Abstract
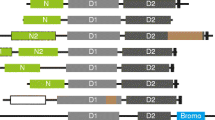
Members of the highly conserved and ubiquitous 14-3-3 protein family modulate a wide variety of cellular processes. To determine the evolutionary relationships among specific 14-3-3 proteins in different plant, animal, and fungal species and to initiate a predictive analysis of isoform-specific differences in light of the latest functional and structural studies of 14-3-3, multiple alignments were constructed from forty-six 14-3-3 sequences retrieved from the GenBank and SwissProt databases and a newly identified second 14-3-3 gene fromCaenorhabditis elegans. The alignment revealed five highly conserved sequence blocks. Blocks 2–5 correlate well with the alpha helices 3, 5, 7, and 9 which form the proposed internal binding domain in the three-dimensional structure model of the functioning dimer. Amino acid differences within the functional and structural domains of plant and animal 14-3-3 proteins were identified which may account for functional diversity amongst isoforms. Protein phylogenic trees were constructed using both the maximum parsimony and neighbor joining methods of the PHYLIP(3.5c) package; 14-3-3 proteins fromEntamoeba histolytica, an amitochondrial protozoa, were employed as an outgroup in our analysis. Epsilon isoforms from the animal lineage form a distinct grouping in both trees, which suggests an early divergence from the other animal isoforms. Epsilons were found to be more similar to yeast and plant isoforms than other animal isoforms at numerous amino acid positions, and thus epsilon may have retained functional characteristics of the ancestral protein. The known invertebrate proteins group with the nonepsilon mammalian isoforms. Most of the current 14-3-3 isoform diversity probably arose through independent duplication events after the divergence of the major eukaryotic kingdoms. Divergence of the seven mammalian isoforms beta, zeta, gamma, eta, epsilon, tau, and sigma (stratifin/ HME1) occurred before the divergence of mammalian and perhaps before the divergence of vertebrate species. A possible ancestral 14-3-3 sequence is proposed.
Similar content being viewed by others
References
Alam R, Hachiya N, Sakaguchi M, Kawabata S, Iwanaga S, Kitajima M, Mihara K, Omura T (1994) cDNA cloning and characterization of mitochondrial import stimulation factor (MSF) purified from rat liver cytosol. J Biochem (Tokyo) 1116(2): 416–425
Aitken A (1995) 14-3-3 proteins on the MAP. Trends Biochem Sci 20(3): 95–97
Aitken A, Collinge DB, van Heusden PH, Isobe T, Roseboom PH, Rosenfeld G, Soll J (1992) 14-3-3 proteins: a highly conserved widespread family of eukaryotc proteins. Trends Biochem Sci 17: 498–501
Aitken A, Howell S, Jones D, Madrazo J, Patel Y (1995) 14-3-3 alpha and delta are the phosphorylated forms of raf-activating 14-3-3 beta and zeta.In vivo stoichiometric phosphorylation in brain at a Ser-Pro-Glu-Lys motif. J Biol Chem 17, 270(11): 5706–5709
Bairoch A, Bucher P (1994) Prosite: recent developments. Nucleic Acids Res 22: 3583–3589
Baldauf SL, Palmer JD (1993) Animals and fungi are each other's closest relatives: congruent evidence from multiple proteins. PNAS 90: 11558–11562
Brandt J, Thordalchristensen H, Vad K, Gregersen PL, Collinge DB (1992) A pathogen-induced gene of barley encodes a protein showing high similarity to a protein-kinase regulator. Plant J 2 (5):815–820
Burbelo PD, Hall A (1995) 14-3-3 proteins. Hot numbers in signal transduction. Curr Biol 5 (2): 95–96
Cavalier-Smith T, Allsopp MTEP, Chao EE (1994) Thraustochytrids are chromists, not Fungi: 18s rRNA signatures of Heterokonta. Philos Trans R Soc Lond Biol 346: 387–397
Conklin DS, Galaktionov K, Beach D (1995) 14-3-3 proteins associate with cdc25 phosphatases. PNAS 92: 7892–7896
deVetten NC, Lu GH, Ferl RJ (1992) A maize pattern associated with the G-box binding complex has homology to brain regulatory proteins. Plant Cell 4 (10): 1295–1307
