Abstract
Oxidatively damaged biomolecules impair cellular functions and contribute to the pathology of a variety of diseases. RNA is also attacked by reactive oxygen species, and oxidized RNA is increasingly recognized as an important contributor to neurodegenerative complications in humans. Recently, evidence has accumulated supporting the notion that tRNA is involved in cellular responses to various stress conditions. This review focuses on the intriguing consequences of oxidative modification of tRNA at the structural and functional level.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
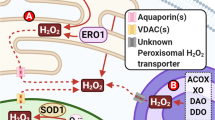
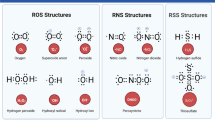
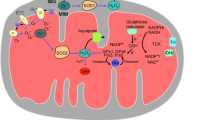
The evolution of higher life forms is firmly associated with the utilization of oxygen, which is essential for energy production in the mitochondrial respiratory chain. Oxidative modifications of biomolecules were adopted to function as signals in intracellular communication. However, the high reactivity of oxygen must be tightly controlled to avoid unwanted oxidative damage. During mitochondrial energy production, oxygen is partially converted into reactive oxygen species (ROS). About 1–2% of oxygen leaks out of the respiratory chain in the form of superoxide (O •−2 ), which is transformed into hydrogen peroxide (H2O2). In the presence of Fenton-reactive metals, such as the ferrous cation (Fe2+), H2O2 is split into a highly aggressive hydroxyl radical (OH•) and a hydroxyl anion (OH−). The majority of ROS arise as byproducts of cellular energy production, whereas a smaller portion is directly created by oxygenases. The intracellular level of ROS is tightly regulated by anti-oxidative enzymes and nonenzymatic antioxidants. ROS act as second messengers and are essential components of signal transduction pathways [1]. However, when the generation of ROS exceeds their degradation by antioxidant enzymes, a condition of oxidative stress arises. Overly elevated levels of ROS cause oxidative damage to proteins, nucleic acids, polysaccharides, and lipids, leading to disturbances of cellular functions and eventually cell death [2].
Oxidative damage of biomolecules is increasingly understood to serve a critical role in the pathogenesis of human diseases, especially those involving neurodegeneration [3]. Neurons are rich in polyunsaturated lipids, which are highly susceptible to oxidative stress damage. Also, oxidative damage of nucleic acids was identified as a key contributor to Alzheimer’s disease; the amount of 8-hydroxy-2′-deoxyguanosine (8-oxo-dG), one of the major products of nucleic acid oxidation, is increased in the brains and cerebrospinal fluid of Alzheimer’s disease patients. Not only DNA but also up to 50% of messenger RNAs (mRNAs) are oxidatively damaged in the affected brain areas of Alzheimer’s patients [4]. DNA damage can result in a loss of genetic information, constituting a more persistent insult than any injury of proteins and RNA molecules that are quickly resynthesized. Most mRNA molecules are relatively short-lived, so RNA oxidation reflects a kind of steady-state balance of oxidative damage [5]. Oxidized mRNAs are not translated properly. Consequently, protein levels are reduced and normal protein function is lost [6]. Protein aggregation, as it is observed in neurodegenerative diseases, may be a consequence. Oxidative damage of mRNA is thought to contribute not only to neuronal cell death in Alzheimer’s disease but also to the pathology of Parkinson’s disease, epilepsy, atherosclerosis, and amyotrophic lateral sclerosis (characterized by the progressive degeneration of motor neurons) [6].
In addition to mRNA, ribosomal RNAs (rRNAs) and transfer RNAs (tRNAs) are also targeted by ROS. The binding capacity to Fenton-reactive ions is higher for rRNA than for tRNA, thus rRNA was found to be more heavily oxidized than tRNA [7]. Although repair mechanisms have been described for RNA modified by alkylation [8], no salvage activity was reported for oxidatively damaged RNA which therefore undergoes enzymatic nucleolysis [9]. How the damaged RNA is distinguished and marked for degradation is largely unknown. Several RNA-binding proteins bind 8-oxo-G with higher affinity than to non-oxidized nucleotides, which could mark oxidized RNA for degradation [10], including the human polynucleotide phosphorylase (Pnp) and the Y box-binding protein-1 (YB-1) [11]. In comparison to mRNA, tRNAs are less susceptible to degradation as a result of stabilization by their tertiary structure and a high content of base modifications. In bacteria, a kind of quality-control process detects mutant tRNAs at the level of tRNA precursors and promotes their degradation [12]. Whether and how an oxidative damage of mature tRNA is recognized and translated into a degradation signal is currently unknown. However, during recent years, evidence has accumulated showing that oxidized tRNA plays a role in stress response regulation.
Stress induces tRNA cleavage
A detailed analysis of the cellular small RNA content by deep sequencing revealed an unexpectedly high abundance of tRNA fragments [13]. The length of tRNA fragments was either 22 nucleotides, which is the typical size of microRNAs (miRNAs), or in the range of 30–40 nucleotides, which corresponds to tRNA halves. miRNAs compose a class of highly expressed, noncoding small RNAs involved in post-transcriptional regulation of mRNA abundance. miRNAs bind to the 3’-untranslated regions of target mRNAs via imperfect base-pairing to inhibit their translation. The second abundant class of RNA fragments was found to be generated under various stress conditions by single cleavage of tRNA molecules in the anticodon loop [14]. Several findings support the assumption that such instances of tRNA cleavage occur as specific events, rather than by degradation in the course of metabolic turnover: (1) tRNA cleavage is triggered by certain stress conditions, including nutritional deficiency, heat shock, hypothermia, hypoxia, and oxidative stress (Fig. 1), but not by others, such as irradiation. Similarly, some apoptotic inducers (which actually also cause stress), such as Fas ligand, promote tRNA cleavage, while others, such as staurosporine, are inactive [15]. (2) Some tRNA species are more vulnerable than others, e.g., nutritional stress-induced cleavage of tRNA is specific for methionine (tRNAMet) and tRNAVal, but not for tRNATyr [14]. Similarly, oxidative stress mediated by H2O2 results in strong cleavage of tRNAArg and some cleavage of tRNATrp, but not of tRNATyr [15]. The data published so far are incomplete; a comprehensive investigation of all tRNA species exposed to various types of stress is still lacking. (3) The specificity of this process is further emphasized by the fact that tRNA cleavage is a conserved response to oxidative stress among many eukaryotic species, from Saccharomyces cerevisiae and Arabidopsis to human cells [15].
