Abstract
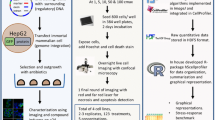
Exposure to oxidative radical species and cytokine-mediated inflammatory stress are established contributors to hepatocyte cell death during cholestasis. Cellular counter measures against those stressors are called adaptive stress response pathways. While in early stages of the disease adaptive stress pathways protect the hepatocytes, in later stages during prolonged stressed conditions they fail. The quantitative imaging-based assessment of cellular stress response pathways using the HepG2 BAC-GFP response reporter platform is a powerful strategy to evaluate the impact of chemical substances and gene knockdown on activation of adaptive stress response pathways, hence allowing systematic screening for positive or negative influences on cholestasis progression. This protocol allows the application of a highly versatile screening tool for a systematic evaluation of the effect of compounds having cholestasis liability and affected genes during cholestatic injury on cellular adaptive stress pathway activation. The approach involves high-throughput live-cell visualization of GFP-tagged key proteins of the oxidative stress response/Nrf2 pathway and inflammatory cytokine signaling. Quantitative image analysis of temporal responses of individual cells is followed by informatics analysis. The overall practical approaches are discussed in this chapter.
Access this chapter
Tax calculation will be finalised at checkout
Purchases are for personal use only
Similar content being viewed by others
References
Wink S, Hiemstra S, Huppelschoten S et al (2014) Quantitative high content imaging of cellular adaptive stress response pathways in toxicity for chemical safety assessment. Chem Res Toxicol 27:338–355
Copple BL, Jaeschke H, Klaassen CD (2010) Oxidative stress and the pathogenesis of cholestasis. Semin Liver Dis 30:195–204
Yamamoto M, Kensler TW, Motohashi H (2018) The KEAP1-NRF2 system: a thiol-based sensor-effector apparatus for maintaining redox homeostasis. Phys Rev 98:1169–1203
Okada K, Shoda J, Taguchi K et al (2009) Nrf2 counteracts cholestatic liver injury via stimulation of hepatic defense systems. Biochem Biophys Res Commun 389:431–436
Tanaka Y, Aleksunes LM, Cui YJ et al (2009) ANIT-induced intrahepatic cholestasis alters hepatobiliary transporter expression via Nrf2-dependent and independent signaling. Toxicol Sci 108:247–257
Weerachayaphorn J, Luo Y, Mennone A et al (2014) Deleterious effect of oltipraz on extrahepatic cholestasis in bile duct-ligated mice. J Hepatol 60:160–166
Gujral JS, Farhood A, Bajt ML et al (2003) Neutrophils aggravate acute liver injury during obstructive cholestasis in bile duct–ligated mice. Hepatology 38:355–363
Woolbright BL, Jaeschke H (2018) Mechanisms of inflammatory liver injury and drug-induced hepatotoxicity. Curr Pharm Rep 5:346–357
Gehrke N, Nagel M, Straub BK et al (2018) Loss of cellular FLICE-inhibitory protein promotes acute cholestatic liver injury and inflammation from bile duct ligation. Am J Physiol Gastrointest Liver Physiol 314:G319–GG33
Allen K, Jaeschke H, Copple BL (2011) Bile acids induce inflammatory genes in hepatocytes: a novel mechanism of inflammation during obstructive cholestasis. Am J Pathol 178:175–186
Qin P, Borges-Marcucci LA, Evans MJ et al (2005) Bile acid signaling through FXR induces intracellular adhesion molecule-1 expression in mouse liver and human hepatocytes. Am J Physiol Gastrointest Liver Physiol 289:G267–GG73
Cai S-Y, Boyer JL (2017) Studies on the mechanisms of bile acid initiated hepatic inflammation in cholestatic liver injury. Inflamm Cell Signal 4:e1561