De Rijk P, Van de Peer Y, Van den Broeck I, De Wachter R (1995) Evolution according to large ribosomal subunit RNA. J Mol Evol 41: 366–375
Doolittle RF (1993) Convergent evolution: the need to be concise. Trends Biochem Sci 19: 15–18
Fantl WJ, Muslin AJ, Kikuchi A, Martin JA, MacNicol AM, Gross R, Williams LT (1994) Activation of Raf-1 by 14-3-3 proteins. Nature 13, 371 (6498): 612–614
Felsenstein J (1993) PHYLIP(phylogeny inference package) version 3.5c. Distributed by the author. Department of Genetics, University of Washington, Seattle
Ferl RJ, Lu G, Bowen BW (1994) Evolutionary implications of the family of 14-3-3 brain protein homologs inArabidopsis thaliana. Genetica 92(2): 129–138
Ford JC, al-Khodairy F, Fotou E, Sheldrick KS, Griffiths DJ, Carr AM (1994) 14-3-3 protein homologs required for the DNA damage checkpoint in fission yeast. Science 265: 533–535
Freed E, Symons M, Macdonald SG, McCormick F, Ruggieri R (1994) Binding of 14-3-3 proteins to the protein kinase Raf and effects on its activation. Science 265: 1713–1716
Fu H, Coburn J, Collier RJ (1993) The eukaryotic host factor that activates exoenzyme S ofPseudomonas aeruginosa is a member of the 14-3-3 protein family. PNAS 90: 2320–2324
Fu H, Xia K, Pallas DC, Cui C, Conroy K, Narsimhan RP, Mamon H, Collier RJ, Roberts TM (1994) Interaction of the protein kinase Raf-1 with 14-3-3 proteins. Science 266: 126–129
Hasegawa M, Hashimoto T, Adachi J, Iwabe N, Miyata T (1993) Early branchings in the evolution of eukaryotes: ancient divergence of Entamoeba that lacks mitochondria revealed by protein sequence data. J Mol Evol 36: 380–388
Hinkle G, Leipe DD, Nerad TA, Sogin ML (1994) The unusually long small subunit ribosomal RNA ofPhreatamoeba balamuthi. Nucleic Acids Res 22: 465–469
Hirsch S, Aitken A, Bertsch U, Soll J (1992) A plant homologue to mammalian brain 14-3-3 protein and protein kinase C inhibitor. FEBS Lett 296: 222–224
Ichimura T, Isobe T, Okuyama T, Takahashi N, Araki K, Kuwano R, Takahashi Y (1988) Molecular cloning of cDNA coding for brain-specific 14-3-3 protein, a protein kinase-dependent activator of tyrosine and tryptophan hydroxylases. PNAS 85 (19): 7084–7087
Ischimura T, Sugano H, Kuwano R, Sunaya T, Okuyama T, Isobe T (1991) Widespread distribution of the 14-3-3 protein in vertebrate brains and bovine tissues: correlation with the distribution of calcium-dependent protein kinases. J Neurochem 56: 1449–1451
Ichimura-Ohshima Y, Mori K, Ichimura T, Araki K, Takahashi Y, Isobe T, Minoshima S, Fukuyama R, Shimizu N, Kuwano R (1992) cDNA cloning and chromosome assignment of the gene for human brain 14-3-3 protein eta-chain. J Neurosci Res 31: 600–605
Irie K, Gotoh Y, Yashar BM, Errede B, Nishida E, Matsumoto K (1994) Stimulatory effects of yeast and mammalian 14-3-3 proteins on the Raf protein kinase. Science 265: 1716–1719
Isobe T, Ichimura T, Sunaya T, Okuyama T, Takahashi N, Kuwano R, Rosenfeld G, Takahashi Y (1991) Distinct forms of the protein kinase dependent activator of tyrosine and tryptophan hydroxylases. J Mol Biol 217: 125–132
Jarillo JA, Capel J, Leyva A, Martinez-Zapater JM, Salinas EBJ (1994) Two related low-temperature-inducible genes of arabidopsis encode proteins showing high homology to 14-3-3 proteins, a family of putative kinase regulators. Plant Mol Biol 25 (4): 693–704
Jones DH, Ley S, Aitken A (1995a) Isoforms of 14-3-3 protein can form homo- and heterodimers in vivo and in vitro: implications for function as adapter proteins. FEBS Lett 368: 55–58
Jones DH, Martin H, Madrazo J, Robinson KA, Neilsen P, Roseboom PH, Patel Y, Howell SA, Aitken A (1995b) Expression and structural analysis of 14-3-3 proteins. J Mol Biol 245: 375–384
Korthout HA, deBoer AH (1994) A fusicoccin binding protein belongs to the family of 14-3-3 brain protein homologs. Plant Cell 6: 1681–1692.
