Abstract
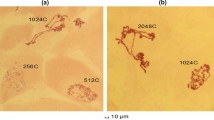
By combining information from microscopical observation and photography a graphical map ofDrosophila virilis salivary gland chromosomes was constructed. About 1,560 individual bands are shown and patterns of transcription at about 360 sites are indicated. The application of the map is demonstrated by using genetic, morphological and in situ hybridization data to identify thewhite-Notch regions ofD. virilis andDrosophila melanogaster as homologous chromosome segments with constant and variable features.
Similar content being viewed by others
References
Aimanova KG, Pereligina LM, Kolesnikov NN (1987a) A biochemical analysis of the salivary gland secretory glycoproteins ofDrosophila virilis. Dros Inf Serv 66:1–3
Aimanova KG, Pereligina LM, Kolesnikov NN (1987b) A genetic analysis of the tissue-specific salivary gland secretion proteins ofD. virilis. Dros Inf Serv 66:3–5
Alexander ML (1976) The genetics ofDrosophila virilis. In: Ashburner M, Novitzki E (eds) The genetics and biology ofDrosophila, vol 1c. Academic Press, New York, pp 1365–1427
Ashburner M (1989)Drosophila. A laboratory handbook. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, NY, p 51
Blackman RK, Meselson M (1986) Interspecific nucleotide sequence comparisons used to identify regulatory and structural features of theDrosophila hsp 82 gene. J Mol Biol 188:499–515
Chen C, Malone T, Beckendorf SK, Davis RL (1987) At least two genes reside within a large intron of the dunce gene ofDrosophila. Nature 329:721–724
Colot HV, Hall JC, Rosbash M (1988) Interspecific comparison of the period gene ofDrosophila reveals large blocks of nonconserved coding DNA. EMBO J 7:3929–3937
Evgen'ev MB (1977) New method of cytological localization of genes in “virilis” group ofDrosophila. Dros Inf Serv 52:59–60
Fujii S (1936) Salivary gland chromosomes ofDrosophila virilis. Cytologia 7:272–275
Fujii S (1942) Further studies on the salivary gland chromosomes ofDrosophila virilis. Cytologia 12:435–455
Gall JG, Atherton DD (1974) Satellite DNA sequences inDrosophila virilis. J Mol Biol 85:633–664
Gubenko IS, Evgen'ev MB (1984) Cytological and linkage maps ofDrosophila virilis chromosomes. Genetica 65:127–139
Hooper JE, Perez-Alonso M, Bermingham JR, Prout MR, Rocklein BA, Wagenbach M, Edström J-E, de Frutos R, Scott MP (1992) Comparative studies ofDrosophila Antennapedia genes. Genetics 132:453–469
Hsu TC (1952) Chromosomal variation and evolution in thevirilis group ofDrosophila. Univ Texas Publ 504:35–56
Hughes RD (1939) An analysis of the chromosomes of the two subspeciesDrosophila virilis virilis andDrosophila virilis americana. Genetics 24:811–834
Jones CW, Dalton MW, Townley LH (1991) Interspecific comparison of the structure and regulation of theDrosophila ecdysone-inducible gene E74. Genetics 127:535–543
Kalisch WE, Schwitalla G, Whitmore T (1986) Electron microscopic band-interband pattern of the X chromosome inDrosophila hydei. Chromosoma 93:387–392
Kassis JA, Poole SJ, Wright DK, O'Farrell PH (1986) Sequence conservation in the protein coding and intron regions of the engrailed transcription unit. EMBO J 5:3583–3589
Kress H (1972) Das Puffmuster der Riesenchromosomen in den larvalen Speicheldrüsen vonDrosophila virilis: seine Veränderungen in der Normalentwicklung und nach Injektion von Ecdyson. Sitz Ber Bayer Akad Wiss Math Naturw Kl 129–149
Kress H (1979) Ecdysone-induced changes in glycoprotein synthesis and puff activities inDrosophila virilis salivary glands. Chromosoma 72:53–66
Kress H (1982) Biochemical and ontogenetic aspects of glycoprotein synthesis inDrosophila virilis salivary glands. Dev Biol 93:231–239