tRNAs are cleaved by specific enzymes
Stress-induced tRNA cleavage is carried out by specific enzymes (Fig. 2). In yeast, tRNAs are cleaved by the Rny1 RNase [16]. In mammalian cells, angiogenin, a secreted ribonuclease, is required for stress-induced endonucleolytic cleavage of tRNAs [14]. Both RNases are normally spatially segregated from cytoplasmic tRNAs, but they are released into the cytoplasm under conditions of oxidative stress [16]. Angiogenin was originally recognized as an angiogenic factor secreted by tumor cells into the surrounding medium exhibiting enhanced secretion under hypoxic conditions. In vivo, receptors on the surface of endothelial cells bind and internalize angiogenin. Importantly, the induction of new blood vessel outgrowth is dependent on its ribonuclease activity. In endothelial cells, angiogenin is concentrated in the nucleolus and also partially localized to the cytoplasm, where it is bound by the ribonuclease inhibitor RNH1 [17]. Depletion of RNH1 increases tRNA cleavage consistent with a role for cytoplasmic angiogenin [18].
tRNAs are cleaved by specific enzymes. Bacterial endonucleases, PrrC, colicin D, and colicin E5, cleave specific subsets of bacterial tRNAs of invading species in the anticodon region. A similar tRNA endonuclease activity is mediated by γ-toxin from the dairy yeast Kluyveromyces lactis, to arrest the growth of Saccharomyces cerevisiae. Under oxidative stress, tRNAs are cleaved by Rny1 in yeast and by angiogenin in mammalian cells. The nucleolytic activity of angiogenin is further regulated by RNH1, an inhibitory protein, and by tRNA methylation
Yet another finding supports the view that tRNA cleavage is not just a simple degradation process: the cleavage is controlled by RNA methylation. The DNA methyltransferase Dnmt2 is capable of methylating tRNA molecules as well, in particular tRNAAsp, tRNAVal and tRNAGly. Dnmt2-mediated methylation was shown to protect tRNAs against ribonuclease cleavage by angiogenin [19].
The levels of full-length tRNAs do not decline significantly during stress-induced tRNA cleavage; only a small proportion of tRNA is targeted. Is there any function, then, of the tRNA halves generated in a controlled way as a response to many, but not all, stress conditions?
tRNA halves promote stress granule assembly
When exposed to environmental stress, eukaryotic cells activate stress response programs. Energy-expensive processes, such as transcription and translation, are reduced to conserve energy for survival and the repair of stress-induced damage. The expression of common housekeeping genes is blocked, whereas the expression of genes that repair stress-induced damage and promote cell survival is increased. Cleaved tRNA may contribute to the translational arrest by inhibitory interactions with the translation machinery. After cleavage, tRNA might persist in a fully folded conformation, but the nicked anticodon loop would prevent correct interaction with its respective codon (Fig. 3). Elongation, therefore, would be stalled [20]. In association with specialized proteins, tRNA halves might directly regulate gene expression (see below). In addition to interfering with protein translation, tRNA halves were demonstrated capable of inducing the formation of stress granules (SG) [21]. SG appear as part of the stress response program as cytoplasmic aggregates in which nontranslated transcripts accumulate. It is thought that SG compose the sites where mRNAs are sorted into stored, degraded, or translated ones. SG are physically associated with P-bodies, another kind of cytoplasmic foci present in resting cells that are enriched in RNA-degrading enzymes (Fig. 3). P-bodies serve as a site of mRNA degradation.
Functions of damaged tRNAs. The nicked anticodon loop in cleaved tRNA prevents correct interaction with its respective codon and elongation is stalled. The 5′-half tRNA halves induce the assembly of stress granules, cytoplasmic aggregates which associate with P-bodies to function in selective degradation of mRNA. At least one tRNA-derived fragment was required for the proliferation of prostate cancer cells. In association with specialized proteins, tRNA halves contribute to the regulation of gene expression by guiding endonucleolytic cleavage of target mRNAs
SG formation was demonstrated to be enhanced by angiogenin [21], which provided a traceable link to tRNA cleavage. When cells were transfected with tRNA fragments derived from stress-induced cleavage, transfection of 5′-tRNA halves induced SG formation, while no such effect was observed with the 3′-tRNA fragments. These results convincingly exhibit the signaling component of tRNA-derived fragments in the cellular response to stress.
tRNA fragments in RNA-mediated translational silencing
Post-transcriptional regulation of gene expression includes the miRNA-mediated gene-silencing pathway, in which selected mRNAs are prevented from translation. One of the major silencing routes employs the RNA-induced silencing complex (RISC). Non-coding RNA transcripts (pre-miRNAs) are processed by Drosha and Dicer nucleases and loaded onto Argonaute (AGO) proteins, which are components of the RISC. RISC-associated miRNAs recognize and interact with complementary sequences of target mRNAs, which are then silenced by degradation or stalled translation. Also, tRNA fragments were found to be associated with AGO proteins (Fig. 3) [22]. In HeLa cells, tRNA-derived ~19 nucleotide-long fragments, mainly processed from the 5′-end, are highly abundant [23]. In murine embryonic stem cells, a small RNA fragment was identified that mapped to a tRNAIle gene. For this tRNAIle, an alternative fold was predicted beside the tRNA cloverleaf structure, which could serve as a substrate for the Dicer RNA processing enzyme [24]. tRNA-derived small fragments with low inherent silencing activity might compete with other small RNA species for association with AGO, and due to their high abundance, significantly diminish the silencing activity of ‘regular’ miRNAs [25]. Some small RNA fragments cleaved from tRNA in prostate cancer cell lines were further characterized [26]. They were 18–25-nucleotides long, representing 5′-ends or 3′-ends of mature tRNAs or tRNA precursors. At least one of them, derived from the 3′-end of a tRNASer precursor transcript, was shown to be necessary for proliferation of these cancer cells (Fig. 3). This fragment was cleaved from a pre-tRNA in the cytoplasm by RNase Z (ELAC2), an enzyme involved in tRNA maturation [27]. During this maturation process, RNase Z removes a 3′-trailer from pre-tRNA.