Kidou S, Umeda M, Kato A, Uchimiya H (1993) Isolation and characterization of a rice cDNA similar to the bovine brain-specific 14-3-3 protein gene. Plant Mol Biol 21: 191–194
Laughner B, Lawrence SD, Ferl RJ (1994) Two tomato fruit homologs of 14-3-3 mammalian brain proteins. Plant Physiol 105 (4): 1457–1458
Leffers H, Madsen P, Rasmussen HH, Honore B, Andersen AH, Walbum E, Vandekerckhove J, Celis JE (1993) Molecular cloning and expression of the transformation sensitive epithelial with the G-box marker stratifin. A member of a protein family involved in the protein kinase C signalling pathway. J Mol Biol 231: 982–998
Liu D, Bienkowska J, Petosa C, Collier RJ, Fu H, Liddington R (1995) Crystal structure of the zeta isoform of the 14-3-3 protein. Nature 376: 191–194
Lu GH, Delisle AJ, deVetten NC, Ferl RJ (1992) Brain proteins in plants—an Arabidopsis homolog to neurotransmitter pathway activators is part of a DNA binding complex. PNAS 89 (23): 1490–1494
Heusden GP, Ferl RJ (1994a) A single Arabidopsis GF14 isoform possesses biochemical characteristics of diverse 14-3-3 homologues. Plant Mol Biol 25 (4): 659–673
Lu G, Sehnke PC, Ferl RJ (1994b) Phosphorylation and calcium binding properties of an Arabidopsis GF14 brain protein homolog. Plant Cell 6 (4): 501–510
Markiewicz E, Rzepecki R, Zopa S (1994) Molecular cloning and sequencing of the cDNA sequence of the cDNA encoding plant nuclear matrix endonuclease. Acta Biochim Pol 41: 137–138
Martens GJM, Piosik PA, Danen EHJ (1992) Evolutionary conservation of the 14-3-3 protein. Biochim Biophys Res Commun 184: 1456–1459
Martin H, Patel Y, Jones D, Howell S, Robinson K, Aitken A (1993) Antibodies against the major brain isoforms of 14-3-3 protein: an antibody specitic for the N-acetylated amino-terminus of a protein. FEBS Lett 331: 296–303
McConnell JE, Armstrong JF, Hodges PE, Bard JB (1995) The mouse 14-3-3 epsilon isoform, a kinase regulator whose expression pattern is modulated in mesenchyme and neuronal differentiation. Dev Biol 169 (1): 218–228
Moore BW, Perez VJ (1967) Specific acidic proteins of the nervous system. In: Carlson FD (ed) Physiological and biochemical aspects of nervous integration. Prentice-Hall, Englewood Cliffs, NJ, pp 343–359
Morgan A, Burgoyne RD (1992) Interaction between protein kinase C and Exo 1 (14-3-3 protein) and its relevance to exocytosis in permeabilized adrenal chromaffin cells. Biochem J 286: 807–811
Morrison D (1994) 14-3-3 modulators of signaling proteins? Science 266: 56–57
Nielsen PJ (1991) Primary structure of a human protein kinase regulator protein. Biochim Biophys Acta 1088: 425–428
Oecking C, Eckerskorn C, Weiler EW (1994) The fusicoccin receptor of plants is a member of the 14-3-3 superfamily of eukaryotic regulatory proteins. FEBS Lett 352: 163–166
Pallas DC, Fu H, Haehnel LC, Weller W, Collier RJ, Roberts TM (1994) Association of polyomavirus middle tumor antigen with 14-3-3 proteins. Science 265: 535–537
Prasad GL, Valverius EM, McDuffie E, Cooper HL (1992) Complementary DNA cloning of a novel epithelial cell marker protein, HME1, that may be down-regulated in neoplastic mammary cells. Cell Growth Differ 3 (8): 507–513
Reuther GW, Fu H, Cripe LD, Collier RJ, Pendergast AM (1994) Association of the protein kinase c-Bcr and Bcr-Abl with proteins of the 14-3-3 family. Science 266: 129–133
Roseboom PH, Weller JL, Babila T, Aitken A, Sellers LA, Moffett JR, Namboodiri MA, Klein DC (1994) Cloning and characterization of the epsilon and zeta isoforms of the 14-3-3 proteins. DNA Cell Biol 13 (6): 629–640
