Abstract
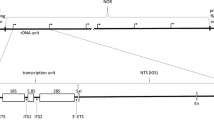
The organization of rat liver ribosomal DNA (rDNA) as matrix-attached DNA loops was examined using a protocol which fractionates chromatin from discrete regions of DNA loops. Southern blot analysis of matrixattached and solubilized chromatin DNA fragments demonstrated that rDNA is associated with the matrix via its 5′ and 3′ nontranscribed spacer sequences (NTS). Although the 45 S rRNA coding sequences were approximately threefold enriched in matrix preparations, the recovery of this DNA (unlike the NTS) was dependent on the extent of nuclease digest and proportional to the length of the matrix-attached DNA fragments. The data suggest that rDNA is organized as matrix-attached DNA loops and only the NTS are directly involved in matrix binding. Further, we demonstrated that while the kinetics and extent of nuclease digestion were similar in all regions of the DNA loops, the nuclease digestion pattern of bulk nuclear and matrix DNA showed a typical nucleosome organization, but the rDNA fragments retained with the nuclear matrix did not.
Similar content being viewed by others
References
Agutter, P. S., and Richardson, J. C. W. (1980). Nuclear non-chromatin proteinaceous structures: Their role in the organization and function of the interphase nucleus. J. Cell Sci. 44395.
Basler, J., Hastie, N. D., Pietras, D., Matsui, S. I., Sandberg, A. A., and Berezney, R. (1981). Hybridization of nuclear matrix attached deoxyribonucleic acid fragments. Biochemistry 206921.
Berezney, R. (1984). Organization and functions of the nuclear matrix. In Hnilica, L. S. (ed). Chromosomal Nonhistone Proteins Structural Associations CRC Press, Boca Raton, Fla., pp. 120–180.
Berezney, R., and Buchholtz, L. A. (1981). Isolation and characterization of rat liver nuclear matrices containing high molecular weight deoxyribonucleic acid. Biochemistry 204995.
Bodnar, J. W., Jones, C. J., Coombs, D. H., Pearson, G. D., and Ward, D. C. (1983). Proteins tightly bound to HeLa cell DNA at nuclear matrix attachment sites. Mol. Cell. Biol. 31567.
Bolla, R. I., Braaten, D. C., Shiomi, Y., Herbert, M. B., and Schlessinger, D. (1985). Localization of specific rDNA spacer sequences to the mouse L-cell nucleolar matrix. Mol. Cell. Biol. 51287.
Borchsenius, S., Bonven, B., Leer, J. C., and Westergaard, O. (1981). Nuclease-sensitive regions on the extrachromosomal r-chromatin from tetrahymena pyriforms. Eur. J. Biochem. 117245.
Bowen, B., Steinberg, J., Laemmli, U. K., and Weintraub, H. (1980). The detection of DNA-binding proteins by protein blotting. Nucleic Acids Res. 81.
Chikaraishi, D. M., Buchanan, L., Danna, K. J., and Harrington, C. A. (1983). Genomic organization of rat rDNA. Nucleic Acids Res. 116437.
Ciejek, E. M., Tsai, M. J., and O'Malley, B. W. (1983). Actively transcribed genes are associated with the nuclear matrix. Nature 306608.
Cockerill, P. N., and Garrard, W. T. (1986). Chromosomal loop anchorage of the kappa immunoglobulin gene occurs next to the enhancer in the region containing topoisomerase II sites. Cell 44273.
Davis, A. H., Reudelhuber, T. L., and Garrard, W. T. (1983). Variegated chromatin structures of mouse ribosomal RNA genes. J. Mol. Biol. 167133.
Denhardt, D. J. (1966). A membrane-filter technique for the detection of complementary DNA. Biochem. Biophys. Res. Commun. 23641.
Earnshaw, W. C., and Heck, M. M. S. (1985). Localization of topoisomerase II in mitotic chromosomes. J. Cell Biol. 1001716.
Earnshaw, W. C., and Laemmli, U. K. (1983). Architecture of metaphase chromosomes and chromosome scaffolds. J. Cell Biol. 9684.
Earnshaw, W. C., Halligan, B., Cooke, C. A., Heck, M. M. S., and Liu, L. F. (1985). Topoisomerase II is a structural component of mitotic chromosomes. J. Cell Biol. 1001706.
Ferrer, P., Caizergues-Ferrer, M., Amalric, F., and Zalta, J. P. C. (1980). Structure of nucleolar and rDNA chromatin in CHO cells. Biol. Cell. 39155.
