Abstract
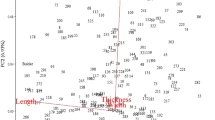
Quantitative trait locus (QTL) and QTL x environment (E) interaction effects for agronomic and malting quality traits were measured using a 123-point linkage map and multi-environment phenotype data from an F1-derived doubled haploid population of barley (Hordeum vulgare). The QTL × E interactions were due to differences in magnitude of QTL effects. Highly significant QTL effects were found for all traits at multiple sites in the genome. Yield QTL peaks and support intervals often coincided with plant height and lodging QTL peaks and support intervals. QTL were detected in the vicinity of a previously mapped Mendelian maturity locus and known function probes forα- andβ-amylase genes. The average map density (9.6 cM) should be adequate for molecular marker-assisted selection, particularly since there were few cases of alternative favorable alleles for different traits mapping to the same or adjacent intervals.
Similar content being viewed by others
References
Allard RW (1988) Genetic changes associated with the evolution of adaptedness in cultivated plants and their wild progenitors. J Hered 79:225–238
Beavis WD, Grant D, Albertsen M, Fincher R (1991) Quantitative trait loci for plant height in four maize populations and their associations with qualitative genetic loci. Theor Appl Genet 83:141–145
Chen F, Hayes PM (1989) A comparison ofHordeum bulbosum-mediated haploid production efficiency in barley using in vitro floret and tiller culture. Theor Appl Genet 77:701–704
Haley CS, Knott (1992) A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69:315–324
Hayes PM, Blake TK, Chen THH, Tragoonrung S, Chen F, Pan A, Liu B (1993) Quantitative trait loci on barley (Hordeum vulgare) chromosome 7 associated with components of winterhardiness. Genome 36:66–71
Heun M (1992) Mapping quantitative powdery mildew resistance of barley using a restriction fragment length polymorphism map. Genome 35:1019–1025
Hockett E, Nilan R (1985) Genetics. In: Rasmusson DC (ed) Barley. Am Soc Agron Madison Wis., pp 190–216
Khursheed B, Rogers JC (1988) Barleyα-amylase genes: quantitative comparison of steady-state mRNA levels from individual members of two different families expressed in aleurone cells. J Bio Chem 263:18953–18960
Kleinhofs A, Kilian A, Saghai Maroof MA, Biyashev RM, Hayes PM, Chen F, Lapitan N, Fenwick A, Blake TK, Kanazin V, Ananiev E, Dahleen L, Kudrna D, Bollinger J, Knapp SJ, Liu B, Sorrells M, Heun M, Franckowiak JD, Hoffman D, Skadsen R, Steffenson BJ (1993) A molecular, isozyme, and morphological map of the barley (Hordeum vulgare) genome Theor Appl Genet (in press)
Knapp SJ (1989) Quasi-Mendelian genetic analysis of quantitative trait loci using molecular marker linkage maps. In: Roebbelen G (ed) Proc 12th Eucarpia Congr. Feb. 27–March 4. Goettingen, Germany, pp 51–67
Knapp SJ, Bridges WC (1990) Programs for mapping quantitative trait loci using flanking molecular markers. J Hered 81:234–235
Knapp SJ, Bridges WC, Birkes D (1990) Mapping quantitative trait loci using molecular marker linkage maps. Theor Appl Genet 79:583–592
Kosambi DD (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Kreis M, Williamson M, Buxton B, Pywell J, Hejgaard J, Svendsen I (1987) Primary structure and differential expression ofβ-amylase in normal and mutant barleys. Eur J Biochem 169:517–525
Liu BH, Knapp SJ (1990) GMENDEL: a program for Mendelian segregation and linkage analysis of individual or multiple progeny using log-likelihood ratios. J Hered 81:407
Nilan RA (1964) The cytology and genetics of barley. Monogr Suppl No. 332:1. Washington State Univ Press, Pullman, Wash.
Paterson A, Tanksley SD, Sorrells ME (1991a) DNA markers in plant improvement. In: Adv Agron 46:39–90
Paterson A, Damon S, Hewitt JD, Zamir D, Rabinowitch HD, Lincoln S, Lander ES, Tanksley SD (1991b) Mendelian factors underlying quantitative traits in tomato: comparison across species, generations, and environments. Genetics 127:181–197
Shewry PR, Faulks AJ, Pickering RA, Jones IT, Finch RA, Miflin BJ (1980) The genetic analysis of barley storage proteins. Heredity 44:383–389
van Ooijen JW (1992) Accuracy of mapping quantitative trait loci in autogamous species. Theor Appl Genet 84:803–811
Young ND, Tanksley SD (1989) RFLP analysis of the size of chromosomal segments retained around theTm-2 locus of tomato during backcross breeding. Theor Appl Genet 77:95–101
Author information
Authors and Affiliations
Additional information
Communicated by G. Wenzel
Oreg Agric Exp Stn J No. 10150
Rights and permissions
About this article
Cite this article
Hayes, P.M., Liu, B.H., Knapp, S.J. et al. Quantitative trait locus effects and environmental interaction in a sample of North American barley germ plasm. Theoret. Appl. Genetics 87, 392–401 (1993). https://doi.org/10.1007/BF01184929
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF01184929