Abstract
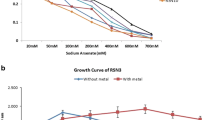
Rapid industrialization and intensive agriculture activities have led to a rise in heavy metal contamination all over the world. Chhattisgarh (India) being an industrial state, the soil and water are thickly contaminated with heavy metals, especially from arsenic (As). In the present study, we isolated 108 arsenic-resistant bacteria (both from soil and water) from different arsenic-contaminated industrial and mining sites of Chhattisgarh to explore the bacterial gene pool. Further, we screened 24 potential isolates out of 108 for their ability to tolerate a high level of arsenic. The sequencing of the 16S rRNA gene of bacterial isolates revealed that all these samples belong to different diverse genera including Bacillus, Enterobacter, Klebsiella, Pantoea, Acinetobacter, Cronobacter, Pseudomonas and Agrobacterium. The metal tolerance ability was determined by amplification of arsB (arsenite efflux gene) and arsC (arsenate reductase gene) from chromosomal DNA of isolated RnASA11, which was identified as Klebsiella pneumoniae through in silico analysis. The bacterial strains RpSWA2 and RnASA11 were found to tolerate 600 mM As (V) and 30 mM As (III) but the growth of strain RpSWA2 was slower than RnASA11. Furthermore, atomic absorption spectroscopy (AAS) of the sample obtained from bioremediation assay revealed that Klebsiella pneumoniae RnASA11 was able to reduce the arsenic concentration significantly in the presence of arsenate (44%) and arsenite (38.8%) as compared to control.




Similar content being viewed by others
References
Dash B, Soni R, Goel R (2019) Rhizobacteria for reducing heavy metal stress in plant and soil. In: Sayyed RZ, Arora NK, Reddy MS (eds) Plant growth promoting rhizobacteria for sustainable stress management volume 1: rhizobacteria in abiotic stress management. Springer, Singapore, pp 179–203
Soni R, Dash B, Kumar P, Mishra UN, Goel R (2019) Microbes for bioremediation of heavy metals. In: Singh DP, Prabha R (eds) Microbial interventions in agriculture and environment volume: soil and crop health management. Springer, Singapore, pp 129–142
Ratnaike RN (2003) Acute and chronic arsenic toxicity. Postgrad Med J 79:391–396. https://doi.org/10.1136/pmj.79.933.391
Sher H, Perry Z, Arbel S, Reuveni H (2019) Prescription patterns of antidepressants: the effect of the black box warning among pediatric patients. Res Health Sci 4:1. https://doi.org/10.22158/rhs.v4n1p1
Roane TM, Pepper IL (1999) Microbial responses to environmentally toxic cadmium. Microb Ecol 38:358–364. https://doi.org/10.1007/s002489901001
Yang HC, Rosen BP (2016) New mechanisms of bacterial arsenic resistance. Biomed J 3(9):5–13. https://doi.org/10.1016/j.bj.2015.08.003
Moghannem S, Refaat B, Elsherbiny G, El-Sayed M, Elsehemy I, Kalaba M (2015) Characterization of heavy metal and antibiotic-resistant bacteria isolated from polluted localities in Egypt. Egypt Pharmaceut J 14:158. https://doi.org/10.4103/1687-4315.172856
Chakraborti D, Rahman MM, Das B, Chatterjee A et al (2017) Groundwater arsenic contamination and its health effects in India. Hydrogeol J 25:1165–1181. https://doi.org/10.1007/s10040-017-1556-6
Patel KS, Shrivas K, Brandt R, Jakubowski N, Corns W, Hoffmann P (2005) Arsenic contamination in water, soil, sediment and rice of central India. Environ Geochem Health 27:131–145. https://doi.org/10.1007/s10653-005-0120-9
Shrivastava A, Ghosh D, Dash A, Bose S (2015) Arsenic contamination in soil and sediment in India: sources, effects, and remediation. Curr Pollution Rep 1(2015):35–46. https://doi.org/10.1007/s40726-015-0004-2
Yadav A et al (2020) Assessment of arsenic and heavy metal pollution in Chhattisgarh. India J Hazard Toxic Radioact Waste. https://doi.org/10.1061/(ASCE)HZ.2153-5515.0000478
Kumar P, Gupta SB, Anurag SR (2019) Bioremediation of cadmium by mixed indigenous isolates Serratia liquefaciens BSWC3 and Klebsiella pneumoniae RpSWC3 isolated from industrial and mining affected water samples. Pollution 5:351–360. https://doi.org/10.22059/POLL.2018.268603.533