Foe, V. E. (1977). Modulation of ribosomal RNA synthesis in Oncopeltus faciatus: An electron microscopic study of the relationship between changes in chromatin structure and transcriptional activity. Cold Spring Harbor Symp. Quant. Biol. 42723.
Foster, K. A., and Collins, J. M. (1985). The interrelation between DNA synthesis rates and DNA polymerases bound to the nuclear matrix in synchronized HeLa cells. J. Biol. Chem. 2604229.
Giri, C. P., and Gorovsky, M. A. (1980). DNase I sensitivity of ribosomal genes in isolated nucleosome core particles. Nucleic Acids Res. 8197.
Goldberg, G. J., Collier, I., and Cassel, A. (1983). Specific DNA sequences associated with the nuclear matrix in synchronized mouse 3T3 cells. Proc. Natl. Acad. Sci. USA 806887.
Gupta, R. C., Dighe, N. R., Randerath, K., and Smith, H. C. (1985). Distribution of initial and persistent 2-acetylaminofluorene-induced DNA adducts within DNA loops. Proc. Natl. Acad. Sci. USA 826605.
Hentzen, P. C., Rho, J. H., and Bekhor, I. (1984). Nuclear matrix DNA from chicken erythrocytes contains B-globin gene sequences. Proc. Natl. Acad. Sci. USA 81304.
Jackson, D. A., Cook, P. R., and Patel, S. B. (1984). Attachment of repeated sequences to the nuclear cage. Nucleic Acids Res. 126709.
Jeppesen, P. G. N., and Bankier, A. T. (1979). A partial characterization of DNA fragments protected from nuclease degradation in histone depleted metaphase chromosomes of the Chinese hamster. Nucleic Acids Res. 749.
Kafatos, F. C., Jones, C. W., and Efstratiadis, A. (1979). Determination of nucleic acid sequence homologies and relative concentrations by a dot hybridization procedure. Nucleic Acids Res. 71541.
Lebkowski, J. S., and Laemmli, U. K. (1982a). Evidence for two levels of DNA folding in histone-depleted HeLa interphase nuclei. J. Mol. Biol. 156309.
Lebkowski, J. S., and Laemmli, U. K. (1982b). Non-histone proteins and long-range organization of HeLa interphase DNA. J. Mol. Biol. 156325.
Matsumoto, L. H. (1981). Enrichment of satellite DNA on the nuclear matrix of bovine cells. Nature (London) 294481.
McCready, S. J., Godwin, J., Mason, D. W., Brazell, I. A., and Cook, P. R. (1980). DNA is replicated at the nuclear cage. J. Cell Sci. 46365.
McKnight, S. L., Martin, K. A., Beyer, A. L., and Miller, O. L. (1979). Visualization of functionally active chromatin. In Busch, H. (ed.), The Cell Nucleus, 7 Academic Press, New York, pp. 97–122.
Miesfeld, R., Krystal, M., and Arnheim, N. (1981). A member of a new repeated sequence family which is conserved throughout eucaryotic evolution is found between the human delta and fetal globin genes. Nucleic Acids Res. 95931.
Mirkovitch, J., Mirault, M. E., and Laemmli, U. K. (1984). Organization of the higher-order chromatin loop: Specific DNA attachment sites on nuclear scaffold. Cell 39223.
Mroczka, D. L., Cassidy, B., Busch, H., and Rothblum, L. I. (1984). Characterization of rat ribosomal DNA. J. Mol. Biol. 174141.
Pardoll, D. M., and Vogelstein, B. (1980). Sequence analysis of nuclear matrix associated DNA from rat liver. Exp. Cell Res. 128466.
Paulson, J. R., and Laemmli, U. K. (1977). The structure of histone-depleted metaphase chromosomes. Cell 12817.
Prior, C. P., Cantor, C. R., Johnson, E. M., Littau, V. C., and Allfrey, V. G. (1983). Reversible changes in nucleosome structure and histone H3 accessibility in transcriptionally active and inactive states of rDNA chromatin. Cell 341033.
Pruitt, S. C., and Reeder, R. H. (1984). Effect of topological constraint on transcription of ribosomal DNA in Xenopus oocytes. J. Mol. Biol. 174121.
Razin, S. V., Mantieva, V. L., and Georgiev, G. P. (1979). The similarity of DNA sequences remaining bound to scaffold upon nuclease treatment of interphase nuclei and metaphase chromosomes. Nucleic Acids Res. 71713.
