Abstract
Objective
To identify CD8+ T cell-related molecular clusters and establish a novel gene signature for predicting the prognosis and efficacy of immunotherapy in bladder cancer (BCa).
Methods
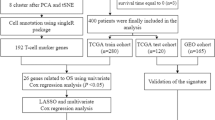
Transcriptome and clinical data of BCa samples were obtained from the Cancer Genome Atlas (TCGA) and GEO databases. The CD8+ T cell-related genes were screened through the CIBERSORT algorithm and correlation analysis. Consensus clustering analysis was utilized to identified CD8+ T cell-related molecular clusters. A novel CD8+ T cell-related prognostic model was developed using univariate Cox regression analysis and Lasso regression analysis. Internal and external validations were performed and the validity of the model was validated in a real-world cohort. Finally, preliminary experimental verifications were carried out to verify the biological functions of SH2D2A in bladder cancer.
Results
A total of 52 CD8+ T cell-related prognostic genes were screened and two molecular clusters with notably diverse immune cell infiltration, prognosis and clinical features were developed. Then, a novel CD8+ T cell-related prognostic model was constructed. The patients with high-risk scores exhibited a significantly worse overall survival in training, test, whole TCGA and validating cohort. The AUC was 0.766, 0.725, 0.739 and 0.658 in the four cohorts sequentially. Subgroup analysis suggested that the novel prognostic model has a robust clinical application for selecting high-risk patients. Finally, we confirmed that patients in the low-risk group might benefit more from immunotherapy or chemotherapy, and validated the prognostic model in a real-world immunotherapy cohort. Preliminary experiment showed that SH2D2A was capable of attenuating proliferation, migration and invasion of BCa cells.
Conclusions
CD8+ T cell-related molecular clusters were successfully identified. Besides, a novel CD8+ T cell-related prognostic model with an excellent predictive performance in predicting survival rates and immunotherapy efficacy of BCa was developed.











Similar content being viewed by others
Availability of data and materials
The data presented in the study are included in the article/Supplementary Material, further inquiries can be directed to the corresponding authors.
References
Yang X, Ye T, Liu H, Lv P, Duan C, Wu X, Jiang K, Lu H, Xia D, Peng E, Chen Z, Tang K, Ye Z. Expression profiles, biological functions and clinical significance of circRNAs in bladder cancer. Mol Cancer. 2021;20:4.
Liu Y, Huang W, Cai Z. Synthesizing AND gate minigene circuits based on CRISPReader for identification of bladder cancer cells. Nat Commun. 2020;11:5486.
Lu Q, Liu T, Feng H, Yang R, Zhao X, Chen W, Jiang B, Qin H, Guo X, Liu M, Li L, Guo H. Circular RNA circSLC8A1 acts as a sponge of miR-130b/miR-494 in suppressing bladder cancer progression via regulating PTEN. Mol Cancer. 2019;18:111.
Kita Y, Hamada A, Saito R, Teramoto Y, Tanaka R, Takano K, Nakayama K, Murakami K, Matsumoto K, Akamatsu S, Yamasaki T, Inoue T, Tabata Y, Okuno Y, Ogawa O, Kobayashi T. Systematic chemical screening identifies disulfiram as a repurposed drug that enhances sensitivity to cisplatin in bladder cancer: a summary of preclinical studies. Br J Cancer. 2019;121:1027–38.
Yadollahvandmiandoab R, Jalalizadeh M, Buosi K, Garcia-Perdomo HA, Reis LO. Immunogenic cell death role in urothelial cancer therapy. Curr Oncol (Tor, Ont). 2022;29:6700–13.
Ren D, Hua Y, Yu B, Ye X, He Z, Li C, Wang J, Mo Y, Wei X, Chen Y, Zhou Y, Liao Q, Wang H, Xiang B, Zhou M, Li X, Li G, Li Y, Zeng Z, Xiong W. Predictive biomarkers and mechanisms underlying resistance to PD1/PD-L1 blockade cancer immunotherapy. Mol Cancer. 2020;19:19.
Wrona E, Borowiec M, Potemski P. CAR-NK cells in the treatment of solid tumors. Int J Mol Sci. 2021;22:5899.