Simeonova D, Lievremont D, Lagarde F, Muller D, Groudeva V, Lett MC (2004) Microplate screening assay for detection of arsenite oxidizing and arsenate-reducing bacteria. FEMS Microbiol Lett 237:249–253
Lu JJ, Perng CL, Lee SY, Wan CC (2000) Use of PCR with universal primers and restriction endonuclease digestions for detection and identification of common bacterial pathogens in cerebrospinal fluid. J Clin Microbiol 38(6):2076–2080
Achour AR, Bauda P, Billard P (2007) Diversity of arsenite transporter genes from arsenic-resistant soil bacteria. Res Microbiol 158:128–137. https://doi.org/10.1016/j.resmic.2006.11.006
Saltikov CW, Olson BH (2002) Homology of Escherichia coli R773 arsA, arsB, and arsC genes in arsenic-resistant bacteria isolated from raw sewage and arsenic-enriched creek waters. Appl Environ Microbiol 68:280–288. https://doi.org/10.1128/aem.68.1.280-288.2002
Sher S, Zajif Hussain S, Rehman A (2020) Multiple resistance mechanisms in Staphylococcus sp. strain AS6 under arsenite stress and its potential use in amelioration of wastewater. J King Saud Univ Sci 32(7):3052–3058. https://doi.org/10.1016/j.jksus.2020.08.012
Mujawar SY, Shamim K, Vaigankar DC, Dubey SK (2019) Arsenite biotransformation and bioaccumulation by Klebsiella pneumoniae strain SSSW7 possessing arsenite oxidase (aioA) gene. Biometals 32:65–76. https://doi.org/10.1007/s10534-018-0158-7
Li X, Krumholz LR (2007) Regulation of arsenate resistance in Desulfovibrio desulfuricans G20 by an arsRBCC operon and an arsC gene. J Bacteriol 189:3705–3711. https://doi.org/10.1128/jb.01913-06
Banerjee S, Datta S, Chattyopadhyay D, Sarkar P (2011) Arsenic accumulating and transforming bacteria isolated from contaminated soil for potential use in bioremediation. J Environ Sci Health A Tox Hazard Subst Environ Eng 46:1736–1747. https://doi.org/10.1080/10934529.2011.623995
Naher U, Rahman F, Islam SMM, Sarkar MI, Biswas J (2016) Isolation of arsenic oxidizing-reducing bacteria and reclamation of As (III) in in vitro condition. Bangladesh Rice J 19:94. https://doi.org/10.3329/brj.v19i2.28170
Shakoori FR, Tabassum S, Rehman A, Shakoori AR (2010) Isolation and characterization of Cr6+ reducing bacteria and their potential use in bioremediation of chromium containing wastewater. Pakistan J Zool 42(6):651–658
Butt AS, Rehman A (2011) Isolation of arsenite-oxidizing bacteria from industrial effluents and their potential use in wastewater treatment. World J Microbiol Biotechnol 27:2435–2441. https://doi.org/10.1007/s11274-011-0716-4
Abbas SZ, Riaz M, Ramzan N, Zahid MT, Shakoori FR, Rafatullah M (2014) Isolation and characterization of arsenic resistant bacteria from wastewater. Braz J Microbiol 45:1309–1315. https://doi.org/10.1590/s1517-83822014000400022
Daware V, Gade WN (2015) Mechanism of arsenic tolerance in Klebsiella pneumoniae (HQ857583). Ind J Sci Res 6:457–469. https://doi.org/10.21608/eajbsg.2015.16483
Titah HS, Abdullah SRS, Idris M, Anuar N et al (2018) Arsenic resistance and biosorption by isolated rhizobacteria from the roots of Ludwigia octovalvis. Int J Microbiol 1–10:3101498. https://doi.org/10.1155/2018/3101498
Ghosh PK, Maiti TK, Pramanik K, Ghosh SK, Mitra S, De TK (2018) The role of arsenic resistant Bacillus aryabhattai MCC3374 in promotion of rice seedlings growth and alleviation of arsenic phytotoxicity. Chemosphere 211:407–419
Anderson CR, Cook GM (2004) Isolation and characterization of arsenate-reducing bacteria from arsenic-contaminated sites in New Zealand. Curr Microbiol 48:341–347. https://doi.org/10.1007/s00284-003-4205-3
Baker-Austin C, Wright MS, Stepanauskas R, McArthur JV (2006) Co-selection of antibiotic and metal resistance. Trends Microbiol 14:176–182. https://doi.org/10.1016/j.tim.2006.02.006