Wink S, Hiemstra SW, Huppelschoten S et al (2018) Dynamic imaging of adaptive stress response pathway activation for prediction of drug induced liver injury. Arch Toxicol 92:1797–1814
Van den Hof WFPM, Coonen MLJ, van Herwijnen M et al (2014) Classification of hepatotoxicants using HepG2 cells: a proof of principle study. Chem Chem Res Toxicol 27:433–442
Niemeijer M, Hiemstra S, Wink S et al (2018) Systems microscopy approaches in unraveling and predicting drug-induced liver injury (DILI). In: Chen M, Will Y (eds) Drug-induced liver toxicity, 1st edn. Springer Nature, New York
Schindelin J, Arganda-Carreras I, Frise E et al (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9:676–682
Du G (2012) FociPicker3D 3D and 2D particle counter. Institute of Modern Physics, CAS, China
Di Z, Herpers B, Fredriksson L et al (2012) Automated analysis of NF-kappaB nuclear translocation kinetics in high-throughput screening. PLoS One 7:e52337
Oppelt A, Kaschek D, Huppelschoten S et al (2018) Model-based identification of TNFα-induced IKKβ-mediated and IκBα-mediated regulation of NFκB signal transduction as a tool to quantify the impact of drug-induced liver injury compounds. npjsba 4:23
Ritz C, Baty F, Streibig JC et al (2016) Dose-response analysis using R. PLoS One 10:e0146021
Davis JA, Gift JS, Zhao QJ (2011) Introduction to benchmark dose methods and U.S. EPA's benchmark dose software (BMDS) version 2.1.1. Toxicol Appl Pharmacol 254:181–191
Akaike H (1973) Information theory and an extension of the maximum likelihood principle. In: Petrov BN, Csáki F (eds) Second international symposium on information theory. Akadémiai Kiadó, Tsahkadsor, Armenia, U.S.S.R.: Budapest, pp 267–281
Wickham H (2016) ggplot2—Elegant graphics for data analysis, vol XVI. Springer, New York, 260 pp
Poser I, Sarov M, Hutchins JR et al (2008) BAC TransgeneOmics: a high-throughput method for exploration of protein function in mammals. Nat Methods 5:409–415
Muyrers JP, Zhang Y, Testa G et al (1999) Rapid modification of bacterial artificial chromosomes by ET-recombination. Nucleic Acids Res 27:1555–1557
Puigvert JC, de Bont H, van de Water B et al (2010) High-throughput live cell imaging of apoptosis. Curr Protoc Cell Biol Chapter 18:Unit 18.0.1-3
Yan K, Verbeek FJ (2012) Segmentation for high-throughput image analysis: watershed masked clustering, Berlin, Heidelberg, 2012. Springer, Berlin, Heidelberg, pp 25–41
R Development Core Team. (2013) R: a language and environment for statistical computing. R foundation for Statistical Computing
Vermes I, Haanen C, Steffens-Nakken H, Reutelingsperger C (1995) A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labelled Annexin V. J Immunol Methods 184:39–51
Hanshaw RG, Smith BD (2005) New reagents for phosphatidylserine recognition and detection of apoptosis. Bioorg Med Chem 13:5035–5042
Acknowledgments
This work was supported by the FP7 DETECTIVE project (grant agreement 266838), IMI IP-DILI project (grant agreement 115336), and H2020 EU-ToxRisk project (grant agreement 681002).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2019 Springer Science+Business Media, LLC, part of Springer Nature
About this protocol
Cite this protocol
Schimming, J.P., ter Braak, B., Niemeijer, M., Wink, S., van de Water, B. (2019). System Microscopy of Stress Response Pathways in Cholestasis Research. In: Vinken, M. (eds) Experimental Cholestasis Research. Methods in Molecular Biology, vol 1981. Humana, New York, NY. https://doi.org/10.1007/978-1-4939-9420-5_13
Download citation
DOI: https://doi.org/10.1007/978-1-4939-9420-5_13
Published:
Publisher Name: Humana, New York, NY
Print ISBN: 978-1-4939-9419-9
Online ISBN: 978-1-4939-9420-5
eBook Packages: Springer Protocols