Abstract
During meiosis, telomeres cluster and promote homologous chromosome pairing. Telomere clustering depends on conserved SUN and KASH domain nuclear membrane proteins, which form a complex called the linker of nucleoskeleton and cytoskeleton (LINC) and connect telomeres with the cytoskeleton. It has been thought that LINC-mediated cytoskeletal forces induce telomere clustering. However, how cytoskeletal forces induce telomere clustering is not fully understood. Recent study of fission yeast has shown that the LINC complex forms the microtubule-organizing center (MTOC) at the telomere, which has been designated as the “telocentrosome”, and that microtubule motors gather telomeres via telocentrosome-nucleated microtubules. This MTOC-dependent telomere clustering might be conserved in other eukaryotes. Furthermore, the MTOC-dependent clustering mechanism appears to function in various other biological events. This review presents an overview of the current understanding of the mechanism of meiotic telomere clustering and discusses the universality of the MTOC-dependent clustering mechanism.





Similar content being viewed by others
Abbreviations
- DHC:
-
Dynein heavy chain
- DLC:
-
Dynein light chain
- LINC:
-
Linker of nucleoskeleton and cytoskeleton
- MTOC:
-
Microtubule-organizing center
- PC:
-
Pairing center
- SPB:
-
Spindle pole body
- γ-TuC:
-
γ-Tubulin complex
References
Scherthan H (2001) A bouquet makes ends meet. Nat Rev Mol Cell Biol 2(8):621–627
Zickler D, Kleckner N (1998) The leptotene-zygotene transition of meiosis. Annu Rev Genet 32:619–697. doi:10.1146/annurev.genet.32.1.619
Fridkin A, Penkner A, Jantsch V, Gruenbaum Y (2009) SUN-domain and KASH-domain proteins during development, meiosis and disease. Cellular Mol Life Sci: CMLS 66(9):1518–1533. doi:10.1007/s00018-008-8713-y
Hiraoka Y, Dernburg AF (2009) The SUN rises on meiotic chromosome dynamics. Dev Cell 17(5):598–605. doi:10.1016/j.devcel.2009.10.014
Razafsky D, Hodzic D (2009) Bringing KASH under the SUN: the many faces of nucleo-cytoskeletal connections. J Cell Biol 186(4):461–472. doi:10.1083/jcb.200906068
Crisp M, Liu Q, Roux K, Rattner JB, Shanahan C, Burke B, Stahl PD, Hodzic D (2006) Coupling of the nucleus and cytoplasm: role of the LINC complex. J Cell Biol 172(1):41–53. doi:10.1083/jcb.200509124
Yoshida M, Katsuyama S, Tateho K, Nakamura H, Miyoshi J, Ohba T, Matsuhara H, Miki F, Okazaki K, Haraguchi T, Niwa O, Hiraoka Y, Yamamoto A (2013) Microtubule-organizing center formation at telomeres induces meiotic telomere clustering. J Cell Biol 200(4):385–395. doi:10.1083/jcb.201207168
Yamamoto A, Hiraoka Y (2001) How do meiotic chromosomes meet their homologous partners? Lessons from fission yeast. Bioessays 23(6):526–533. doi:10.1002/bies.1072
Harper L, Golubovskaya I, Cande WZ (2004) A bouquet of chromosomes. J Cell Sci 117(Pt 18):4025–4032. doi:10.1242/jcs.01363
Koszul R, Kleckner N (2009) Dynamic chromosome movements during meiosis: a way to eliminate unwanted connections? Trends Cell Biol 19(12):716–724. doi:10.1016/j.tcb.2009.09.007
Yamamoto M, Imai Y, Watanabe Y (1997) Mating and sporulation in Schizosaccharomyces pombe. In: Pringle JR, Broach JR, Jones EW (eds) The molecular and cellular biology of the yeast Saccharomyces; cell cycle and cell biology, vol 3. Cold Spring Harbor Laboratory Press, New York, pp 1037–1106
Chikashige Y, Ding DQ, Funabiki H, Haraguchi T, Mashiko S, Yanagida M, Hiraoka Y (1994) Telomere-led premeiotic chromosome movement in fission yeast. Science 264(5156):270–273
Ding DQ, Chikashige Y, Haraguchi T, Hiraoka Y (1998) Oscillatory nuclear movement in fission yeast meiotic prophase is driven by astral microtubules, as revealed by continuous observation of chromosomes and microtubules in living cells. J Cell Sci 111(Pt 6):701–712