Razin, S. V., Chernokohovostou, V. V., Roodyn, A. V., Zbarsky, I. B., and Georgiev, G. P. (1981). Proteins tightly bound to DNA in the regions of DNA attachment to the skeletal structures of interphase nuclei and metaphase chromosomes. Cell 2765.
Reeves, R. (1976). Ribosomal genes of Xenopus laevis: Evidence of nucleosomes in transcriptionally active chromatin. Science 194529.
Rigby, P. W. J., Dieckmann, M., Rhodes, C., and Berg, P. (1977). Labeling deoxyribonucleic acid to high specific activity in vitro by nick translation with DNA polymerase I. J. Mol. Biol. 113237.
Robinson, S. I., Small, D., Idzerda, R., McKnight, G. S., and Vogelstein, B. (1983). The association of transcriptionally active genes with the nuclear matrix of the chicken oviduct. Nucleic Acids Res. 115113.
Rothblum, L. I., Mamrack, P. M., Kunkle, H. M., Olson, O. J., and Busch, H. (1977). Fractionation of nucleoli. Enzymatic and two-dimensional polyacrylamide gel electrophoretic analysis. Biochemistry 164716.
Rothblum, L. I., Parker, D. L., and Cassidy, B. (1982a). Isolation and characterization of rat ribosomal DNA clones. Gene 1775.
Rothblum, L. I., Reddy, R., and Cassidy, B. (1982b). Transcription initiation site of rat ribosomal DNA. Nucleic Acids Res. 107345.
Scheer, U., and Zentgraf, H. (1982). Morphology of nucleolar chromatin in electron microscopic spread preparations. In Busch, H., and Rothblum, L. I. (eds.), The Cell Nucleus, 11 Academic Press, New York, pp. 143–191.
Shaper, J. H., Pardoll, D. M., Kaufmann, S. H., Barrack, E. R., Vogelstein, B., and Coffee, D. S. (1979). The relationship of the nuclear matrix to cellular structure and function. Adv Enzyme Regul. 17213.
Small, D., Nelkin, B., and Vogelstein, B. (1982). Nonrandom distribution of repeated DNA sequences with respect to supercoiled loops and the nuclear matrix. Proc. Natl. Acad. Sci. USA 795911.
Smith, H. C., and Berezney, R. (1982). Nuclear matrix bound deoxyribonucleic acid synthesis: An in vitro system. Biochemistry 216751.
Smith, H. C., and Berezney, R. (1983). Dynamic domains of DNA polymerase alpha in regenerating rat liver. Biochemistry 223042.
Smith, H. C., Puvion, E., Buchholtz, L. A., and Berezney, R. (1984). Spatial distribution of DNA loop attachment and replicational sites in the nuclear matrix. J. Cell Biol. 991794.
Smith, H. C., Ochs, R. L., Fernandez, E. A., and Spector, D. L. (1986). Macromolecular domains containing nuclear protein p107 and U-snRNP protein p28: Further evidence for an in situ nuclear matrix. Mol. Cell. Biochem. 70151.
Smith, H. C., and Ochs, R. L. Lin, D., and Chinault, A. C. (1987). Ultrastructural and biochemical comparisons of nuclear matrices prepared by high salt or LIS extraction. Mol. Cell. Biochem. 7749.
Treco, D., Brownell, E., and Arnheim, N. (1982). The ribosomal gene nontranscribed spacer. In Busch, H., and Rothblum, R. I. (eds.), The Cell Nucleus, 12 Academic Press, New York, pp. 101–126.
Vogelstein, B., Pardoll, D. M., and Coffey, D. S. (1980). Supercoiled loops and eukaryotic DNA replication. Cell 2279.
Wahl, G. M., Stern, M., and Stark, G. R. (1979). Efficient transfer of large DNA fragments from agarose gels to diazobenzyloxymethyl-paper and rapid hybridization by using dextran sulfate. Proc. Natl. Acad. Sci. USA 763683.
Werner, D., Chanpu, S., Muller, M., Spiess, E., and Plagens, U. (1984). Antibodies to the most tightly bound proteins in eukaryotic DNA. Exp. Cell Res. 151384.
Author information
Authors and Affiliations
Additional information
These studies were supported by Cancer Research Center Grant CA-10893, P4, P9, awarded by the National Cancer Institute, Department of Health the Human Services, Public Health Service.
Rights and permissions
About this article
Cite this article
Smith, H.C., Rothblum, L.I. Ribosomal DNA sequences attached to the nuclear matrix. Biochem Genet 25, 863–879 (1987). https://doi.org/10.1007/BF00502606
Received:
Revised:
Issue Date:
DOI: https://doi.org/10.1007/BF00502606