RNase Z not only trims tRNA precursors but is also capable of cleaving target RNA at any desired site by a silencing mechanism similar to that exerted by the RISC [28]. For that silencing activity, RNase Z has to be loaded with a guide RNA that brings the enzyme into contact with the target RNA. By co-immunoprecipitation, small RNA species associated with RNase Z were identified in human cell lines. Among those small RNAs was the 5′-half of tRNAGlu, which indeed functioned as guide RNA in RNase Z-mediated cleavage (Fig. 3). A mRNA targeted by the tRNAGlu-half/RNase Z complex was identified as coding for PPM1F, a serine/threonine protein phosphatase which specifically dephosphorylates Thr-286 of the calcium/calmodulin-dependent protein kinase II (CaMKII) [29]. CaMKII is a multifunctional enzyme that is involved in inducing apoptosis upon endoplasmatic reticulum stress [30]. Downregulation of PPM1F, as a direct consequence of tRNA cleavage, could therefore promote apoptosis in cells under stress. How gene silencing with small tRNA-derived fragments might be associated with stress is a fascinating question that warrants further research.
Cleavage of tRNA in the anticodon loop by bacterial and fungal endonucleases
Halves of tRNA molecules were also found in bacterial species, in which tRNA anticodon damage is an activity of a kind of innate immune system against invading species. The bacterial tRNA endonucleases, PrrC, colicin D, and colicin E5, cleave specific subsets of bacterial tRNAs in the anticodon region (Fig. 2) [31–33]. Notably, the presence of modified nucleosides within the anticodon loop seems to determine which tRNAs are targeted. PrrC cleaves tRNALys at the 5′ phosphate of the 5-methylaminomethyl-2-thiouridine (mnm5S2U) wobble base in position 34 [31, 34, 35]. Colicin D shows specificity for all four tRNAArg isoaccepting molecules, and cleaving them at nucleotide 38 impairs protein synthesis and promotes cell death [33]. tRNAs for Tyr, His, Asn, and Asp, which contain the modified base queuine at the wobble position, are cut by colicin E5 at the 3′ phosphate of the wobble nucleotide. A similar tRNA endonuclease activity mediated by γ-toxin was found in the dairy yeast Kluyveromyces lactis, which arrests the growth of Saccharomyces cerevisiae. The γ-toxin is secreted as a component of a larger protein complex and transported into the cytoplasm of target cells, where it cleaves three tRNA species specific for Glu, Gln and Lys at the 3′ site of the modified wobble uridine (mcm5S2U, Fig. 2) [36]. Modifications of the ribose in the wobble base suppressed the cleavage [37]. In contrast to the stress signaling function of tRNA halves described above, no such function was described for the cleavage products of bacterial and fungal tRNA endonucleases. Instead, tRNA cleavage in bacteria and yeast results in inhibition of protein translation and induction of cell death.
Oxidative dethiolation of tRNAs
In addition to tRNA cleavage under stress conditions, single nucleotides in tRNAs are chemically altered by high levels of ROS. tRNA contains a variety of modified nucleotides, some of which serve as targets for oxidative modifications. The oxidative dethiolation of sulfur-containing nucleotides is one example that has been investigated in detail.
Sulfur-containing tRNAs
There are more than 100 post-transcriptionally modified nucleosides present in all types of RNA, and most of them are located in tRNA. They function in modifying the translation process by precise decoding of the genetic information [38–42]. The tRNA Modification Databases [43, 44] contain a comprehensive list of modified nucleosides identified to date in tRNA. Among them, 18 are modified pyrimidine nucleosides containing either sulfur or selenium at the nucleobase moiety. The structures of known thio- and seleno-pyrimidine nucleosides are shown in Fig. 4. Most of them are 2-thiouridines, which are substituted with a side chain at the carbon C5, in addition to a sulfur atom at C2. The 2-thiouridines most typically appear in the first position of the tRNA anticodon (position 34, wobble position). Three tRNAs, specific for lysine (tRNALys3), glutamic acid (tRNAGlu) and glutamine (tRNAGln), contain hypermodified 2-thiouridines in the wobble position. The 2-thiouridine (S2U) unit in position 34 facilitates predominantly Watson–Crick base pairing with adenosine and restricts the wobble pairing with G in the third position of the codon [39, 45]. The S2U modification in the anticodon loop of tRNA is also a recognition element for the cognate aminoacyl tRNA synthetases (aa-RS) [46–48]. Notably, the human tRNALys3 serves as a primer for the reverse transcription of the human immunodeficiency virus type 1 (HIV-1) RNA; it contains 5-methoxycarbonylmethyl-2-thiouridine (mcm5S2U) in position 34, which is required for the formation of the initial complex with the viral RNA [49, 50]. In addition to S2U, selenium-containing uridine (Se2U) has been identified in tRNAs from bacterial, mammalian and plant species, and this modified unit is located at the wobble position of three tRNAs specific for Glu, Gln and Lys (the same as for S2U) [51–56]. All these modified units contain thiocarbonyl/selenocarbonyl function, which is prone to attack of oxidizing agents [57].