Yamamoto A, West RR, McIntosh JR, Hiraoka Y (1999) A cytoplasmic dynein heavy chain is required for oscillatory nuclear movement of meiotic prophase and efficient meiotic recombination in fission yeast. J Cell Biol 145(6):1233–1249
Yamamoto A, Tsutsumi C, Kojima H, Oiwa K, Hiraoka Y (2001) Dynamic behavior of microtubules during dynein-dependent nuclear migrations of meiotic prophase in fission yeast. Mol Biol Cell 12(12):3933–3946
Yamamoto A, Hiraoka Y (2003) Cytoplasmic dynein in fungi: insights from nuclear migration. J Cell Sci 116(Pt 22):4501–4512. doi:10.1242/jcs.00835
Ananthanarayanan V, Schattat M, Vogel SK, Krull A, Pavin N, Tolic-Norrelykke IM (2013) Dynein motion switches from diffusive to directed upon cortical anchoring. Cell 153(7):1526–1536. doi:10.1016/j.cell.2013.05.020
Vogel SK, Pavin N, Maghelli N, Julicher F, Tolic-Norrelykke IM (2009) Self-organization of dynein motors generates meiotic nuclear oscillations. PLoS Biol 7(4):e1000087. doi:10.1371/journal.pbio.1000087
Ding DQ, Yamamoto A, Haraguchi T, Hiraoka Y (2004) Dynamics of homologous chromosome pairing during meiotic prophase in fission yeast. Dev Cell 6(3):329–341
Miki F, Okazaki K, Shimanuki M, Yamamoto A, Hiraoka Y, Niwa O (2002) The 14-kDa dynein light chain-family protein Dlc1 is required for regular oscillatory nuclear movement and efficient recombination during meiotic prophase in fission yeast. Mol Biol Cell 13(3):930–946. doi:10.1091/mbc.01-11-0543
Shimanuki M, Miki F, Ding DQ, Chikashige Y, Hiraoka Y, Horio T, Niwa O (1997) A novel fission yeast gene, kms1+, is required for the formation of meiotic prophase-specific nuclear architecture. Mol Gen Genet: MGG 254(3):238–249
Nimmo ER, Pidoux AL, Perry PE, Allshire RC (1998) Defective meiosis in telomere-silencing mutants of Schizosaccharomyces pombe. Nature 392(6678):825–828. doi:10.1038/33941
Cooper JP, Watanabe Y, Nurse P (1998) Fission yeast Taz1 protein is required for meiotic telomere clustering and recombination. Nature 392(6678):828–831. doi:10.1038/33947
Tuzon CT, Borgstrom B, Weilguny D, Egel R, Cooper JP, Nielsen O (2004) The fission yeast heterochromatin protein Rik1 is required for telomere clustering during meiosis. J Cell Biol 165(6):759–765. doi:10.1083/jcb.200312061
Chikashige Y, Hiraoka Y (2001) Telomere binding of the Rap1 protein is required for meiosis in fission yeast. Curr Biol 11(20):1618–1623
Kanoh J, Ishikawa F (2001) spRap1 and spRif1, recruited to telomeres by Taz1, are essential for telomere function in fission yeast. Curr Biol 11(20):1624–1630
Niwa O, Shimanuki M, Miki F (2000) Telomere-led bouquet formation facilitates homologous chromosome pairing and restricts ectopic interaction in fission yeast meiosis. EMBO J 19(14):3831–3840. doi:10.1093/emboj/19.14.3831
Trelles-Sticken E, Dresser ME, Scherthan H (2000) Meiotic telomere protein Ndj1p is required for meiosis-specific telomere distribution, bouquet formation and efficient homologue pairing. J Cell Biol 151(1):95–106
Trelles-Sticken E, Adelfalk C, Loidl J, Scherthan H (2005) Meiotic telomere clustering requires actin for its formation and cohesin for its resolution. J Cell Biol 170(2):213–223
Conrad MN, Lee CY, Chao G, Shinohara M, Kosaka H, Shinohara A, Conchello JA, Dresser ME (2008) Rapid telomere movement in meiotic prophase is promoted by NDJ1, MPS3, and CSM4 and is modulated by recombination. Cell 133(7):1175–1187
Koszul R, Kim KP, Prentiss M, Kleckner N, Kameoka S (2008) Meiotic chromosomes move by linkage to dynamic actin cables with transduction of force through the nuclear envelope. Cell 133(7):1188–1201
Scherthan H, Wang H, Adelfalk C, White EJ, Cowan C, Cande WZ, Kaback DB (2007) Chromosome mobility during meiotic prophase in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 104(43):16934–16939
Conrad MN, Dominguez AM, Dresser ME (1997) Ndj1p, a meiotic telomere protein required for normal chromosome synapsis and segregation in yeast. Science 276(5316):1252–1255
