Abstract
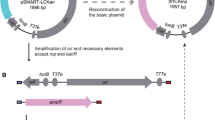
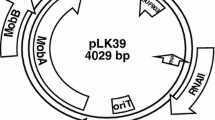
A high frequency of plasmids was detected in the rumen bacteriumSelenomonas ruminantium. Plasmids 0.9–20 kb in size were detected in more than 50% tested strains. Densitometric analysis indicated that plasmid DNA could represents more than 25% of total cellular DNA. Up to six plasmids were detected in strainS. ruminantium 18. Two smallest cryptic plasmids pSRD181 and pSRD182 from this strain were cloned intoEscherichia coli vector pBluescriptSK+ and partially characterized. The plasmid pSRD181 is 1.4 kb and pSRD182 is 2.0 kb. While computer analysis of pSRD181 sequence data showed high homology with replication protein ofStaphylococcus aureus plasmids, the pSRD182 sequence showed no significant homology in GenBank data. StrainS. ruminantium 28 was successfully transformed with pJW1 derived plasmid pJ1B1 using ampicillin resistance gene as marker. This is the first report on transformation of selenomonads with foreign DNA.
Similar content being viewed by others
References
Alexeyev M.F., Shokolenko I.N.: Mini-Tn10 transposon derivatives for insertion mutagenesis and gene delivery into the chromosome of Gram-negative bacteria.Gene160, 59–62 (1995).
Al-Khaldi S.F., Evans J.D., Martin S.A.: Complete nucleotide sequence of a cryptic plasmid from the ruminal bacteriumSelenomonas ruminantium HD4 and identification of two predicted open reading frames.Plasmid42, 45–52 (1999).
Antoine R., Locht C.: Isolation and molecular characterization of a novel broad-host-range plasmid fromBordetella bronchiseptica with sequence similarities to plasmid from gram-positive organisms.Mol. Microbiol.6, 1785–1799 (1992).
Attwood G.T., Brooker J.D.: Complete nucleotide sequence of aSelenomonas ruminantium plasmid and definition of a region necessary for its replication inEscherichia coli.Plasmid28, 123–129 (1992).
Birnboim H.C., Doly J.: A rapid alkaline lysis procedure for screening recombinant plasmid DNA.Nucl. Acids. Res.7, 1515–1523 (1979).
Bojovic B., Djordjevic G., Topisirovic L.: Improved vector for promoter screening in lactococci.Appl. Environ. Microbiol.2, 385–388 (1991).
Cocconcelli P.S., Elli M., Riboli B., Morelli L.: Genetic analysis of the replication region of theLactobacillus plasmid vector pPSC22.Res. Microbiol.8, 619–624 (1996).
Fliegerová K., Pažoutová S., Benada O., Flint H.J.: Small plasmids from rumen strains ofSelenomonas ruminantium.Proc. Nutr. Physiol.3, 149–150 (1994).
Fliegerová K., Pažoutová S., Pristaš P., Flint H.J.: Highly conserved DNA sequence present in small plasmids fromSelenomonas ruminantium.Plasmid44, 94–99 (2000).
Martin S.A., Dean R.G.: Characterization of a plasmid from ruminal bacteriaSelenomonas ruminantim.Appl. Environ. Microbiol.55, 3035–3038 (1989).
Nakamura M., Nagamine T., Ogata K., Tajima K., Aminov R.I., Benno Y.: Sequence analysis of small cryptic plasmids isolated fromSelenomonas ruminantium S20.Curr. Microbiol.38, 107–112 (1999).
Pristaš P., Molnárová V., Javorský P.: Restriction and modification systems of ruminal bacteria.Folia Microbiol.46, 71–72 (2001).
Pristaš P., Vanat I., Javorský P.: Variability of endonucleolytic activity indicates high genetic diversity among natural population ofSelenomonas ruminantium.Folia Microbiol.42, 121–124 (1997).
Tiwari A.D., Bryant M.P., Wolfe S.R.: Simple method for isolation ofSelenomonas ruminantium and some nutritional characteristics of the species.J. Dairy Sci.52, 2054–2056 (1969).
Zhang N., Brooker J.D.: Characterization, sequence, and replication of a small cryptic plasmid fromSelenomonas ruminantium subsp.lactilytica.Plasmid29, 125–134 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Hermanová, A., Pristaš, P., Molnárová, V. et al. Plasmids ofSelenomonas ruminantium and development of host-vector system. Folia Microbiol 46, 289–291 (2001). https://doi.org/10.1007/BF02815615
Received:
Issue Date:
DOI: https://doi.org/10.1007/BF02815615