The structures of sulfur- and selenium-containing pyrimidine nucleosides. S2U and Se2U nucleosides present in transfer RNAs, according to Agris et al., http://rna-mdb.cas.albany.edu/RNAmods/ [43] and Dunin-Horkawicz et al., http://modomics.genesilico.pl/ [44]
Loss of function of sulfur-containing tRNAs in oxidative conditions in vitro
The earliest report on oxidative damage of tRNA describes the inactivation of specific E. coli or rabbit liver tRNA species for amino acid acceptor activity via treatment with dilute iodine–potassium iodide and recovery of functional tRNAs via the action of appropriate reducing agents [58]. Chemical modification of sulfur-containing pyrimidine nucleosides in E. coli tRNA with various oxidizing reagents showed that, in the presence of hydrogen peroxide or cyanogen bromide (CNBr), the entire 5-methylaminomethyl-2-thiouridine nucleoside was dethiolated, although a detailed structure of products was not reported [59]. The CNBr-oxidized tRNA species specific for Glu, Gln and Lys lost their potential to be aminoacylated by their cognate aa-RSs. It has been shown (either at the nucleoside or entire tRNA level) that the 2-thiouridines are oxidized by H2O2 to uridine and to another “H2O2-altered”, unidentified product, but not to a ring-opened or disulfide structure [60]. Dethiolation of E. coli tRNAGlu in oxidative conditions led to more than 95% conversion of the 2-thiouridine 34 to uridine, as assigned by base composition analysis [61]. Notably, the H2O2-oxidized tRNAGlu lost much of its binding affinity to tRNAPhe, which has the complementary anticodon sequence. The stability of the tRNAGlu/tRNAPhe complex after mnm5S2U oxidation was significantly reduced, as determined by the temperature jump relaxation method. The melting temperature of this complex dropped by 20°C in comparison to the Tm of the parental tRNAGlu/tRNAPhe complex. The rationale for the observed phenomena was the decrease of the stacking interaction and change in uridine conformational flexibility, in comparison to the S2U nucleoside. Other experiments have shown that probing the tRNALys3 (mcm5S2U at the wobble position) with potassium peroxynitrite (ONOOK), which generates hydroxyl radicals in the absence of heavy metal ions, results in dethiolation and subsequent strand scission at the wobble position, but the non-thiolated substrate is not cleaved [62]. Hydroxyl radicals are thought to promote cleavage of the polynucleotide backbone. Despite these intense studies, only very limited information has been made available on the structure of the dethiolated nucleosides and the origin of the observed loss of tRNA function.
Dethiolation of S2U on nucleoside level
The removal of the sulfur atom from 2-thiopyrimidine nucleosides proceeds either under reductive conditions or under various oxidative conditions in a process called oxidative desulfuration (desulfurization or dethiolation). Reduction of S2U gives the corresponding 4-pyrimidinone nucleoside (H2U). The action of oxidants may remove sulfur in favor of hydrogen (desulfuration to H2U) or in favor of oxygen (oxidation to U) (Fig. 5), and these reactions usually occur simultaneously with a ratio dependent on the kind of oxygen agent and reaction conditions [57, 63].
Dethiolation of S2U under reductive conditions
It has been reported that dethiolation of 2-thiopyrimidine nucleosides proceeds in moderate yield under reductive conditions in the reaction with dibenzoyldiazene [64] or under treatment with dipotassium diazenedicarboxylate [65]. However, desulfuration of 2-thiothymidine by Raney-nickel reduction worked more efficiently to produce corresponding 4-pyrimidinone derivatives at an isolated yield of 61% [66].
Dethiolation of S2U under oxidative conditions
Oxidative dethiolation of the 2-thiopyrimidine moiety has been observed upon treatment with various oxidizing agents: aqueous iodine [67], m-chloroperbenzoic acid [67], dimethyldioxirane [68] and trans-2-phenylsulfonyl-3-phenyloxaziridine (PSO) [69]. The reactivity of the 2-thiocarbonyl function towards aqueous iodine was found to be the main cause of the low efficiency of 2-thionucleoside incorporation into the oligonucleotide chain during the standard phosphoramidite oligonucleotide synthesis, as desulfuration occurred during the oxidation step routinely performed with this oxidizing agent [67, 70–73]. During the evaluation of alternative oxidizing agents, it was discovered that treatment of 2-thiothymidine with m-chloroperbenzoic acid/pyridine solution rapidly produced the corresponding 4-pyrimidinone 2′-deoxynucleoside in high yield [67]. Recently, the same oxidizing reagent was successfully applied in a RNA nucleotide synthesis for the efficient, selective transformation of 2-thiouridine into 4-pyrimidinone ribonucleoside [74]. Treatment of the sugar-protected 2-thiouridine derivative with dimethyldioxirane revealed incomplete removal of the sulfur atom. Under these conditions, the thiocarbonyl function at the C2 position of the pyrimidine ring was partially dethiolated, leading to the 4-pyrimidinone nucleoside (yield 43%), and the disulfide of 2-thiouridine was formed with 20% yield [68]. We have reported that the S2U nucleoside in the presence of PSO, undergoes selective transformation to the H2U nucleoside. The loss of the sulfur atom from 2-thiouridine was also observed under H2O2 treatment in aqueous solution. Under these conditions, the 2-thionucleoside is converted to the 4-pyrimidinone nucleoside and uridine [64, 75], and the process is pH-dependent (Sochacka, unpublished results). The development of the synthesis of the corresponding S2U- and H2U-phosphoramidites has provided model oligonucleotides for further studies on tRNA dethiolation.
Dethiolation of S2U on the RNA oligonucleotide level
For the first time, the effective desulfuration of the 2-thiouridine built into the 5′-TdA(S2U)dGdC-3′ pentamer could be achieved by the treatment of oligomer in ‘organic conditions’ with PSO in a water/acetonitrile solution [69]. The obtained product, as identified by MALDI-TOF mass spectrometry, did not contain sulfur and, in accordance with the results obtained at the nucleoside level, was identified as the 4-pyrimidinone nucleoside. Further studies, carried out under in vitro oxidative stress conditions in the presence of aqueous H2O2, demonstrated that the S2U-containing RNA, which was homo-sequential to the anticodon loop of tRNALys3, was transformed predominantly to H2U-RNA [76]. The affinity of H2U-RNA to its Watson-Crick complement (A opposite of H2U), as determined by UV/VIS melting experiments, was much smaller than the affinity of S2U-RNA or U-RNA to the complementary RNA. These novel results indicated that the sulfur-containing nucleosides present in tRNA chains are transformed preferentially to the H2U analog than to uridine when dethiolated under oxidative stress conditions, which is commonly accepted as the sole dethiolation product.