Chua PR, Roeder GS (1997) Tam1, a telomere-associated meiotic protein, functions in chromosome synapsis and crossover interference. Genes Dev 11(14):1786–1800
Rockmill B, Roeder GS (1998) Telomere-mediated chromosome pairing during meiosis in budding yeast. Genes Dev 12(16):2574–2586
Goldman AS, Lichten M (2000) Restriction of ectopic recombination by interhomolog interactions during Saccharomyces cerevisiae meiosis. Proc Natl Acad Sci USA 97(17):9537–9542
Schlecht HB, Lichten M, Goldman AS (2004) Compartmentalization of the yeast meiotic nucleus revealed by analysis of ectopic recombination. Genetics 168(3):1189–1203. doi:10.1534/genetics.104.029157
MacQueen AJ, Phillips CM, Bhalla N, Weiser P, Villeneuve AM, Dernburg AF (2005) Chromosome sites play dual roles to establish homologous synapsis during meiosis in C. elegans. Cell 123(6):1037–1050
Phillips CM, Wong C, Bhalla N, Carlton PM, Weiser P, Meneely PM, Dernburg AF (2005) HIM-8 binds to the X chromosome pairing center and mediates chromosome-specific meiotic synapsis. Cell 123(6):1051–1063
Penkner AM, Fridkin A, Gloggnitzer J, Baudrimont A, Machacek T, Woglar A, Csaszar E, Pasierbek P, Ammerer G, Gruenbaum Y, Jantsch V (2009) Meiotic chromosome homology search involves modifications of the nuclear envelope protein Matefin/SUN-1. Cell 139(5):920–933. doi:10.1016/j.cell.2009.10.045
Baudrimont A, Penkner A, Woglar A, Machacek T, Wegrostek C, Gloggnitzer J, Fridkin A, Klein F, Gruenbaum Y, Pasierbek P, Jantsch V (2010) Leptotene/zygotene chromosome movement via the SUN/KASH protein bridge in Caenorhabditis elegans. PLoS Genet 6(11):e1001219. doi:10.1371/journal.pgen.1001219
Wynne DJ, Rog O, Carlton PM, Dernburg AF (2012) Dynein-dependent processive chromosome motions promote homologous pairing in C. elegans meiosis. J Cell Biol 196(1):47–64. doi:10.1083/jcb.201106022
Sato A, Isaac B, Phillips CM, Rillo R, Carlton PM, Wynne DJ, Kasad RA, Dernburg AF (2009) Cytoskeletal forces span the nuclear envelope to coordinate meiotic chromosome pairing and synapsis. Cell 139(5):907–919. doi:10.1016/j.cell.2009.10.039
Scherthan H, Weich S, Schwegler H, Heyting C, Harle M, Cremer T (1996) Centromere and telomere movements during early meiotic prophase of mouse and man are associated with the onset of chromosome pairing. J Cell Biol 134(5):1109–1125
Morimoto A, Shibuya H, Zhu X, Kim J, Ishiguro K, Han M, Watanabe Y (2012) A conserved KASH domain protein associates with telomeres, SUN1, and dynactin during mammalian meiosis. J Cell Biol 198(2):165–172. doi:10.1083/jcb.201204085
Bass HW, Marshall WF, Sedat JW, Agard DA, Cande WZ (1997) Telomeres cluster de novo before the initiation of synapsis: a three-dimensional spatial analysis of telomere positions before and during meiotic prophase. J Cell Biol 137(1):5–18
Carlton PM, Cande WZ (2002) Telomeres act autonomously in maize to organize the meiotic bouquet from a semipolarized chromosome orientation. J Cell Biol 157(2):231–242. doi:10.1083/jcb.200110126
Bass HW, Riera-Lizarazu O, Ananiev EV, Bordoli SJ, Rines HW, Phillips RL, Sedat JW, Agard DA, Cande WZ (2000) Evidence for the coincident initiation of homolog pairing and synapsis during the telomere-clustering (bouquet) stage of meiotic prophase. J Cell Sci 113(Pt 6):1033–1042
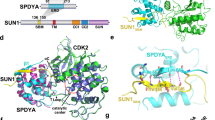
Sosa BA, Rothballer A, Kutay U, Schwartz TU (2012) LINC complexes form by binding of three KASH peptides to domain interfaces of trimeric SUN proteins. Cell 149(5):1035–1047. doi:10.1016/j.cell.2012.03.046
Zhou Z, Du X, Cai Z, Song X, Zhang H, Mizuno T, Suzuki E, Yee MR, Berezov A, Murali R, Wu SL, Karger BL, Greene MI, Wang Q (2012) Structure of Sad1-UNC84 homology (SUN) domain defines features of molecular bridge in nuclear envelope. J Biol Chem 287(8):5317–5326. doi:10.1074/jbc.M111.304543
Starr DA, Han M (2003) ANChors away: an actin based mechanism of nuclear positioning. J Cell Sci 116(Pt 2):211–216
