Abstract
Paracellular barrier properties of tissues are mainly determined by the composition of claudin heteropolymers. To analyze the molecular organization of tight junctions (TJ), we investigated the ability of claudins (Cld) to form homo- and heteromers. Cld1, -2, -3, -5, and -12 expressed in cerebral barriers were investigated. TJ-strands were reconstituted by claudin-transfection of HEK293-cells. cis-Interactions and/or spatial proximity were analyzed by fluorescence resonance energy transfer inside and outside of strands and ranked: Cld5/Cld5 > Cld5/Cld1 > Cld3/Cld1 > Cld3/Cld3 > Cld3/Cld5, no Cld3/Cld2. Classic Cld1, -3, and -5 but not non-classic Cld12 showed homophilic trans-interaction. Freeze-fracture electron microscopy revealed that, in contrast to classic claudins, YFP-tagged Cld12 does not form homopolymers. Heterophilic trans-interactions were analyzed in cocultures of differently monotransfected cells. trans-Interaction of Cld3/Cld5 was less pronounced than that of Cld3/Cld1, Cld5/Cld1, Cld5/Cld5 or Cld3/Cld3. The barrier function of reconstituted TJ-strands was demonstrated by a novel imaging assay. A model of the molecular organization of TJ was generated.










Similar content being viewed by others
References
Angelow S, Ahlstrom R, Yu AS (2008) Biology of claudins. Am J Physiol Renal Physiol 295:F867–F876
Staehelin LA (1974) Structure and function of intercellular junctions. Int Rev Cytol 39:191–283
Piontek J, Winkler L, Wolburg H, Muller SL, Zuleger N, Piehl C, Wiesner B, Krause G, Blasig IE (2008) Formation of tight junction: determinants of homophilic interaction between classic claudins. FASEB J 22:146–158
Morita K, Furuse M, Fujimoto K, Tsukita S (1999) Claudin multigene family encoding four-transmembrane domain protein components of tight junction strands. Proc Natl Acad Sci USA 96:511–516
Krause G, Winkler L, Mueller SL, Haseloff RF, Piontek J, Blasig IE (2008) Structure and function of claudins. Biochim Biophys Acta 1778:631–645
Nitta T, Hata M, Gotoh S, Seo Y, Sasaki H, Hashimoto N, Furuse M, Tsukita S (2003) Size-selective loosening of the blood–brain barrier in claudin-5-deficient mice. J Cell Biol 161:653–660
Hou J, Renigunta A, Konrad M, Gomes AS, Schneeberger EE, Paul DL, Waldegger S, Goodenough DA (2008) Claudin-16 and claudin-19 interact and form a cation-selective tight junction complex. J Clin Invest 118:619–628
Hou J, Renigunta A, Gomes AS, Hou M, Paul DL, Waldegger S, Goodenough DA (2009) Claudin-16 and claudin-19 interaction is required for their assembly into tight junctions and for renal reabsorption of magnesium. Proc Natl Acad Sci USA 106:15350–15355
Yu AS, Cheng MH, Angelow S, Gunzel D, Kanzawa SA, Schneeberger EE, Fromm M, Coalson RD (2009) Molecular basis for cation selectivity in claudin-2-based paracellular pores: identification of an electrostatic interaction site. J Gen Physiol 133:111–127
Daugherty BL, Ward C, Smith T, Ritzenthaler JD, Koval M (2007) Regulation of heterotypic claudin compatibility. J Biol Chem 282:30005–30013
Piehl C, Piontek J, Cording J, Wolburg H, Blasig IE (2010) Participation of the second extracellular loop of claudin-5 in paracellular tightening against ions, small and large molecules. Cell Mol Life Sci 67:2131–2140
Zhang J, Piontek J, Wolburg H, Piehl C, Liss M, Otten C, Christ A, Willnow TE, Blasig IE, Abdelilah-Seyfried S (2010) Establishment of a neuroepithelial barrier by Claudin5a is essential for zebrafish brain ventricular lumen expansion. Proc Natl Acad Sci USA 107:1425–1430
Furuse M, Sasaki H, Tsukita S (1999) Manner of interaction of heterogeneous claudin species within and between tight junction strands. J Cell Biol 147:891–903
Wolburg H, Lippoldt A (2002) Tight junctions of the blood–brain barrier: development, composition and regulation. Vascul Pharmacol 38:323–337
Kniesel U, Risau W, Wolburg H (1996) Development of blood–brain barrier tight junctions in the rat cortex. Dev Brain Res 96:229–240
Liebner S, Fischmann A, Rascher G, Duffner F, Grote EH, Kalbacher H, Wolburg H (2000) Claudin-1 and claudin-5 expression and tight junction morphology are altered in blood vessels of human glioblastoma multiforme. Acta Neuropathol 100:323–331