Biological consequences of desulfuration of 2-thiouridine in tRNA
It has been recognized that the 2-thiocarbonyl group of 2-thiouracil nucleosides strongly influences their conformation and plays a key role in the modulation of base pair recognition [77, 78]. These properties are crucial for the decoding ability of natural 2-thiouracil ribonucleosides located in the anticodon wobble position of many tRNAs [38–40]. 2-Thiouridines preferentially adopt a rigid C3′-endo sugar ring conformation [79, 80], for which the long-range electrostatic effect between the 2′-OH group and the sulfur atom plays a dominant role [81]. The S2U-A base pair in RNA duplexes is more stable than that of a parent U-A one [82, 83]. Furthermore, due to steric hindrance and the weaker H-bonding ability of sulfur relative to oxygen, 2-thiouracil ribonucleosides make the S2U-G wobble base pairs less stable than base pairs containing uridine [80, 82–84]. The specific hybridization properties of 2-thiouracil ribonucleosides led to their practical use in the antisense strategy and single-nucleotide polymorphism (SNP) analysis [85, 86, 87, 88].
Our recent studies on 4-pyrimidinone ribonucleoside conformation in a solid state and in solution (NMR) have demonstrated that the conformational characteristics of the ribofuranose ring is dramatically affected by desulfuration [74]. The 4-pyrimidinone nucleoside predominantly adopts the C2′-endo form (S conformer) in aqueous solution (Fig. 6), and this ribose pucker is also fixed for H2U molecules in the crystal state. The loss of 2-thiocarbonyl/2-carbonyl function significantly influences the conformation of nucleosides. The observed differences between the S2U, U and H2U ribose folding (Table 1) may have important biological consequences on tRNA fate and cellular function, because dethiolation of natural 2-thiouridines is plausible under conditions of oxidative stress in the cell.
Notably, 4-pyrimidinone nucleosides are nucleoside analogs lacking both the N3-amide hydrogen and the thiocarbonyl/carbonyl function at the C2 position in the heterobase moiety, characteristic for 2-thiouridines and natural uridines, and thus offer entirely different possibilities for base pairing within RNA and DNA duplexes (Fig. 7). Interestingly, the UV-melting profiles of S2U-, U- and H2U-RNAs hybridized to wobble complements (G opposite of modification site) are almost identical, what indicates that the mode of wobble base pairing is less affected by dethiolation [76]. Plausible modes of hydrogen bond patterns of H2U-A and H2U-G are shown in Fig. 7.
It is clear that shifting guanine residues along the 4-pyrimidinone moiety offers two hydrogen bonds that might compensate for the original wobble hydrogen bonds. This hypothesis should be addressed in future studies.
Moreover, some data reveal that dH2U-containing DNA oligonucleotides undergo strand cleavage in certain conditions [89]. We have also reported that H2U-containing RNA is unstable in basic conditions and decomposes to its abasic form, followed by strand scission [76]. These preliminary results need to be further confirmed to consider the H2U-containing anticodon loop as a strand scission site, whereby such a possibility would offer new insight into tRNA oxidative damage.
Conclusions
Oxidative damage of biomolecules, including RNA, contributes to neurodegenerative and other diseases. tRNA is cleaved under oxidative stress, and the resulting tRNA fragments participate in stress signaling pathways, like the induction of stress granule formation. Modified nucleosides, which are found in wide variety in tRNA, might constitute a primary target for oxidative attack. An example is the transformation of S2U-tRNA to H2U-tRNA which creates a site of tRNA damage. Such damage may disturb codon–anticodon interactions and recognition of tRNA by aa-tRNA synthetases and other enzymes involved in tRNA metabolism and function. Oxidative stress may, in fact, cause loss of function of S2U-tRNAs, disrupting their participation in the production of vital proteins, which might also be crucial for cellular ageing through the accumulation of nonfunctional tRNA species. Another potential implementation of the oxidative desulfuration of 2-thiopyrimidine nucleosides might be as anti-HIV-1 therapeutic strategies. The use of oxidizing agents might block the function of tRNALys3 in priming reverse transcription by preventing its binding to viral RNA. Moreover, some data suggest that H2U-containing RNA undergoes strand cleavage at the modification site. Future research will reveal whether there is any connection between the S2U-to-H2U transition and the enzymatic cleavage of tRNAs with functions in stress response signaling and nonself defense.
References
Brown DI, Griendling KK (2009) Nox proteins in signal transduction. Free Radic Biol Med 47:1239–1253
Avery SV (2011) Molecular targets of oxidative stress. Biochem J 434:201–210
Sayre LM, Perry G, Smith MA (2008) Oxidative stress and neurotoxicity. Chem Res Toxicol 21:172–188
Shan X, Chang Y, Lin CL (2007) Messenger RNA oxidation is an early event preceding cell death and causes reduced protein expression. FASEB J 21:2753–2764
Nunomura A, Perry G, Pappolla MA, Wade R, Hirai K, Chiba S, Smith MA (1999) RNA oxidation is a prominent feature of vulnerable neurons in Alzheimer’s disease. J Neurosci 19:1959–1964
Kong Q, Lin CL (2010) Oxidative damage to RNA: mechanisms, consequences, and diseases. Cell Mol Life Sci 67:1817–1829
Honda K, Smith MA, Zhu X, Baus D, Merrick WC, Tartakoff AM, Hattier T, Harris PL, Siedlak SL, Fujioka H et al (2005) Ribosomal RNA in Alzheimer disease is oxidized by bound redoxactive iron. J Biol Chem 280:20978–20986
Aas PA, Otterlei M, Falnes PO, Vagbo CB, Skorpen F, Akbari M, Sundheim O, Bjoras M, Slupphaug G, Seeberg E, Krokan HE (2003) Human and bacterial oxidative demethylases repair alkylation damage in both RNA and DNA. Nature 421:859–863