Jiang H, Yu T, Yang Y, Yu S, Wu J, Lin R, Li Y, Fang J, Zhu C (2020) Co-occurrence of antibiotic and heavy metal resistance and sequence type diversity of vibrio parahaemolyticus isolated from Penaeus vannamei at freshwater farms, seawater farms, and markets in Zhejiang province, China. Front Microbiol 11(1294):1–14. https://doi.org/10.3389/fmicb.2020.01294
Seiler C, Berendonk TU (2012) Heavy metal driven co-selection of antibiotic resistance in soil and water bodies impacted by agriculture and aquaculture. Front Microbiol 3:399. https://doi.org/10.3389/fmicb.2012.00399
Pal C, Asiani K, Arya S, Rensing C, Stekel DJ, Larsson DGJ, Hobman JL (2017) Metal resistance and its association with antibiotic resistance. Adv Microb Physiol 70:261–313. https://doi.org/10.1016/bs.ampbs.2017.02.001
Zhang M, Wan K, Zeng J, Lin W, Ye C, Yu X (2020) Co-selection and stability of bacterial antibiotic resistance by arsenic pollution accidents in source water. Environ Int 135(105351):1–11. https://doi.org/10.1016/j.envint.2019.105351
Tewari S, Ramteke P, Tripathi M, Kumar S, Garg S (2013) Plasmid mediated transfer of antibiotic resistance and heavy metal tolerance in thermotolerant water borne coliforms. Afr J Microbiol Res 7:130–136. https://doi.org/10.5897/AJMR12.1563
Chen J, Rosen BP (2020) The arsenic methylation cycle: how microbial communities adapted methylarsenicals for use as weapons in the continuing war for dominance. Front Environ Sci 8(43):1–14. https://doi.org/10.3389/fenvs.2020.00043
Kaur S, Kamli MR, Ali A (2011) Role of arsenic and its resistance in nature. Can J Microbiol 57:769–774. https://doi.org/10.1139/w11-062
Dash B, Sahu N, Singh AK, Gupta SB, Soni R (2021) Arsenic efflux in Enterobacter cloacae RSN3 isolated from arsenic-rich soil. Folia Microbiol 66:189–196. https://doi.org/10.1007/s12223-020-00832-2
Chen J, Rosen BP (2014) Biosensors for inorganic and organic arsenicals. Biosensors 4:494–512. https://doi.org/10.3390/bios4040494
Acknowledgements
Author RS wants to acknowledge his university, i.e. Indira Gandhi Krishi Vishwavidyalaya, Raipur, India, for financial assistance. However, this research did not receive any specific grant from funding agencies in the public, commercial, or nonprofit sectors. We also acknowledge the Department of Soil Science and Agril. Chemistry, Indira Gandhi Krishi Vishwavidyalaya, Raipur, for providing instrumentation facilities during this research.
Author information
Authors and Affiliations
Contributions
The concept, design, data analyses, manuscript preparation, and revision were performed by RS and PK. The experiments, data curation, manuscript preparation, and validation were performed by RS, PK, BD, and SBG. The project supervision, final validation, fund management, and final manuscript revision were performed by RS, SBG, and TC. All authors read and approved the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declared no conflict of interest regarding the publication of this manuscript.
Ethical Approval
This study does not describe any experimental work related to human.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
284_2021_2602_MOESM1_ESM.jpg
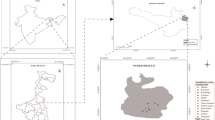
Supplementary file1 Pictorial view of sample collection sites in Industrial and Mining affected area of Chhattisgarh (India). (JPG 109 kb)
284_2021_2602_MOESM2_ESM.jpg
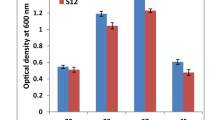
Supplementary file2 Comparative screening of strain RpSWA2 and RnASA11. The experiment was conducted using seven treatments with three replications of each. Treatment without metal salt was treated as a positive control whereas absolute control did not contain any metal salts and bacterial culture. ( JPG 34 kb)
Rights and permissions
About this article
Cite this article
Kumar, P., Dash, B., Suyal, D.C. et al. Characterization of Arsenic-Resistant Klebsiella pneumoniae RnASA11 from Contaminated Soil and Water Samples and Its Bioremediation Potential. Curr Microbiol 78, 3258–3267 (2021). https://doi.org/10.1007/s00284-021-02602-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-021-02602-w