Wolburg H, Wolburg-Buchholz K, Kraus J, Rascher-Eggstein G, Liebner S, Hamm S, Duffner F, Grote EH, Risau W, Engelhardt B (2003) Localization of claudin-3 in tight junctions of the blood–brain barrier is selectively lost during experimental autoimmune encephalomyelitis and human glioblastoma multiforme. Acta Neuropathol 105:586–592
Winkler L, Gehring C, Wenzel A, Muller SL, Piehl C, Krause G, Blasig IE, Piontek J (2009) Molecular determinants of the interaction between Clostridium perfringens enterotoxin fragments and claudin-3. J Biol Chem 284:18863–18872
Blasig IE, Winkler L, Lassowski B, Mueller SL, Zuleger N, Krause E, Krause G, Gast K, Kolbe M, Piontek J (2006) On the self-association potential of transmembrane tight junction proteins. Cell Mol Life Sci 63:505–514
Shen L, Weber CR, Turner JR (2008) The tight junction protein complex undergoes rapid and continuous molecular remodeling at steady state. J Cell Biol 181:683–695
Yu D, Marchiando AM, Weber CR, Raleigh DR, Wang YM, Shen L, Turner JR (2010) MLCK-dependent exchange and actin binding region-dependent anchoring of ZO-1 regulate tight junction barrier function. Proc Natl Acad Sci USA 107:8237–8241
Mack AF, Wolburg H (2006) Growing axons in fish optic nerve are accompanied by astrocytes interconnected by tight junctions. Brain Res 1103:25–31
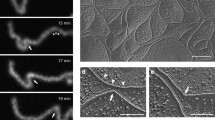
Matsuda M, Kubo A, Furuse M, Tsukita S (2004) A peculiar internalization of claudins, tight junction-specific adhesion molecules, during the intercellular movement of epithelial cells. J Cell Sci 117:1247–1257
Wallrabe H, Elangovan M, Burchard A, Periasamy A, Barroso M (2003) Confocal FRET microscopy to measure clustering of ligand-receptor complexes in endocytic membranes. Biophys J 85:559–571
Banning C, Votteler J, Hoffmann D, Koppensteiner H, Warmer M, Reimer R, Kirchhoff F, Schubert U, Hauber J, Schindler M (2010) A flow cytometry-based FRET assay to identify and analyse protein–protein interactions in living cells. Plos One 5
van Wageningen S, Pennings AH, van der Reijden BA, Boezeman JB, de Lange F, Jansen JH (2006) Isolation of FRET-positive cells using single 408-nm laser flow cytometry. Cytom Part A 69A:291–298
Di WL, Gu Y, Common JEA, Aasen T, O’Toole EA, Kelsell DP, Zicha D (2005) Connexin interaction patterns in keratinocytes revealed morphologically and by FRET analysis. J Cell Sci 118:1505–1514
Amiri H, Schultz G, Schaefer M (2003) FRET-based analysis of TRPC subunit stoichiometry. Cell Calcium 33:463–470
Morita K, Sasaki H, Furuse M, Tsukita S (1999) Endothelial claudin: claudin-5/TMVCF constitutes tight junction strands in endothelial cells. J Cell Biol 147:185–194
Furuse M, Sasaki H, Fujimoto K, Tsukita S (1998) A single gene product, claudin-1 or -2, reconstitutes tight junction strands and recruits occludin in fibroblasts. J Cell Biol 143:391–401
Morita K, Sasaki H, Fujimoto K, Furuse M, Tsukita S (1999) Claudin-11/OSP-based tight junctions of myelin sheaths in brain and sertoli cells in testis. J Cell Biol 145:579–588
Inai T, Kamimura T, Hirose E, Iida H, Shibata Y (2010) The protoplasmic or exoplasmic face association of tight junction particles cannot predict paracellular permeability or heterotypic claudin compatibility. Eur J Cell Biol 89:547–556
Coyne CB, Gambling TM, Boucher RC, Carson JL, Johnson LG (2003) Role of claudin interactions in airway tight junctional permeability. Am J Physiol Lung Cell Mol Physiol 285:L1166–L1178
Sekar RB, Periasamy A (2003) Fluorescence resonance energy transfer (FRET) microscopy imaging of live cell protein localizations. J Cell Biol 160:629–633
Mitic LL, Unger VM, Anderson JM (2003) Expression, solubilization, and biochemical characterization of the tight junction transmembrane protein claudin-4. Protein Sci 12:218–227
Van Itallie CM, Mitic LL, Anderson JM (2010) Claudin-2 forms homodimers and is a component of a high molecular weight protein complex. J Biol Chem
Harris HJ, Davis C, Mullins JG, Hu K, Goodall M, Farquhar MJ, Mee CJ, McCaffrey K, Young S, Drummer H, Balfe P, McKeating JA (2010) Claudin association with CD81 defines hepatitis C virus entry. J Biol Chem 285:21092–21102
Liebner S, Kniesel U, Kalbacher H, Wolburg H (2000) Correlation of tight junction morphology with the expression of tight junction proteins in blood–brain barrier endothelial cells. Eur J Cell Biol 79:707–717