Starr DA, Fischer JA (2005) KASH ‘n Karry: the KASH domain family of cargo-specific cytoskeletal adaptor proteins. Bioessays 27(11):1136–1146. doi:10.1002/bies.20312
Wilhelmsen K, Ketema M, Truong H, Sonnenberg A (2006) KASH-domain proteins in nuclear migration, anchorage and other processes. J Cell Sci 119(Pt 24):5021–5029
Patterson K, Molofsky AB, Robinson C, Acosta S, Cater C, Fischer JA (2004) The functions of Klarsicht and nuclear lamin in developmentally regulated nuclear migrations of photoreceptor cells in the Drosophila eye. Mol Biol Cell 15(2):600–610. doi:10.1091/mbc.E03-06-0374
Haque F, Lloyd DJ, Smallwood DT, Dent CL, Shanahan CM, Fry AM, Trembath RC, Shackleton S (2006) SUN1 interacts with nuclear lamin A and cytoplasmic nesprins to provide a physical connection between the nuclear lamina and the cytoskeleton. Mol Cell Biol 26(10):3738–3751. doi:10.1128/MCB.26.10.3738-3751.2006
Libotte T, Zaim H, Abraham S, Padmakumar VC, Schneider M, Lu W, Munck M, Hutchison C, Wehnert M, Fahrenkrog B, Sauder U, Aebi U, Noegel AA, Karakesisoglou I (2005) Lamin A/C-dependent localization of Nesprin-2, a giant scaffolder at the nuclear envelope. Mol Biol Cell 16(7):3411–3424. doi:10.1091/mbc.E04-11-1009
Mislow JM, Kim MS, Davis DB, McNally EM (2002) Myne-1, a spectrin repeat transmembrane protein of the myocyte inner nuclear membrane, interacts with lamin A/C. J Cell Sci 115(Pt 1):61–70
Zhang Q, Ragnauth CD, Skepper JN, Worth NF, Warren DT, Roberts RG, Weissberg PL, Ellis JA, Shanahan CM (2005) Nesprin-2 is a multi-isomeric protein that binds lamin and emerin at the nuclear envelope and forms a subcellular network in skeletal muscle. J Cell Sci 118(Pt 4):673–687. doi:10.1242/jcs.01642
Lee KK, Starr D, Cohen M, Liu J, Han M, Wilson KL, Gruenbaum Y (2002) Lamin-dependent localization of UNC-84, a protein required for nuclear migration in Caenorhabditis elegans. Mol Biol Cell 13(3):892–901. doi:10.1091/mbc.01-06-0294
Emery AE (1989) Emery-Dreifuss syndrome. J Med Genet 26(10):637–641
Bione S, Maestrini E, Rivella S, Mancini M, Regis S, Romeo G, Toniolo D (1994) Identification of a novel X-linked gene responsible for Emery-Dreifuss muscular dystrophy. Nat Genet 8(4):323–327. doi:10.1038/ng1294-323
Yadlapalli S, Yamashita YM (2013) Chromosome-specific nonrandom sister chromatid segregation during stem-cell division. Nature 498(7453):251–254. doi:10.1038/Nature12106
Yamashita YM (2013) Nonrandom sister chromatid segregation of sex chromosomes in Drosophila male germline stem cells. Chromosome Res 21(3):243–254. doi:10.1007/S10577-013-9353-0
Hagan I, Yanagida M (1995) The product of the spindle formation gene sad1+ associates with the fission yeast spindle pole body and is essential for viability. J Cell Biol 129(4):1033–1047
Chikashige Y, Tsutsumi C, Yamane M, Okamasa K, Haraguchi T, Hiraoka Y (2006) Meiotic proteins Bqt1 and Bqt2 tether telomeres to form the bouquet arrangement of chromosomes. Cell 125(1):59–69
Chikashige Y, Yamane M, Okamasa K, Tsutsumi C, Kojidani T, Sato M, Haraguchi T, Hiraoka Y (2009) Membrane proteins Bqt3 and -4 anchor telomeres to the nuclear envelope to ensure chromosomal bouquet formation. J Cell Biol 187(3):413–427. doi:10.1083/jcb.200902122
Tang X, Jin Y, Cande WZ (2006) Bqt2p is essential for initiating telomere clustering upon pheromone sensing in fission yeast. J Cell Biol 173(6):845–851. doi:10.1083/jcb.200602152
Conrad MN, Lee CY, Wilkerson JL, Dresser ME (2007) MPS3 mediates meiotic bouquet formation in Saccharomyces cerevisiae. Proc Natl Acad Sci USA 104(21):8863–8868
Rao HB, Shinohara M, Shinohara A (2011) Mps3 SUN domain is important for chromosome motion and juxtaposition of homologous chromosomes during meiosis. Genes Cells 16(11):1081–1096. doi:10.1111/j.1365-2443.2011.01554.x
Penkner A, Tang L, Novatchkova M, Ladurner M, Fridkin A, Gruenbaum Y, Schweizer D, Loidl J, Jantsch V (2007) The nuclear envelope protein Matefin/SUN-1 is required for homologous pairing in C. elegans meiosis. Dev Cell 12(6):873–885