Deng C, Xu Y, Fu J, Zhu X, Chen H, Xu H, Wang G, Song Y, Song G, Lu J, Liu R, Tang Q, Huang W, Wang J. Reprograming the tumor immunologic microenvironment using neoadjuvant chemotherapy in osteosarcoma. Cancer Sci. 2020;111:1899–909.
Leclerc M, Voilin E, Gros G, Corgnac S, de Montpréville V, Validire P, Bismuth G, Mami-Chouaib F. Regulation of antitumour CD8 T-cell immunity and checkpoint blockade immunotherapy by Neuropilin-1. Nat Commun. 2019;10:3345.
Liu A, Xia Y, Li W, Zhang G, Liu Y, Ye S, Zhao ZR, Yang Y, Jia Y, Guo Y, Liu X, Chen H, Yu J. The predictive value of changes in the absolute counts of peripheral lymphocyte subsets for progression and prognosis in breast cancer patients. Contrast Media Mol Imaging. 2022;2022:3444360.
Chabab G, Boissière-Michot F, Mollevi C, Ramos J, Lopez-Crapez E, Colombo PE, Jacot W, Bonnefoy N, Lafont V. Diversity of tumor-infiltrating, γδ T-cell abundance in solid cancers. Cells. 2020;9:1537.
Luo D, Kuang F, Du J, Zhou M, Peng F, Gan Y, Fang C, Yang X, Li B, Su S. Characterization of the immune cell infiltration profile in pancreatic carcinoma to aid in immunotherapy. Front Oncol. 2021;11: 677609.
Shimizu S, Hiratsuka H, Koike K, Tsuchihashi K, Sonoda T, Ogi K, Miyakawa A, Kobayashi J, Kaneko T, Igarashi T, Hasegawa T, Miyazaki A. Tumor-infiltrating CD8(+) T-cell density is an independent prognostic marker for oral squamous cell carcinoma. Cancer Med. 2019;8:80–93.
He D, Zhang X, Tu J. Diagnostic significance and carcinogenic mechanism of pan-cancer gene POU5F1 in liver hepatocellular carcinoma. Cancer Med. 2020;9:8782–800.
Sun M, Ji X, Xie M, Chen X, Zhang B, Luo X, Feng Y, Liu D, Wang Y, Li Y, Liu B, Xia L, Huang W. Identification of necroptosis-related subtypes, development of a novel signature, and characterization of immune infiltration in colorectal cancer. Front Immunol. 2022;13: 999084.
Goutas D, Palamaris K, Stofas A, Politakis N, Despotidi A, Giannopoulou I, Goutas N, Vlachodimitropoulos D, Kavantzas N, Lazaris AC, Gakiopoulou H. Immunohistochemical study of bladder cancer molecular subtypes and their association with PD-L1 expression. Cancers. 2022;15:188.
Voutsadakis IA. Urothelial bladder carcinomas with high tumor mutation burden have a better prognosis and targetable molecular defects beyond immunotherapies. Curr Oncol (Tor, Ont). 2022;29:1390–407.
Liu Z, Lu H, Jiang Z, Pastuszyn A, Hu CA. Apolipoprotein l6, a novel proapoptotic Bcl-2 homology 3-only protein, induces mitochondria-mediated apoptosis in cancer cells. Mol Cancer Res. 2005;3:21–31.
Scorrano L, Korsmeyer SJ. Mechanisms of cytochrome c release by proapoptotic BCL-2 family members. Biochem Biophys Res Commun. 2003;304:437–44.
Raji GR, Poyyakkara A, Krishnan AK, Maurya AK, Changmai U, Shankar SS, Kumar VBS. Horizontal transfer of miR-643 from cisplatin-resistant cells confers chemoresistance to recipient drug-sensitive cells by targeting APOL6. Cells. 2021;10:1341.
Liu K, Chen Y, Li B, Li Y, Liang X, Lin H, Luo L, Chen T, Dai Y, Pang W, Zeng L. Upregulation of apolipoprotein L6 improves tumor immunotherapy by inducing immunogenic cell death. Biomolecules. 2023;13:415.
