Summary
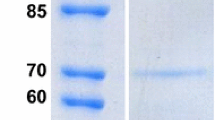
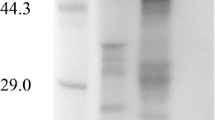
Endo-β-1,4-glucanase genemcenA ofMicromonospora cellulolyticum 86W-16 was cloned, and the nucleotide sequence was determined. An open reading frame (ORF) of 1374 bases, coding for a peptide (McenA) of 457 amino acids and 46742 Da, was found. It is preceded by a Gram-positive type of ribosomebinding site and followed by an imperfect inverted repeat. A putative signal peptide containing 23 amino acids is at the N-terminus and a linker region possessing 37 amino acids is in the midpart of McenA. The N-half of McenA functions as the catalytic domain and the C-half might serve as a cellulosebinding domain (CBD). Deletion of the latter did not decrease the CMCase activity of McenA. Significant similarity (70%) was found between the amino acid sequences of McenA and MbcelA, an endoglucanase fromMicrobispora bispora.
Similar content being viewed by others
References
Cheng, Y.R., F. Lin, Z. Zhang and B.S. Yu. 1988. Utilization of cellulose byMicromonospora spp I. A. newly isolatedMicromonospora spp. no. 86W-6. In: Recent Advances in Biotechnology and Applied Biology (Chang, S.T., K.Y. Chan and N.Y.S. Woo, eds), pp. 451–460. Proceedings of 8th International Conference on Global Impacts of Applied Microbiology, August 1–5, 1988, Hong Kong.
Chung, C.T., S.L. Niemala and R.H. Miller. 1989. One-step preparation of competentEscherichia coli: transformation and storage of bacterial cells in the same solution. Proc. Natl. Acad. Sci. USA 86: 2172–2175.
Coutinho, J.B., B. Moser, D.G. Kilburn, R.A.J. Warren and R.C. Miller, Jr. 1991. Nucleotide sequence of the endoglucanase C gene (cenC) ofCellulomonas fimi, its high-level expression inEscherichia coli, and characterization of its products. Mol. Microbiol. 5: 1221–1233.
Eveleigh, D.E. 1991. Personal communication, Rutgers University, New Brunswick, NJ, USA.
Foong, F. and R.H. Doi. 1992. Characterization and comparison ofClostridium cellulovorans endoglucanase-xylanases EngB and EngD hyperexpressed inEscherichia coli. J. Bacteriol. 174: 1403–1409.
Gilkes N.R., D.G. Kilburn, R. C. Miller, Jr. and R.A.J. Warren. 1991. Bacterial cellulases. Bioresource Technol. 36: 21–35.
Gilkes N.R., B. Henrissat, D.G. Kilburn, R.C. Miller, Jr and R.A.J. Warren. 1991. Domains in microbial β-1,4-glycanase: sequence conservation, function, and enzyme families. Microbiol. Rev. 55: 303–315.
Grépinet, O., M.C. Chebrou and P. Béguin. 1988. Nucleotide sequence and deletion analysis of the xylanase gene (xynZ) ofClostridium thermocellum. J. Bacteriol. 170: 4582–4588.
Hamamoto, T., F. Foong, O. Shoseyov and R.H. Doi. 1992. Analysis of functional domains of endoglucanases fromClostridium cellulovorans by gene cloning, nucleotide sequencing and chimeric protein construction. Mol. Gen. Genet. 231: 472–479.
Hopwood, D.A., M.J. Bibb, K.F. Chater, T. Kieser, C.J. Bruton, H.M. Kieser, D.J. Lydiate, C.P. Smith, J.M. Ward and H. Schrempf. 1985. Isolation ofStreptomyces ‘total’ DNA: procedure 1. In: Genetic Manipulation ofStreptomyces, A Laboratory Manual (Joklik, W.K. ed.), pp. 72–74. The John Innes Foundation, Norwich.