Labella S, Woglar A, Jantsch V, Zetka M (2011) Polo kinases establish links between meiotic chromosomes and cytoskeletal forces essential for homolog pairing. Dev Cell 21(5):948–958. doi:10.1016/j.devcel.2011.07.011
Harper NC, Rillo R, Jover-Gil S, Assaf ZJ, Bhalla N, Dernburg AF (2011) Pairing centers recruit a Polo-like kinase to orchestrate meiotic chromosome dynamics in C. elegans. Dev Cell 21(5):934–947. doi:10.1016/j.devcel.2011.09.001
Woglar A, Daryabeigi A, Adamo A, Habacher C, Machacek T, La Volpe A, Jantsch V (2013) Matefin/SUN-1 phosphorylation is part of a surveillance mechanism to coordinate chromosome synapsis and recombination with meiotic progression and chromosome movement. PLoS Genet 9(3):e1003335. doi:10.1371/journal.pgen.1003335
Ding X, Xu R, Yu J, Xu T, Zhuang Y, Han M (2007) SUN1 is required for telomere attachment to nuclear envelope and gametogenesis in mice. Dev Cell 12(6):863–872
Schmitt J, Benavente R, Hodzic D, Hoog C, Stewart CL, Alsheimer M (2007) Transmembrane protein Sun2 is involved in tethering mammalian meiotic telomeres to the nuclear envelope. Proc Natl Acad Sci USA 104(18):7426–7431. doi:10.1073/pnas.0609198104
Horn HF, Kim DI, Wright GD, Wong ES, Stewart CL, Burke B, Roux KJ (2013) A mammalian KASH domain protein coupling meiotic chromosomes to the cytoskeleton. J Cell Biol 202:1023–1039. doi:10.1083/jcb.201304004
Fridolfsson HN, Ly N, Meyerzon M, Starr DA (2010) UNC-83 coordinates kinesin-1 and dynein activities at the nuclear envelope during nuclear migration. Dev Biol 338(2):237–250. doi:10.1016/j.ydbio.2009.12.004
Fridolfsson HN, Starr DA (2010) Kinesin-1 and dynein at the nuclear envelope mediate the bidirectional migrations of nuclei. J Cell Biol 191(1):115–128. doi:10.1083/jcb.201004118
Meyerzon M, Fridolfsson HN, Ly N, McNally FJ, Starr DA (2009) UNC-83 is a nuclear-specific cargo adaptor for kinesin-1-mediated nuclear migration. Development 136(16):2725–2733. doi:10.1242/dev.038596
Whited JL, Cassell A, Brouillette M, Garrity PA (2004) Dynactin is required to maintain nuclear position within postmitotic Drosophila photoreceptor neurons. Development 131(19):4677–4686. doi:10.1242/dev.01366
Wilson MH, Holzbaur EL (2012) Opposing microtubule motors drive robust nuclear dynamics in developing muscle cells. J Cell Sci 125(Pt 17):4158–4169. doi:10.1242/jcs.108688
Cowan CR, Cande WZ (2002) Meiotic telomere clustering is inhibited by colchicine but does not require cytoplasmic microtubules. J Cell Sci 115(Pt 19):3747–3756
Trelles-Sticken E, Loidl J, Scherthan H (2003) Increased ploidy and KAR3 and SIR3 disruption alter the dynamics of meiotic chromosomes and telomeres. J Cell Sci 116(Pt 12):2431–2442. doi:10.1242/jcs.00453
Drummond DR, Cross RA (2000) Dynamics of interphase microtubules in Schizosaccharomyces pombe. Curr Biol 10(13):766–775
Tran PT, Marsh L, Doye V, Inoue S, Chang F (2001) A mechanism for nuclear positioning in fission yeast based on microtubule pushing. J Cell Biol 153(2):397–411
Sagolla MJ, Uzawa S, Cande WZ (2003) Individual microtubule dynamics contribute to the function of mitotic and cytoplasmic arrays in fission yeast. J Cell Sci 116(Pt 24):4891–4903. doi:10.1242/jcs.00796
Horio T, Uzawa S, Jung MK, Oakley BR, Tanaka K, Yanagida M (1991) The fission yeast gamma-tubulin is essential for mitosis and is localized at microtubule organizing centers. J Cell Sci 99(Pt 4):693–700
Funaya C, Samarasinghe S, Pruggnaller S, Ohta M, Connolly Y, Muller J, Murakami H, Grallert A, Yamamoto M, Smith D, Antony C, Tanaka K (2012) Transient structure associated with the spindle pole body directs meiotic microtubule reorganization in S. pombe. Curr Biol 22(7):562–574. doi:10.1016/j.cub.2012.02.042
Sawin KE, Tran PT (2006) Cytoplasmic microtubule organization in fission yeast. Yeast 23(13):1001–1014. doi:10.1002/yea.1404