Kress H (1986) Stoffwechselaktivitäten und Transkriptionsmuster in den larvalen Speicheldrüsen vonDrosophila virilis. Naturwissenschaften 73:180–187
Kress H, Lucka L, Swida U, Thüroff E, Klemm U (1990) Genes from two intermoult puffs inDrosophila virilis polytene chromosomes are differentially transcribed during larval development. Development 108:261–267
Laird CD (1974) DNA ofDrosophila chromosomes. Annu Rev Gen 7:177–203
Lindsley DL, Zimm GG (1992) The genome ofDrosophila melanogaster. Academic Press, New York
Lozovskaya ER, Petrov DA, Hartl DL (1993) A combined molecular and cytogenetic approach to genome evolution inDrosophila using large-fragment DNA cloning. Chromosoma 102:253–266
Merriam J, Ashburner M, Hartl DL, Kafatos FC (1991) Toward cloning and mapping the genome ofDrosophila. Science 254:221–225
Mulder MP, Duijn P van, Gloor HJ (1968) The replicative organization of DNA in polytene chromosomes ofDrosophila hydei. Genetica 39:385–428
Neufeld TP, Carthew RW, Rubin GM (1991) Evolution of gene position: chromosomal arrangement and sequence comparison of theDrosophila virilis sina and Rh4 genes. Proc Natl Acad Sci USA 88:10203–10207
O'Neil MT, Belote JM (1992) Interspecific comparison of the transformer gene ofDrosophila reveals an unusually high degree of evolutionary divergence. Genetics 131:113–128
Patterson JT, Stone WS, Griffen AB (1940) Evolution of the virilis group inDrosophila. Univ Texas Publ 4032:218–250
Rudkin GT (1969) Non-replicating DNA inDrosophila. Genetics 61: [Suppl] 227–238
Rykowski MC, Parmelee SJ, Agard DA, Sedat JW (1988) Precise determination of the molecular limits of a polytene chromosome band: regulation sequences for the Notch gene are in the interband. Cell 54:461–472
Sorsa V (1988) Chromosome maps ofDrosophila, vol II. CRC Press, Boca Raton, Florida
Sturtevant AH, Novitzki E (1941) The homologies of the chromosome elements in the genusDrosophila. Genetics 26:517–541
Swida U, Lucka L, Kress H (1990) Glue protein genes inDrosophila virilis: their organization, developmental control of transcription and specific mRNA degradation. Development 108:269–280
Thüroff E, Stöven S, Kress H (1992)Drosophila salivary glands exhibit a regional reprogramming of gene expression during the third larval instar. Mech Dev 37:81–93
Treier T, Pfeifle C, Tautz D (1989) Comparison of the gap segmentation gene hunchback betweenDrosophila melanogaster andDrosophila virilis reveals novel modes of evolutionary change. EMBO J 8:1517–1525
Wainwright SM, Ish-Horowicz D (1992) Point mutations in theDrosophila hairy gene demonstrate in vivo requirements for basic, helix-loop-helix, and WRPW domains. Mol Cell Biol 12:2475–2483
Wallrath LL, Friedman TB (1991) Species differences in the temporal pattern ofDrosophila urate oxidase gene expression are attributed to trans-acting regulatory changes. Proc Natl Acad Sci USA 88:5489–5493
Wharton KA, Johansen KM, Xu T, Artavanis-Tsakonas S (1985) Nucleotide sequence from the neurogenic locus Notch implies a gene product that shares homology with proteins containing EGF-like repeats. Cell 43:567–581
Whiting JH, Pliley MD, Farmer JL, Jeffery DE (1989) In situ hybridization analysis of chromosomal homologies inDrosophila melanogaster andDrosophila virilis. Genetics 122:99–109
Author information
Authors and Affiliations
Additional information
Communicated by: E.R. Schmidt
Rights and permissions
About this article
Cite this article
Kress, H. The salivary gland chromosomes ofDrosophila virilis: a cytological map, pattern of transcription and aspects of chromosome evolution. Chromosoma 102, 734–742 (1993). https://doi.org/10.1007/BF00650901
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1007/BF00650901