Steidl C, Shah SP, Woolcock BW, Rui L, Kawahara M, Farinha P, Johnson NA, Zhao Y, Telenius A, Neriah SB, McPherson A, Meissner B, Okoye UC, Diepstra A, van den Berg A, Sun M, Leung G, Jones SJ, Connors JM, Huntsman DG, Savage KJ, Rimsza LM, Horsman DE, Staudt LM, Steidl U, Marra MA, Gascoyne RD. MHC class II transactivator CIITA is a recurrent gene fusion partner in lymphoid cancers. Nature. 2011;471:377–81.
Mottok A, Woolcock B, Chan FC, Tong KM, Chong L, Farinha P, Telenius A, Chavez E, Ramchandani S, Drake M, Boyle M, Ben-Neriah S, Scott DW, Rimsza LM, Siebert R, Gascoyne RD, Steidl C. Genomic alterations in CIITA are frequent in primary mediastinal large B cell lymphoma and are associated with diminished MHC class II expression. Cell Rep. 2015;13:1418–31.
Neuwelt AJ, Kimball AK, Johnson AM, Arnold BW, Bullock BL, Kaspar RE, Kleczko EK, Kwak JW, Wu MH, Heasley LE, Doebele RC, Li HY, Nemenoff RA, Clambey ET. Cancer cell-intrinsic expression of MHC II in lung cancer cell lines is actively restricted by MEK/ERK signaling and epigenetic mechanisms. J Immunother Cancer. 2020;8:e000441.
Redelman-Sidi G, Binyamin A, Antonelli AC, Catalano W, Bean J, Al-Ahmadie H, Jungbluth AA, Glickman MS. BCG-induced tumor immunity requires tumor-intrinsic CIITA independent of MHC-II. Cancer Immunol Res. 2022;10:1241–53.
Pezzè L, Meškytė EM, Forcato M, Pontalti S, Badowska KA, Rizzotto D, Skvortsova II, Bicciato S, Ciribilli Y II. ETV7 regulates breast cancer stem-like cell features by repressing IFN-response genes. Cell Death Dis. 2021;12:742.
Qu H, Zhao H, Zhang X, Liu Y, Li F, Sun L, Song Z. Integrated analysis of the ETS family in melanoma reveals a regulatory role of ETV7 in the immune microenvironment. Front Immunol. 2020;11: 612784.
Harwood FC, Klein Geltink RI, O’Hara BP, Cardone M, Janke L, Finkelstein D, Entin I, Paul L, Houghton PJ, Grosveld GC. ETV7 is an essential component of a rapamycin-insensitive mTOR complex in cancer. Sci Adv. 2018;4:3938.
Sang Y, Cheng C, Zeng YX, Kang T. Snail promotes metastasis of nasopharyngeal carcinoma partly by down-regulating TEL2. Cancer Commun (Lond, Engl). 2018;38:58.
Li H, Zhang Y, Zheng S. Comprehensive analysis identified ETV7 as a potential prognostic biomarker in bladder cancer. Biomed Res Int. 2021;2021:8530186.
Wedel J, Stack MP, Seto T, Sheehan MM, Flynn EA, Stillman IE, Kong SW, Liu K, Briscoe DM. T cell-specific adaptor protein regulates mitochondrial function and CD4(+) T regulatory cell activity in vivo following transplantation. J Immuno (Baltimore, MD). 1950;203(2019):2328–38.
Abrahamsen G, Sundvold-Gjerstad V, Habtamu M, Bogen B, Spurkland A. Polarity of CD4+ T cells towards the antigen presenting cell is regulated by the Lck adapter TSAd. Sci Rep. 2018;8:13319.
Berge T, Grønningsæter IH, Lorvik KB, Abrahamsen G, Granum S, Sundvold-Gjerstad V, Corthay A, Bogen B, Spurkland A. SH2D2A modulates T cell mediated protection to a B cell derived tumor in transgenic mice. PLoS ONE. 2012;7: e48239.
Shi Y, Zhang D, Chen J, Jiang Q, Song S, Mi Y, Wang T, Ye Q. Interaction between BEND5 and RBPJ suppresses breast cancer growth and metastasis via inhibiting Notch signaling. Int J Biol Sci. 2022;18:4233–44.