Sawin KE, Lourenco PC, Snaith HA (2004) Microtubule nucleation at non-spindle pole body microtubule-organizing centers requires fission yeast centrosomin-related protein mod20p. Curr Biol 14(9):763–775. doi:10.1016/j.cub.2004.03.042
Venkatram S, Tasto JJ, Feoktistova A, Jennings JL, Link AJ, Gould KL (2004) Identification and characterization of two novel proteins affecting fission yeast gamma-tubulin complex function. Mol Biol Cell 15(5):2287–2301. doi:10.1091/mbc.E03-10-0728
Tanaka K, Kohda T, Yamashita A, Nonaka N, Yamamoto M (2005) Hrs1p/Mcp6p on the meiotic SPB organizes astral microtubule arrays for oscillatory nuclear movement. Curr Biol 15(16):1479–1486. doi:10.1016/j.cub.2005.07.058
Samejima I, Miller VJ, Rincon SA, Sawin KE (2010) Fission yeast Mto1 regulates diversity of cytoplasmic microtubule organizing centers. Curr Biol 20(21):1959–1965. doi:10.1016/j.cub.2010.10.006
Manandhar G, Schatten H, Sutovsky P (2005) Centrosome reduction during gametogenesis and its significance. Biol Reprod 72(1):2–13. doi:10.1095/biolreprod.104.031245
Schatten H, Sun QY (2009) The functional significance of centrosomes in mammalian meiosis, fertilization, development, nuclear transfer, and stem cell differentiation. Environ Mol Mutagen 50(8):620–636. doi:10.1002/em.20493
Schuh M, Ellenberg J (2007) Self-organization of MTOCs replaces centrosome function during acentrosomal spindle assembly in live mouse oocytes. Cell 130(3):484–498. doi:10.1016/j.cell.2007.06.025
Khodjakov A, Cole RW, Oakley BR, Rieder CL (2000) Centrosome-independent mitotic spindle formation in vertebrates. Curr Biol 10(2):59–67
Heald R, Tournebize R, Blank T, Sandaltzopoulos R, Becker P, Hyman A, Karsenti E (1996) Self-organization of microtubules into bipolar spindles around artificial chromosomes in Xenopus egg extracts. Nature 382(6590):420–425. doi:10.1038/382420a0
Heald R, Tournebize R, Habermann A, Karsenti E, Hyman A (1997) Spindle assembly in Xenopus egg extracts: respective roles of centrosomes and microtubule self-organization. J Cell Biol 138(3):615–628
Jaspersen SL, Giddings TH Jr, Winey M (2002) Mps3p is a novel component of the yeast spindle pole body that interacts with the yeast centrin homologue Cdc31p. J Cell Biol 159(6):945–956. doi:10.1083/jcb.200208169
Tomita K, Cooper JP (2007) The telomere bouquet controls the meiotic spindle. Cell 130(1):113–126. doi:10.1016/j.cell.2007.05.024
Mitchison TJ, Kirschner MW (1985) Properties of the kinetochore in vitro. I. Microtubule nucleation and tubulin binding. J Cell Biol 101(3):755–765
Khodjakov A, Copenagle L, Gordon MB, Compton DA, Kapoor TM (2003) Minus-end capture of preformed kinetochore fibers contributes to spindle morphogenesis. J Cell Biol 160(5):671–683. doi:10.1083/jcb.200208143
Maiato H, Rieder CL, Khodjakov A (2004) Kinetochore-driven formation of kinetochore fibers contributes to spindle assembly during animal mitosis. J Cell Biol 167(5):831–840. doi:10.1083/jcb.200407090
Kitamura E, Tanaka K, Komoto S, Kitamura Y, Antony C, Tanaka TU (2010) Kinetochores generate microtubules with distal plus ends: their roles and limited lifetime in mitosis. Dev Cell 18(2):248–259. doi:10.1016/j.devcel.2009.12.018
Mishra RK, Chakraborty P, Arnaoutov A, Fontoura BM, Dasso M (2010) The Nup107-160 complex and gamma-TuRC regulate microtubule polymerization at kinetochores. Nat Cell Biol 12(2):164–169. doi:10.1038/ncb2016
Torosantucci L, De Luca M, Guarguaglini G, Lavia P, Degrassi F (2008) Localized RanGTP accumulation promotes microtubule nucleation at kinetochores in somatic mammalian cells. Mol Biol Cell 19(5):1873–1882. doi:10.1091/mbc.E07-10-1050
Salina D, Enarson P, Rattner JB, Burke B (2003) Nup358 integrates nuclear envelope breakdown with kinetochore assembly. J Cell Biol 162(6):991–1001. doi:10.1083/jcb.200304080
Joseph J, Liu ST, Jablonski SA, Yen TJ, Dasso M (2004) The RanGAP1-RanBP2 complex is essential for microtubule-kinetochore interactions in vivo. Curr Biol 14(7):611–617. doi:10.1016/j.cub.2004.03.031