Li Z, Wu J, Deleo CJ (2006) RNA damage and surveillance under oxidative stress. IUBMB Life 58:81–588
Hayakawa H, Fujikane A, Ito R, Matsumoto M, Nakayama KI, Sekiguchi M (2010) Human proteins that specifically bind to 8-oxoguanine-containing RNA and their responses to oxidative stress. Biochem Biophys Res Commun 403:220–224
Castellani RJ, Nunomura A, Rolston RK, Moreira PI, Takeda A, Perry G, Smith MA (2008) Sublethal RNA oxidation as a mechanism for neurodegenerative disease. Int J Mol Sci 9:789–806
Deutscher MP (2006) Degradation of RNA in bacteria: comparison of mRNA and stable RNA. Nucl Acids Res 34:659–666
Kawaji H, Nakamura M, Takahashi Y, Sandelin A, Katayama S, Fukuda S, Daub CO, Kai C, Kawai J, Yasuda J et al (2008) Hidden layers of human small RNAs. BMC Genomics 9:157–178
Fu H, Feng J, Liu Q, Sun F, Tie Y, Zhu J, Xing R, Sun Z, Zheng X (2009) Stress induces tRNA cleavage by angiogenin in mammalian cells. FEBS Lett 583:437–442
Thompson DM, Lu C, Green PJ, Parker R (2008) tRNA cleavage is a conserved response to oxidative stress in eukaryotes. RNA 14:2095–2103
Thompson DM, Parker R (2009) The RNase Rny1p cleaves tRNAs and promotes cell death during oxidative stress in Saccharomyces cerevisiae. J Cell Biol 185:43–50
Tsuji T, Sun Y, Kishimoto K, Olson KA, Liu S, Hirukawa S, Hu GF (2005) Angiogenin is translocated to the nucleus of HeLa cells and is involved in ribosomal RNA transcription and cell proliferation. Cancer Res 65:1352–1360
Yamasaki S, Ivanov P, Hu GF, Anderson P (2009) Angiogenin cleaves tRNA and promotes stress-induced translational repression. J Cell Biol 185:35–42
Schaefer M, Pollex T, Hanna K, Tuorto F, Meusburger M, Helm M, Lyko F (2010) RNA methylation by Dnmt2 protects transfer RNAs against stress-induced cleavage. Genes Dev 24:1590–1595
Thompson DM, Parker R (2009) Stressing out over tRNA cleavage. Cell 138:215–219
Emara MM, Ivanov P, Hickman T, Dawra N, Tisdale S, Kedersha N, Hu GF, Anderson P (2010) Angiogenin-induced tRNA-derived stress-induced RNAs promote stress-induced stress granule assembly. J Biol Chem 285:10959–10968
Burroughs AM, Ando Y, Hoon ML, Tomaru Y, Suzuki H, Hayashizaki Y, Daub CO (2011) Deep-sequencing of human Argonaute-associated small RNAs provides insight into miRNA sorting and reveals Argonaute association with RNA fragments of diverse origin. RNA Biol 8:158–177
Cole C, Sobala A, Lu C, Thatcher SR, Bowman A, Brown JW, Green PJ, Barton GJ, Hutvagner G (2009) Filtering of deep sequencing data reveals the existence of abundant dicer-dependent small RNAs derived from tRNAs. RNA 15:2147–2160
Babiarz JE, Ruby JG, Wang Y, Bartel DP, Blelloch R (2008) Mouse ES cells express endogenous shRNAs, siRNAs, and other Microprocessor-independent, Dicer-dependent small RNAs. Genes Dev 22:2773–2785
Haussecker D, Huang Y, Lau A, Parameswaran P, Fire AZ, Kay MA (2010) Human tRNA-derived small RNAs in the global regulation of RNA silencing. RNA 16:673–695
Lee YS, Shibata Y, Malhotra A, Dutta A (2009) A novel class of small RNAs: tRNA-derived RNA fragments (tRFs). Genes Dev 23:2639–2649
Phizicky EM, Hopper AK (2010) tRNA biology charges to the front. Genes Dev 24:1832–1860
Elbarbary RA, Takaku H, Uchiumi N, Tamiya H, Abe M, Takahashi M, Nishida H, Nashimoto M (2009) Modulation of gene expression by human cytosolic tRNase Z(L) through 5′-half-tRNA. PLoS One 4:e5908
Harvey BP, Banga SS, Ozer HL (2004) Regulation of the multifunctional Ca2+/calmodulin-dependent protein kinase II by the PP2C phosphatase PPM1F in fibroblasts. J Biol Chem 279:24889–24898
Timmins JM, Ozcan L, Seimon TA, Li G, Malagelada C, Backs J, Backs T, Bassel-Duby R, Olson EN, Anderson ME, Tabas I (2009) Calcium/calmodulin-dependent protein kinase II links ER stress with Fas and mitochondrial apoptosis pathways. J Clin Invest 119:2925–2941
Amitsur M, Levitz R, Kaufmann G (1987) Bacteriophage T4 anticodon nuclease, polynucleotide kinase and RNA ligase reprocess the host lysine tRNA. EMBO J 6:2499–2503
Ogawa T, Tomita K, Ueda T, Watanabe K, Uozumi T, Masaki H (1999) A cytotoxic ribonuclease targeting specific transfer RNA anticodons. Science 283:2097–2100
Tomita K, Ogawa T, Uozumi T, Watanabe K, Masaki H (2000) A cytotoxic ribonuclease which specifically cleaves four isoaccepting arginine tRNAs at their anticodon loops. Proc Natl Acad Sci USA 97:8278–8283
Jiang Y, Meidler R, Amitsur M, Kaufmann G (2001) Specific interaction between anticodon nuclease and the tRNA(Lys) wobble base. J Mol Biol 305:377–388
Jiang Y, Blanga S, Amitsur M, Meidler R, Krivosheyev E, Sundaram M, Bajji AC, Davis DR, Kaufmann G (2002) Structural features of tRNALys favored by anticodon nuclease as inferred from reactivities of anticodon stem and loop substrate analogs. J Biol Chem 277:3836–3841
Lu J, Huang B, Esberg A, Johansson MJ, Byström AS (2005) The kluyveromyces lactis gamma-toxin targets tRNA anticodons. RNA 11:1648–1654
Keppetipola N, Jain R, Meineke B, Diver M, Shuman S (2009) Structure-activity relationships in Kluyveromyces lactis gamma-toxin, a eukaryal tRNA anticodon nuclease. RNA 15:1036–1044
Agris PF (2004) Decoding the genome: a modified view. Nucl Acids Res 32:223–238
Agris PF, Vendeix FAP, Graham WD (2007) tRNA’s wobble decoding of the genome: 40 years of modification. J Mol Biol 366:1–13
Yokoyama S, Nishimura S (1995) Modified nucleosides and codon recognition. In: Söll D, RajBhandary U (eds) tRNA: structure, biosynthesis and function. ASM Press, Washington DC, pp 207–223
Curran JF (1998) Modified nucleosides in translation. In: Grosjean H, Benne B (eds) Modification and editing of RNA. ASM Press, Washington DC, pp 493–516
Watanabe K (2007) Role of modified nucleosides in the translation function of tRNAs from extreme thermophilic bacteria and animal mitochondria. Bull Chem Soc Jpn 80:1253–1267
Agris PF, Crain PF, Rozenski J, Fabris D, Vendeix FAP. The RNA modification database. http://biochem.ncsu.edu/RNAmods/.
