Abstract.
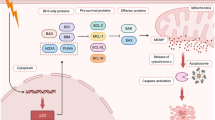
We describe a general strategy for the identification of functional genes that, when downregulated, result in a selectable phenotype. This strategy is based on expression selection of cDNA fragments that counteract their cognate genes. A cDNA library containing random fragments expressed in human HepG2, A375 and CLS-354 cells was used to identify functional genes whose inhibition conferred resistance to Fas-induced apoptosis. Thirty-five clones were isolated, 28 of which were derived from unknown genes, that tagged 19 individual genes and 7 of which referred to known genes that tagged the apoptosis-related protein (APR)-1, -2 and indoleamine-pyrrole 2,3,-dioxygenase (IDO). The ability of APR-1-, -2- and IDO-derived antisense RNAs to induce resistance to Fas in HepG2, A375 and CLS-354 cells suggested that APR-1, -2 and IDO genes are involved in the machinery of Fas-mediated apoptosis. Our gene discovery strategy provides a generally applicable procedure to identify functional genes that interfere with apoptosis, and may therefore be clinically relevant for tumor therapy.
Similar content being viewed by others
Author information
Authors and Affiliations
Corresponding author
Additional information
Received 28 April 2005; received after revision 20 June 2005; accepted 5 July 2005
Rights and permissions
About this article
Cite this article
Hassan, M., Mirmohammadsadegh, A., Selimovic, D. et al. Identification of functional genes during Fas-mediated apoptosis using a randomly fragmented cDNA library. Cell. Mol. Life Sci. 62, 2015–2026 (2005). https://doi.org/10.1007/s00018-005-5172-6
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00018-005-5172-6