Zuccolo M, Alves A, Galy V, Bolhy S, Formstecher E, Racine V, Sibarita JB, Fukagawa T, Shiekhattar R, Yen T, Doye V (2007) The human Nup107-160 nuclear pore subcomplex contributes to proper kinetochore functions. EMBO J 26(7):1853–1864. doi:10.1038/sj.emboj.7601642
Feng J, Huang H, Yen TJ (2006) CENP-F is a novel microtubule-binding protein that is essential for kinetochore attachments and affects the duration of the mitotic checkpoint delay. Chromosoma 115(4):320–329. doi:10.1007/s00412-006-0049-5
Vergnolle MA, Taylor SS (2007) Cenp-F links kinetochores to Ndel1/Nde1/Lis1/dynein microtubule motor complexes. Curr Biol 17(13):1173–1179. doi:10.1016/j.cub.2007.05.077
Moynihan KL, Pooley R, Miller PM, Kaverina I, Bader DM (2009) Murine CENP-F regulates centrosomal microtubule nucleation and interacts with Hook2 at the centrosome. Mol Biol Cell 20(22):4790–4803. doi:10.1091/mbc.E09-07-0560
Chen JS, Lu LX, Ohi MD, Creamer KM, English C, Partridge JF, Ohi R, Gould KL (2011) Cdk1 phosphorylation of the kinetochore protein Nsk1 prevents error-prone chromosome segregation. J Cell Biol 195(4):583–593. doi:10.1083/jcb.201105074
Buttrick GJ, Meadows JC, Lancaster TC, Vanoosthuyse V, Shepperd LA, Hoe KL, Kim DU, Park HO, Hardwick KG, Millar JB (2011) Nsk1 ensures accurate chromosome segregation by promoting association of kinetochores to spindle poles during anaphase B. Mol Biol Cell 22(23):4486–4502. doi:10.1091/mbc.E11-07-0608
Allan VJ, Thompson HM, McNiven MA (2002) Motoring around the Golgi. Nat Cell Biol 4(10):E236–E242. doi:10.1038/ncb1002-e236
Rogalski AA, Singer SJ (1984) Associations of elements of the Golgi apparatus with microtubules. J Cell Biol 99(3):1092–1100
Burkhardt JK, Echeverri CJ, Nilsson T, Vallee RB (1997) Overexpression of the dynamitin (p50) subunit of the dynactin complex disrupts dynein-dependent maintenance of membrane organelle distribution. J Cell Biol 139(2):469–484
Corthesy-Theulaz I, Pauloin A, Pfeffer SR (1992) Cytoplasmic dynein participates in the centrosomal localization of the Golgi complex. J Cell Biol 118(6):1333–1345
Harada A, Takei Y, Kanai Y, Tanaka Y, Nonaka S, Hirokawa N (1998) Golgi vesiculation and lysosome dispersion in cells lacking cytoplasmic dynein. J Cell Biol 141(1):51–59
Fath KR, Trimbur GM, Burgess DR (1994) Molecular motors are differentially distributed on Golgi membranes from polarized epithelial cells. J Cell Biol 126(3):661–675
Efimov A, Kharitonov A, Efimova N, Loncarek J, Miller PM, Andreyeva N, Gleeson P, Galjart N, Maia AR, McLeod IX, Yates JR 3rd, Maiato H, Khodjakov A, Akhmanova A, Kaverina I (2007) Asymmetric CLASP-dependent nucleation of noncentrosomal microtubules at the trans-Golgi network. Dev Cell 12(6):917–930. doi:10.1016/j.devcel.2007.04.002
Chabin-Brion K, Marceiller J, Perez F, Settegrana C, Drechou A, Durand G, Pous C (2001) The Golgi complex is a microtubule-organizing organelle. Mol Biol Cell 12(7):2047–2060
Rios RM, Sanchis A, Tassin AM, Fedriani C, Bornens M (2004) GMAP-210 recruits gamma-tubulin complexes to cis-Golgi membranes and is required for Golgi ribbon formation. Cell 118(3):323–335. doi:10.1016/j.cell.2004.07.012
Verde I, Pahlke G, Salanova M, Zhang G, Wang S, Coletti D, Onuffer J, Jin SL, Conti M (2001) Myomegalin is a novel protein of the Golgi/centrosome that interacts with a cyclic nucleotide phosphodiesterase. J Biol Chem 276(14):11189–11198. doi:10.1074/jbc.M006546200
Ho WC, Allan VJ, van Meer G, Berger EG, Kreis TE (1989) Reclustering of scattered Golgi elements occurs along microtubules. Eur J Cell Biol 48(2):250–263
Reinsch S, Gonczy P (1998) Mechanisms of nuclear positioning. J Cell Sci 111(Pt 16):2283–2295
Salina D, Bodoor K, Eckley DM, Schroer TA, Rattner JB, Burke B (2002) Cytoplasmic dynein as a facilitator of nuclear envelope breakdown. Cell 108(1):97–107
Hebbar S, Mesngon MT, Guillotte AM, Desai B, Ayala R, Smith DS (2008) Lis1 and Ndel1 influence the timing of nuclear envelope breakdown in neural stem cells. J Cell Biol 182(6):1063–1071. doi:10.1083/jcb.200803071