Hu, Y.-J. and D.B. Wilson. 1988. Cloning ofThermomonospora fusca genes coding for beta 1–4 endoglucanases E1, E2 and E5. Gene 71: 331–337.
Kalakoutskii, L. V., N.S. Agre, H. Prauser and L.I. Evtushenko. 1986. GenusPromicromonospora Krasil'nikov, Kalakoutskii and Kirillova 1961, 107AL. In: Bergey's Manual of Systematic Bacteriology, Vol. 2 (Sneath, P.H.A., ed.), pp. 1501–1503, Williams & Wilkins Press, Baltimore.
Kreil, G. 1981. Transfer of proteins across membranes. Annu. Rev. Biochem. 50: 317–348.
Luedemann, G. M. 1974. Genus I.Micromonospora Ørskov 1923, 147 In: Bergey's Manual of Determinative Bacteriology, 8th edn (Buchanan R.E., ed.), pp. 846–855, The Williams & Wilkins Company, Baltimore.
Maniatis, T., E.F. Fritsch and J. Sambrook. 1982. Molecular Cloning, A Laboratory Manual, Cold Spring Harbor Laboratory, Cold Spring Harbor, NY.
Meinke, A., N.R. Gilkes, D.G. Kilburn, R.C. Miller, Jr and R.A.J. Warren. 1991. Multiple domains in endoglucanase D (CenB) fromCellulomonas fimi: functions and relatedness to domains in other polypeptides. J. Bacteriol 173:7126–7135.
Nakai, R., S. Horinouchi and T. Beppu. 1988. Cloning and nucleotide sequence of a cellulase gene,casA, from an alkalophilicStreptomyces strain. Gene 65: 229–238.
Neuwald, A.F. and G.V. Stauffer. 1989. AnEscherichia coli membrane protein with a unique signal sequence. Gene 82: 219–228.
Ozaki, K., S. Shikata, S. Kawai, S., Ito and K. Okamoto. 1990. Molecular cloning and nucleotide sequence of a gene for alkaline cellulase fromBacillus sp. KSM-635. J. Gen. Microbiol. 136: 1327–1334.
Presutti, D.G., T.A. Hughes and F.J. Stutzenberger. 1993. Characterization of aThermomonospora curvata endoglucanase expressed inEscherichia coli. J. Biotechnol. 29: 307–320.
Rosenberg, M. and D. Court. 1974. Regulatory sequences involved in the promotion and termination of RNA transcription. Ann. Rev. Genet. 13: 319–353.
Sanger, F., A.R. Coulson, B.G. Barrell, A.J.H. Smith and B.A. Roe. 1980. Cloning in single-stranded bacteriophage as an aid to rapid DNA sequencing. J. Mol. Biol. 143: 161–178.
Staden, R. 1980. A new computer method for the storage and manipulation of DNA gel reading data. Nucleic Acids Res. 16: 3673–3694.
Teather, R.M. and P.J. Wood. 1982. Use of Congo Red-polysaccharide interactions in enumeration and characterization of cellulolytic bacteria from the bovine rumen. Appl. Environ. Microbiol. 43: 777–780.
Wilson D.B. 1992. Biochemistry and genetics of actionmycete cellulases. Crit. Rev. Biotechnol. 12: 45–63.
Yablonsky, M.D., T. Bartley, K.O. Elliston, S.K. Kahrs, Z.P. Shalita and D.E. Eveleigh. 1988. Characterization and cloning of the cellulase complex ofMicrobispora bispora. In: Biochemistry and Genetics of Cellulose Degradation. FEMS Symposium 43 (J.P. Aubert, P. Bégiun and J. Millet, eds), pp. 249–266, Academic Press, London.
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Lin, F., Marchenko, G. & Cheng, YR. Cloning and sequencing of an endo-β-1,4-glucanase genemcenA fromMicromonospora cellulolyticum 86W-16. Journal of Industrial Microbiology 13, 344–350 (1994). https://doi.org/10.1007/BF01577217
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01577217