Wu ZS, Wu S. The era of personalized treatments: updates on immunotherapy within urothelial of bladder cancer. Curr Urol. 2022;16:117–20.
Liu J, Tan Z, He J, Jin T, Han Y, Hu L, Song J, Huang S. Identification of three molecular subtypes based on immune infiltration in ovarian cancer and its prognostic value. Biosci Rep. 2020;40:BSR20201431.
Gabrilovich DI, Ostrand-Rosenberg S, Bronte V. Coordinated regulation of myeloid cells by tumours. Nat Rev Immunol. 2012;12:253–68.
Cristescu R, Aurora-Garg D, Albright A, Xu L, Liu XQ, Loboda A, Lang L, Jin F, Rubin EH, Snyder A, Lunceford J. Tumor mutational burden predicts the efficacy of pembrolizumab monotherapy: a pan-tumor retrospective analysis of participants with advanced solid tumors. J Immunother Cancer. 2022. https://doi.org/10.1136/jitc-2021-003091.
Rizvi H, Sanchez-Vega F, La K, Chatila W, Jonsson P, Halpenny D, Plodkowski A, Long N, Sauter JL, Rekhtman N, Hollmann T, Schalper KA, Gainor JF, Shen R, Ni A, Arbour KC, Merghoub T, Wolchok J, Snyder A, Chaft JE, Kris MG, Rudin CM, Socci ND, Berger MF, Taylor BS, Zehir A, Solit DB, Arcila ME, Ladanyi M, Riely GJ, Schultz N, Hellmann MD. Molecular determinants of response to anti-programmed cell death (PD)-1 and anti-programmed death-ligand 1 (PD-L1) blockade in patients with non-small-cell lung cancer profiled with targeted next-generation sequencing. J Clin Oncol. 2018;36:633–41.
Xie M, Chen D, Li Y, Liu X, Kuang D, Li X. Genetic mutation profiles and immune microenvironment analysis of pulmonary enteric adenocarcinoma. Diagn Pathol. 2022;17:30.
Hu FF, Liu CJ, Liu LL, Zhang Q, Guo AY. Expression profile of immune checkpoint genes and their roles in predicting immunotherapy response. Br Bioinform. 2021. https://doi.org/10.1093/bib/bbaa176.
Funding
The study was supported by the “Eyas Plan” Youth Top-notch Talent Project of Fujian Province (Grant number: SCYJHBJRC-XN2021) and Class B Talent Research Project of the First Affiliated Hospital of Fujian Medical University (Grant number: YJCRC-B-XN2022).
Author information
Authors and Affiliations
Contributions
FL, Z-BK, Y-TX wrote the original draft. NX and X-YX reviewed and edited the original draft. J-YC and HC contributed to the methodology. FL and Y-ZL did the investigation. Z-BK and X-DL contributed to formal analysis. Y-TX and YW curated the data. FL contributed to the conceptualization and the visualization. NX supervised the project. X-YX administrated the project. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
All authors declared that there were no conflicts of interest.
Ethical approval and informed consent
This study was approved by the Ethics Committee of the First Affiliated Hospital of Fujian Medical University and all patients provided written informed consent.
Consent for publication
Informed consent for publication of the manuscript was obtained from all authors.
Additional information
Responsible Editor: John Di Battista
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
11_2023_1772_MOESM1_ESM.tif
Supplementary file1 Supplemental Figure 1. Lasso regression analysis of key genes significantly related to CD8+ T cell infiltration (A-B). The 8‐year ROC curves of the risk model and other clinical characteristics in training cohort (C). Correlations between the two risk groups and TME score (D). The subgroup survival analysis between low-risk and high-risk groups in patients with age ≤65 and age >65 (E-G). The subgroup survival analysis between low-risk and high-risk groups in the female and male subgroups (H-J). *, P < 0.05; ***, P < 0.001. (TIF 21493 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lin, F., Ke, ZB., Xue, YT. et al. A novel CD8+ T cell-related gene signature for predicting the prognosis and immunotherapy efficacy in bladder cancer. Inflamm. Res. 72, 1665–1687 (2023). https://doi.org/10.1007/s00011-023-01772-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00011-023-01772-6