Dunin-Horkawicz S, Czerwoniec A, Gajda MJ, Feder M, Grosjean H, Bujnicki JM (2006) MODOMICS: a database of RNA modification pathways. Nucl Acids Res 34:D145–D149. http://modomics.genesilico.pl/
Yarian C, Townsend H, Czestkowski W, Sochacka E, Malkiewicz AJ, Guenther R, Miskiewicz A, Agris PF (2002) Accurate translation of the genetic code depends on tRNA modified nucleosides. J Biol Chem 277:16391–16395
Beuning PJ, Musier-Forsyth K (1999) Transfer RNA recognition by aminoacyl-tRNA synthetases. Biopolymers 52:1–28
Sylvers LA, Rogers KC, Shimizu M, Ohtsuka E, Söll D (1993) A 2-thiouridine derivative in tRNAGlu is a positive determinant for aminoacylation by Escherichia coli glutamyl-tRNA synthetase. Biochemistry 32:3836–3841
Gustilo EM, Dubois DY, Lapointe J, Agris PF (2007) E. coli glutamyl-tRNA synthetase is inhibited by anticodon stem-loop domains and a minihelix. RNA Biol 4:85–92
Isel C, Marquet R, Keith G, Ehresmann C, Ehresmann B (1993) Modified nucleotides of tRNA(3Lys) modulate primer/template loop-loop interaction in the initiation complex of HIV-1 reverse transcription. J Biol Chem 268:25269–25272
Bilbille Y, Vendeix FA, Guenther R, Malkiewicz A, Ariza X, Vilarrasa J, Agris PF (2009) The structure of the human tRNALys3 anticodon bound to the HIV genome is stabilized by modified nucleosides and adjacent mismatch base pairs. Nucl Acids Res 37:3342–3353
Ching WM, Wittwer AJ, Tsai L, Stadtman TC (1984) Distribution of two selenonucleosides among the selenium-containing tRNAs from Methanococcus vannielii. Proc Natl Acad Sci USA 81:57–60
Ching WM (1984) Occurrence of selenium-containing tRNAs in mouse leukemia cells. Proc Natl Acad Sci USA 81:3010–3013
Chen CS, Stadtman TC (1980) Selenium-containing tRNAs from Clostridium sticklandii: cochromatography of one species with l-prolyl-tRNA. Proc Natl Acad Sci USA 77:1403–1407
Ching WM, Stadtman TC (1982) Selenium-containing tRNAGlu from Clostridium sticklandii: correlation of aminoacylation with selenium content. Proc Natl Acad Sci USA 79:374–377
Wittwer AJ (1983) Specific incorporation of selenium into lysine- and glutamate-accepting tRNAs from Escherichia coli. J Biol Chem 258:8637–8641
Wen TN, Li C, Chen CS (1988) Ubiquity of selenium-containing tRNA in plants. Plant Sci 57:185–193
Corsaro A, Pistara V (1998) Conversion of the thiocarbonyl group into the carbonyl group. Tetrahedron 54:15027–15062
Carbon J, Hung L, Jones DS (1965) A reversible oxidative inactivation of specific transfer RNA species. Proc Natl Acad Sci USA 53:979–986
Rao PYS, Cherayil JD (1974) Studies on chemical modification of thionucleosides in transfer ribonucleic acid of Escherichia coli. Biochem J 143:285–294
Watanabe K (1980) Reaction of 2-thioribothymidine and 4-thiouridine with hydrogen peroxide in transfer ribonucleic acids from Thermus thermophilus and Escherichia coli studied by circular dichroism. Biochemistry 19:5542–5549
Houssier C, Degee P, Nicoghosian K, Grosjean H (1988) Effect of uridine dethiolation in anticodon triplet of tRNA (Glu) on its association with tRNA(Phe). J Biomol Struct Dynam 5:1259–1265
Götte M, Marquet R, Isel C, Anderson VE, Keith G, Gross HJ, Ehresmann C, Ehresmann B, Heumann H (1996) Probing the higher order structure of RNA with peroxonitrous acid. FEBS Lett 390:226–228
Brown DJ, Evans RF, Cowden WB, Fenn MD (2008) Thiopyrimidines. In: Taylor EC (ed) Chemistry of heterocyclic compounds: the pyrimidines, vol chap 52. Wiley, London, pp 553–609
Mitsunobu O, Ito N, Saita S, Ogihara T, Tamaoki H, Nagasawa H, Suzuki H, Kimira J (1982) Studies on nucleosides and nucleotides. X. Desulfurization of 2′,3′-O-isopropylidene-2-thiouridine by dibenzoyldiazene. Tetrahedron Lett 23:517–520
Ogihara T, Mitsunobu O (1982) Desulfurization of 2-thiouridines by dipotassium diazenedicarboxylate. Chem Lett 11:1621–1624
Rajur SB, McLaughlin LW (1992) The synthesis of oligodeoxynucleotides containing 2-thiothymidine and 5-methyl-4-pyrimidinone base analoques. Tetrahedron Lett 41:6081–6084
Kuimelis RG, Nambiar KP (1993) Efficient desulfurization of 2-thiopyrimidine nucleosides to the corresponding 4-pyrimidinones. Tetrahedron Lett 34:3813–3816
Saladino R, Mincione E, Cearsini C, Mezzetti M (1996) Transformations of thiopyrimidine and thiopurine nucleosides following oxidation with dimethyldioxirane. Tetrahedron 52:6759–6780
Sochacka E, Fratczak I (2004) Efficient desulfurization of 2-thiopyrimidine nucleosides to corresponding 4-pyrimidinone analogues using trans-2-phenylsulfonyl-3-phenyloxaziridine. Tetrahedron Lett 45:6729–6731
Kuimelis RG, Nambiar KP (1994) Synthesis of oligodeoxynucleotides containing 2-thiopyrimidine residues–new protection scheme. Nucl Acids Res 22:1429–1436
