Abstract
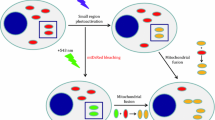
With the availability of increasing numbers of fluorescent protein variants and state-of-the-art imaging techniques, live cell microscopy has become a standard procedure in modern cell biology. Fluorescent markers are used to visualize the dynamic processes that take place in living cells, including the behavior of membrane-bound organelles. Here, we provide two examples of how we analyze the membrane dynamics of mitochondria in living yeast cells using wide field and confocal microscopy: (1) Long-term observation of mitochondrial shape changes using mitochondria-targeted fluorescent proteins and (2) monitoring the behavior of individual mitochondria using a mitochondria-targeted version of a photoconvertible fluorescent protein.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Stephens DJ, Allan VJ (2003) Light microscopy techniques for live cell imaging. Science 300:82–86
Merz S, Hammermeister M, Altmann K, Dürr M, Westermann B (2007) Molecular machinery of mitochondrial dynamics in yeast. Biol Chem 388:917–926
Westermann B (2010) Mitochondrial fusion and fission in cell life and death. Nat Rev Mol Cell Biol 11:872–884
Altmann K, Dürr M, Westermann B (2007) Saccharomyces cerevisiae as a model organism to study mitochondrial biology: general considerations and basic procedures. Methods Mol Biol 372:81–90
Swayne TC, Gay AC, Pon LA (2007) Fluorescence imaging of mitochondria in yeast. Methods Mol Biol 372:433–459
Westermann B, Neupert W (2000) Mitochondria-targeted green fluorescent proteins: convenient tools for the study of organelle biogenesis in Saccharomyces cerevisiae. Yeast 16:1421–1427
Swayne TC, Gay AC, Pon LA (2007) Visualization of mitochondria in budding yeast. Methods Cell Biol 80:591–626
Nunnari J, Marshall WF, Straight A, Murray A, Sedat JW, Walter P (1997) Mitochondrial transmission during mating in Saccharomyces cerevisiae is determined by mitochondrial fusion and fission and the intramitochondrial segregation of mitochondrial DNA. Mol Biol Cell 8:1233–1242
Chen H, Detmer SA, Ewald AJ, Griffin EE, Fraser SE, Chan DC (2003) Mitofusins Mfn1 and Mfn2 coordinately regulate mitochondrial fusion and are essential for embryonic development. J Cell Biol 160:189–200
Legros F, Lombes A, Frachon P, Rojo M (2002) Mitochondrial fusion in human cells is efficient, requires the inner membrane potential, and is mediated by mitofusins. Mol Biol Cell 13:4343–4354
Mattenberger Y, James DI, Martinou JC (2003) Fusion of mitochondria in mammalian cells is dependent on the mitochondrial inner membrane potential and independent of microtubules or actin. FEBS Lett 538:53–59
Ishihara N, Jofuku A, Eura Y, Mihara K (2003) Regulation of mitochondrial morphology by membrane potential, and DRP1-dependent division and FZO1-dependent fusion reaction in mammalian cells. Biochem Biophys Res Commun 301:891–898
Sheahan MB, McCurdy DW, Rose RJ (2005) Mitochondria as a connected population: ensuring continuity of the mitochondrial genome during plant cell dedifferentiation through massive mitochondrial fusion. Plant J 44:744–755
Chudakov DM, Matz MV, Lukyanov S, Lukyanov KA (2010) Fluorescent proteins and their applications in imaging living cells and tissues. Physiol Rev 90:1103–1163
Lukyanov KA, Chudakov DM, Lukyanov S, Verkhusha VV (2005) Innovation: photoactivatable fluorescent proteins. Nat Rev Mol Cell Biol 6:885–891
Jakobs S, Schauss AC, Hell SW (2003) Photoconversion of matrix targeted GFP enables analysis of continuity and intermixing of the mitochondrial lumen. FEBS Lett 554:194–200
Karbowski M, Arnoult D, Chen H, Chan DC, Smith CL, Youle RJ (2004) Quantitation of mitochondrial dynamics by photolabeling of individual organelles shows that mitochondrial fusion is blocked during the Bax activation phase of apoptosis. J Cell Biol 164:493–499
Molina AJA, Shirihai OS (2009) Monitoring mitochondrial dynamics with photoactivatable green fluorescent protein. Methods Enzymol 457:289–304
Arimura S, Yamamoto J, Aida GP, Nakazono M, Tsutsumi N (2004) Frequent fusion and fission of plant mitochondria with unequal nucleoid distribution. Proc Natl Acad Sci USA 101:7805–7808
Brachmann CB, Davies A, Cost GJ, Caputo E, Li J, Hieter P, Boeke JD (1998) Designer deletion strains derived from Saccharomyces cerevisiae S288C: a useful set of strains and plasmids for PCR-mediated gene disruption and other applications. Yeast 14:115–132
Keppler-Ross S, Noffz C, Dean N (2008) A new purple fluorescent color marker for genetic studies in Saccharomyces cerevisiae and Candida albicans. Genetics 179:705–710
Sikorski RS, Hieter P (1989) A system of shuttle vectors and host strains designed for efficient manipulation of DNA in Saccharomyces cerevisiae. Genetics 122:19–27
Chudakov DM, Lukyanov S, Lukyanov KA (2007) Tracking intracellular protein movements using photoswitchable fluorescent proteins PS-CFP2 and Dendra2. Nat Protoc 2:2024–2032
Wallace W, Schaefer LH, Swedlow JR (2001) A workingperson’s guide to deconvolution in light microscopy. Biotechniques 31:1076–1078
Acknowledgment
This work was supported by the Deutsche Forschungsgemeinschaft through grants We 2174/4-2 and 5-1.
Author information
Authors and Affiliations
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2013 Springer Science+Business Media, LLC
About this protocol
Cite this protocol
Scholz, D., Förtsch, J., Böckler, S., Klecker, T., Westermann, B. (2013). Analyzing Membrane Dynamics with Live Cell Fluorescence Microscopy with a Focus on Yeast Mitochondria. In: Rapaport, D., Herrmann, J. (eds) Membrane Biogenesis. Methods in Molecular Biology, vol 1033. Humana Press, Totowa, NJ. https://doi.org/10.1007/978-1-62703-487-6_17
Download citation
DOI: https://doi.org/10.1007/978-1-62703-487-6_17
Published:
Publisher Name: Humana Press, Totowa, NJ
Print ISBN: 978-1-62703-486-9
Online ISBN: 978-1-62703-487-6
eBook Packages: Springer Protocols