Roth D, Morgan A, Martin H, Jones D, Martens GJ, Aitken A, Burgoyne RD (1994) Characterization of 14-3-3 proteins in adrenal chromaffin cells and demonstration of isoform-specific phospholipid binding. Biochem J 301: 305–310
Stankovic B, Garic-Stankovic A, Smith CM, Davies E (1995) Isolation, sequencing, and analysis of a 14-3-3 brain protein homolog from pea (Pisum sativum L.). Plant Physiol 107 (4): 1481–1482
Swanson KD, Ganguly R (1992) Characterization of aDrosophila melanogaster gene similar to the mammalian genes encoding the tyrosine/tryptophan hydroxylase activator and protein kinase C inhibitor proteins. Gene 113: 183–190
Tanji M, Horwitz R, Rosenfeld G, Waymire JC (1994) Activation of protein kinase C by purified bovine brain 14-3-3: comparison with tyrosine hydroxylase activation. J Neurochem 63 (5): 1908–1916
Toker A, Sellers LA, Amess B, Patel Y, Harris A, Aitken A (1992) Multiple isoforms of a protein kinase C inhibitor (KCIP/14-3-3) from sheep brain: amino acid sequence of phosphorylated forms. Eur J Biochem 206: 453–461
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position specific gap penalties and weight matrix choices. Nucleic Acids Res 22: 4673–4680
van Heusden GPH, Wenzel TJ, Lagendijk EL, de Steensma HY, van den Berg JA (1992) Characterization of the yeast BMH1 gene encoding a putative protein homologous to mammalian protein kinase II activators and proteins kinase C inhibitors. FEBS Lett 302 (2): 145–150
van Heusden GPH, Griffiths DJ, Ford JC, Chin A, Woeng TF, Schrader PA, Carr AM, Steensma HY (1995) The 14-3-3 proteins encoded by the BMH1 and BMH2 genes are essential in the yeastSaccharomyces cerevisiae and can be replaced by a plant homologue. Eur J Biochem 229 (1): 45–53
Wang W, Shakes DC (1994) Isolation and sequence analysis of aCaenorhabditis elegans cDNA which encodes a 14-3-3 homologue. Gene 147: 215–218
Watanabe M, Isobe T, Okuyama T, Ichimura T, Kuwano R, Takahashi Y, Kondo H (1991) Molecular cloning of cDNA to rat 14-3-3 eta chain polypeptide and the neuronal expression of the mRNA in the central nervous system. Mol Brain Res 10 (2): 151–158
Watanabe M, Isobe T, Ichimura T, Kuwano R, Takahashi Y, Kondo H (1993a) Molecular cloning of rat cDNAs for beta and gamma subtypes of 14-3-3 protein and developmental change in expression of their mRNAs in the nervous system. Mol Brain Res 17 (1–2): 135–146
Watanabe M, Isobe T, Ichimura, Kuwano R, Takahashi Y, Kondo H (1993b) Developmental regulation of neuronal expression for the eta subtype of the 14-3-3 protein, a putative regulatory protein for protein kinase C. Dev Brain Res 73: 225–235
Watanabe M, Isobe T, Ichimura T, Kuwano R, Takahashi Y, Kondo H, Inoue Y (1994) Molecular cloning of rat cDNAs for the zeta and theta subtypes of 14-3-3 protein and differential distributions of their mRNAs in brain. Mol Brain Res 25 (1–2): 113–121
Xiao B, Smerdon SJ, Johns DH, Dodson GG, Soneji Y, Aitken A, Gamblin SJ (1995) Structure of a 14-3-3 protein and implications for coordination of multiple signalling pathways. Nature 376: 188–191
Zupan LA, Steffens DL, Berry CA, Landt M, Gross RW (1992) Cloning and expression of a human 14-3-3 protein mediating phospholipolysis. J Biol Chem 13: 8707–8710
Author information
Authors and Affiliations
Additional information
Correspondence to: D.C. Shakes
Rights and permissions
About this article
Cite this article
Wang, W., Shakes, D.C. Molecular evolution of the 14-3-3 protein family. J Mol Evol 43, 384–398 (1996). https://doi.org/10.1007/BF02339012
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF02339012