Kumar RK, Davis DR (1995) Synthesis of oligoribonucleotides containing 2-thiouridine: Incorporation of 2-thiouridine phosphoramidite without base protection. J Org Chem 60:7726–7727
Sochacka E (2001) Efficient assessment of modified nucleoside stability under conditions of automated oligonucleotide synthesis: characterization of the oxidation and oxidative desulfurization of 2-thiouridine. Nucleosides Nucleotides Nucleic Acids 20:1871–1879
Okamoto I, Seio K, Sekine M (2006) Improved synthesis of oligonucleotides containing 2-thiouridine derivatives by use of diluted iodine solution. Tetrahedron Lett 47:583–585
Kraszewska K, Kaczyńska I, Jankowski S, Karolak-Wojciechowska J, Sochacka E (2011) Desulfurization of 2-thiouracil nucleosides: conformational studies of 4-pyrimidinone nucleosides. Bioorg Med Chem 19:2443–2449
Sochacka E, Malkiewicz A (1991) XIV-th international tRNA workshop, Rydzyna, Poland
Sochacka E, Kraszewska K, Sochacki M, Sobczak M, Janicka M, Nawrot B (2011) The 2-thiouridine unit in the RNA strand is desulfured predominantly to 4-pyrimidinone nucleoside under in vitro oxidative stress conditions. Chem Commun 47:4914–4916
Davis RD (1998) Biophysical and conformational properties of modified nucleosides in RNA (nuclear magnetic resonance studies). In: Grosjean H, Benne R (eds) Modification and editing of RNA. ASM Press, Washington DC, pp 85–102
Guerra CF, Baerends EJ, Bickelhaupt FM (2008) Watson–Crick base pairs with thiocarbonyl groups: how sulfur changes the hydrogen bonds in DNA. Cent Eur J Chem 6:15–21
Yamamoto Y, Yokoyama S, Miyazawa T, Watanabe K, Higuchi S (1983) NMR analyses on the molecular mechanism of the conformational rigidity of 2-thioribothymidine, a modified nucleoside in extreme thermophile tRNAs. FEBS Lett 157:95–99
Smith WS, Sierzputowska-Gracz H, Sochacka E, Malkiewicz A, Agris PF (1992) Chemistry and structure of modified uridine dinucleosides are determined by thiolation. J Am Chem Soc 114:7990–7997
Zhang R, Erikson LA (2010) Theoretical study on conformational preferences of ribose in 2-thiouridine—the role of the 2′OH group. Phys Chem Chem Phys 12:3690–3697
Kumar RK, Davis DR (1997) Synthesis and studies on the effect of 2-thiouridine and 4-thiouridine on sugar conformation and RNA duplex stability. Nucl Acids Res 25:1272–1280
Testa SM, Disney MD, Turner DH, Kierzek R (1997) Thermodynamics of RNA-RNA duplexes with 2- or 4-thiouridines: implications for antisense design and targeting a group I intron. Biochemistry 38:16655–16662
Agris PF, Sierzputowska-Gracz H, Smith W, Malkiewicz A, Sochacka E, Nawrot B (1992) Thiolation of uridine carbon-2 restricts the motional dynamics of the transfer RNA wobble position nucleoside. J Am Chem Soc 114:2652–2656
Shohda K, Okamoto I, Wada S, Seio K, Sekine M (2000) Synthesis and properties of 2′-O-methyl-2-thiouridine and oligoribonucleotides containing 2′-O-methyl-2-thiouridine. Bioorg Med Chem Lett 10:1795–1798
Okamoto I, Seio K, Sekine M (2008) Study of the base discrimination ability of DNA and 2′-O-metylated RNA oligomers containing 2-thiouracil bases towards complementary RNA or DNA strands and their application to single base mismatch detection. Bioorg Med Chem 16:6034–6041
Sipa K, Sochacka E, Kazmierczak-Baranska J, Maszewska M, Janicka M, Nowak G, Nawrot B (2007) Effect of base modifications on structure, thermodynamic stability, and gene silencing activity of short interfering RNA. RNA 13:1301–1316
Sierant M, Paduszynska A, Kazmierczak-Baranska J, Nacmias B, Sorbi S, Bagnoli S, Sochacka E, Nawrot B (2011) Specific silencing of L392V PSEN1 mutant allele by RNA interference. Int J Alzheimers Dis 2011:809218
Iocono JA, Gildea B, McLaughlin LM (1990) Mild acid hydrolysis of 2-pyrimidinone—containing DNA fragment generates apurinic/apyrimidinic sites. Tetrahedron Lett 31:175–178
Sierzputowska-Gracz H, Sochacka E, Małkiewicz A, Kuo K, Gehrke CW, Agris PF (1987) Chemistry and structure of modified uridines in the anticodon, wobble position of transfer RNA are determined by thiolation. J Am Chem Soc 109:7171–7177
Schleich T, Blackburn BJ, Lapper RD, Smith ICP (1972) A nuclear magnetic resonance study of the influence of aqueous sodium perchlorate and temperature on the solution conformation of uracil nucleosides and nucleotides. Biochemistry 11:137–145
Acknowledgments
The work was financially supported by the Centre for Molecular and Macromolecular Studies Lodz, Polish Academy of Sciences.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Nawrot, B., Sochacka, E. & Düchler, M. tRNA structural and functional changes induced by oxidative stress. Cell. Mol. Life Sci. 68, 4023–4032 (2011). https://doi.org/10.1007/s00018-011-0773-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-011-0773-8