Burke B, Roux KJ (2009) Nuclei take a position: managing nuclear location. Dev Cell 17(5):587–597. doi:10.1016/j.devcel.2009.10.018
Mejat A, Misteli T (2010) LINC complexes in health and disease. Nucleus Austin 1(1):40–52. doi:10.4161/Nucl.1.1.10530
Rothballer A, Schwartz TU, Kutay U (2013) LINCing complex functions at the nuclear envelope: what the molecular architecture of the LINC complex can reveal about its function. Nucleus 4(1):29–36. doi:10.4161/nucl.23387
Tapley EC, Starr DA (2013) Connecting the nucleus to the cytoskeleton by SUN-KASH bridges across the nuclear envelope. Curr Opin Cell Biol 25(1):57–62. doi:10.1016/j.ceb.2012.10.014
Malone CJ, Misner L, Le Bot N, Tsai MC, Campbell JM, Ahringer J, White JG (2003) The C. elegans hook protein, ZYG-12, mediates the essential attachment between the centrosome and nucleus. Cell 115(7):825–836
Kracklauer MP, Banks SM, Xie X, Wu Y, Fischer JA (2007) Drosophila klaroid encodes a SUN domain protein required for Klarsicht localization to the nuclear envelope and nuclear migration in the eye. Fly 1(2):75–85
Anderson MA, Jodoin JN, Lee E, Hales KG, Hays TS, Lee LA (2009) Asunder is a critical regulator of dynein-dynactin localization during Drosophila spermatogenesis. Mol Biol Cell 20(11):2709–2721. doi:10.1091/mbc.E08-12-1165
Sitaram P, Anderson MA, Jodoin JN, Lee E, Lee LA (2012) Regulation of dynein localization and centrosome positioning by Lis-1 and asunder during Drosophila spermatogenesis. Development 139(16):2945–2954. doi:10.1242/dev.077511
Jodoin JN, Shboul M, Sitaram P, Zein-Sabatto H, Reversade B, Lee E, Lee LA (2012) Human Asunder promotes dynein recruitment and centrosomal tethering to the nucleus at mitotic entry. Mol Biol Cell 23(24):4713–4724. doi:10.1091/mbc.E12-07-0558
Zhang X, Lei K, Yuan X, Wu X, Zhuang Y, Xu T, Xu R, Han M (2009) SUN1/2 and Syne/Nesprin-1/2 complexes connect centrosome to the nucleus during neurogenesis and neuronal migration in mice. Neuron 64(2):173–187. doi:10.1016/j.neuron.2009.08.018
Bolhy S, Bouhlel I, Dultz E, Nayak T, Zuccolo M, Gatti X, Vallee R, Ellenberg J, Doye V (2011) A Nup133-dependent NPC-anchored network tethers centrosomes to the nuclear envelope in prophase. J Cell Biol 192(5):855–871. doi:10.1083/jcb.201007118
Splinter D, Tanenbaum ME, Lindqvist A, Jaarsma D, Flotho A, Yu KL, Grigoriev I, Engelsma D, Haasdijk ED, Keijzer N, Demmers J, Fornerod M, Melchior F, Hoogenraad CC, Medema RH, Akhmanova A (2010) Bicaudal D2, dynein, and kinesin-1 associate with nuclear pore complexes and regulate centrosome and nuclear positioning during mitotic entry. PLoS Biol 8(4):e1000350. doi:10.1371/journal.pbio.1000350
Tanaka T, Serneo FF, Higgins C, Gambello MJ, Wynshaw-Boris A, Gleeson JG (2004) Lis1 and doublecortin function with dynein to mediate coupling of the nucleus to the centrosome in neuronal migration. J Cell Biol 165(5):709–721. doi:10.1083/jcb.200309025
Erhardt M, Stoppin-Mellet V, Campagne S, Canaday J, Mutterer J, Fabian T, Sauter M, Muller T, Peter C, Lambert AM, Schmit AC (2002) The plant Spc98p homologue colocalizes with gamma-tubulin at microtubule nucleation sites and is required for microtubule nucleation. J Cell Sci 115(Pt 11):2423–2431
Acknowledgments
I thank Akira Shinohara, Kayoko Tanaka, Takashi Ushimaru, and Masahiro Uritani for critical reading of the manuscript and helpful comments. This work was supported by Grants-in-aid for Scientific Research (C) to A. Y. and the Cooperative Research Program of the Institute for Protein Research, Osaka University.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yamamoto, A. Gathering up meiotic telomeres: a novel function of the microtubule-organizing center. Cell. Mol. Life Sci. 71, 2119–2134 (2014). https://doi.org/10.1007/s00018-013-1548-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-013-1548-1