Acknowledgments
We gratefully acknowledge the help of Ria Knittel in freeze-fracturing. This work was funded by DFG BL308/7-3, 7-4 and PI 837/2-1 and NIH DK61931.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
18_2011_680_MOESM1_ESM.pdf
Supplementary material 1 The FRET-efficiency is strongly dependent on the ratio of acceptor/donor (YFP/CFP) fluorescence intensity (A, R2 = 0.51) but not on the YFP fluorescence intensity (B, R2 = 0.02) or the CFP+YFP fluorescence intensity (C, R2 = 0.02) as a measure of the expression level. Individual data points and non-linear regression curve for Cld5-CFP/Cld5-YFP. (PDF 45 kb)
18_2011_680_MOESM2_ESM.pdf
Supplementary material 2 (A) Flow cytometric expression analysis of HEK cells transfected with different constructs. Non- (mock), CFP- or YFP-transfection were used to account for auto fluorescence. CFP/YFP cotransfection and CFP-YFP tandem were used as FRET negative or positive control, respectively. Cdl5-CFP/CRFR1-YFP and Cld5-CFP/Cld5-YFP were used as pairs of transmembrane proteins shown to be FRET-negative or -positive, respectively, by conventional method (Piontek et al., 2008). On the axes the channels are given. FSC, forward scatter. (B) Visualization of FRET signal by comparison of CFP/YFP - FRET negative -, CFP-YFP tandem - FRET positive - (left panel), Cld5-CFP/CRFR1-YFP - FRET negative - and Cld5-CFP/Cld5-YFP - FRET positive - (right panel). FRET-signal is indicated by left shift. (C) Normalization of FRET and YFP values relative to CFP intensity with the corresponding exponential decay fits (left: CFP/YFP (black dots, green line) and CFP-YFP (red dots, blue line); right: Cld5-CFP/CRFR1-YFP (black dots, green line) and Cld5-CFP/Cld5-YFP (red dots, blue line). The fits were used to calculate the FRET-ratioFC as a measure of FRET- efficiency (see Methods). (PDF 455 kb)
18_2011_680_MOESM3_ESM.pdf
Supplementary material 3 FRET-analysis in intracellular compartments by confocal microscopy. For Cld5-CFP/Cld5-YFP, FRET was significant higher than for Cld3-CFP/Cld3-YFP for which FRET was still higher than for the negative control (CFP-tagged endoplasmic reticulum marker, CFP-ER (BD Biosciences) coexpressed with Cld5-YFP, CFP-ER/Cld5-YFP). 36, 34, and 19 cell–cell contacts were analyzed, respectively; p < 0.001. (PDF 35 kb)
18_2011_680_MOESM4_ESM.pdf
Supplementary material 4 In intracellular compartments claudins did not considerably colocalize with markers for endosomes, Golgi apparatus or lysosomes but partly with that for the endoplasmic reticulum (A, arrow). HEK 293 cells were cotransfected with Cld5-YFP and (A) CFP-ER (marker for endoplasmic reticulum, Clontech), (B) CFP-Endo (marker for endosomes, Clontech) or (C, left) CFP-Golgi (marker for Golgi apparatus, Clontech) and 3 days later analyzed by confocal microscopy. In addition, cells were transfected with Cld5 (C, middle) or Cld3 (C, right) and 3 days later incubated with 50 nM LysoTracker (Invitrogen) in growth medium for 30 min at 37°C, the medium exchanged by DMEM with 10 mM N-(2-hydroxyethyl)piperazine-N’(2-ethanesulfonic acid) pH 7.5 without phenol red and analyzed by confocal microscopy. Bar, 5 µm. (PDF 105 kb)
18_2011_680_MOESM5_ESM.pdf
Supplementary material 5 CellMask labels the apical and basolateral plasma membrane; reconstituted TJs delay the labeling. HEK cells stably transfected with Cld5 were labeled with 0.25 μg/ml CellMask for 5 min (A, B) or 20 min (C, D). After 5 min, CellMask labeling (red) is detected in the apical (ap) and basal (bl) part of the lateral plasma membrane but not at the Cld5-positive (green) area at cell–cell contacts (A,B). In contrast, after 20 min, CellMask (red) colocalizes with claudin-5 (green) at cell–cell contacts (C, D). Merge (A, C); CellMalsk (B, D); arrow, Cld5-enrichment at cell–cell contact; arrowhead, apical and basolateral plasma membrane adjacent to claudin-5; bar, 5 µm. (PDF 272 kb)
Rights and permissions
About this article
Cite this article
Piontek, J., Fritzsche, S., Cording, J. et al. Elucidating the principles of the molecular organization of heteropolymeric tight junction strands. Cell. Mol. Life Sci. 68, 3903–3918 (2011). https://doi.org/10.1007/s00018-011-0680-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-011-0680-z