.2015.4010
Regel I, Merkl L, Friedrich T, Burgermeister E, Zimmermann W, Einwachter H, Herrmann K, Langer R, Rocken C, Hofheinz R, Schmid R, Ebert MP (2012) Pan-histone deacetylase inhibitor panobinostat sensitizes gastric cancer cells to anthracyclines via induction of CITED2. Gastroenterology 143(1):99–109. doi:10.1053/j.gastro.2012.03.035 (e110)
Song S, Wang Y, Xu P, Yang R, Ma Z, Liang S, Zhang G (2015) The inhibition of histone deacetylase 8 suppresses proliferation and inhibits apoptosis in gastric adenocarcinoma. Int J Oncol. doi:10.3892/ijo.2015.3182
Yoon SN, Roh SA, Cho DH, Kim MB, Hyun YL, Ro S, Kim BS, Kim SY, Kim YS, Kim JC (2010) In vitro chemosensitivity of gastric adenocarcinomas to histone deacetylase inhibitors, compared to established drugs. Hepatogastroenterology 57(99–100):657–662
Zhang X, Yashiro M, Ren J, Hirakawa K (2006) Histone deacetylase inhibitor, trichostatin A, increases the chemosensitivity of anticancer drugs in gastric cancer cell lines. Oncol Rep 16(3):563–568
Li Y, Chen K, Zhou Y, Xiao Y, Deng M, Jiang Z, Ye W, Wang X, Wei X, Li J, Liang J, Zheng Z, Yao Y, Wang W, Li P, Xu B (2015) A new strategy to target acute myeloid leukemia stem and progenitor cells using chidamide, a histone deacetylase inhibitor. Curr Cancer Drug Targets 15(6):493–503
Garcia-Manero G, Yang H, Bueso-Ramos C, Ferrajoli A, Cortes J, Wierda WG, Faderl S, Koller C, Morris G, Rosner G, Loboda A, Fantin VR, Randolph SS, Hardwick JS, Reilly JF, Chen C, Ricker JL, Secrist JP, Richon VM, Frankel SR, Kantarjian HM (2008) Phase 1 study of the histone deacetylase inhibitor vorinostat (suberoylanilide hydroxamic acid [SAHA]) in patients with advanced leukemias and myelodysplastic syndromes. Blood 111(3):1060–1066. doi:10.1182/blood-2007-06-098061
Marks PA, Breslow R (2007) Dimethyl sulfoxide to vorinostat: development of this histone deacetylase inhibitor as an anticancer drug. Nat Biotechnol 25(1):84–90. doi:10.1038/nbt1272
McCarthy N (2013) Epigenetics: showing a more sensitive side. Nat Rev Cancer 13(10):680. doi:10.1038/nrc3605
Clozel T, Yang S, Elstrom RL, Tam W, Martin P, Kormaksson M, Banerjee S, Vasanthakumar A, Culjkovic B, Scott DW, Wyman S, Leser M, Shaknovich R, Chadburn A, Tabbo F, Godley LA, Gascoyne RD, Borden KL, Inghirami G, Leonard JP, Melnick A, Cerchietti L (2013) Mechanism-based epigenetic chemosensitization therapy of diffuse large B-cell lymphoma. Cancer Discov 3(9):1002–1019. doi:10.1158/2159-8290.CD-13-0117
Acknowledgments
Our research is supported by the National Science Foundation of China (No. 81272649, No. 81430072, No. 81272203).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Tie, J., Zhang, X. & Fan, D. Epigenetic roles in the malignant transformation of gastric mucosal cells. Cell. Mol. Life Sci. 73, 4599–4610 (2016). https://doi.org/10.1007/s00018-016-2308-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-016-2308-9