Abstract
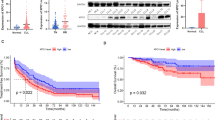
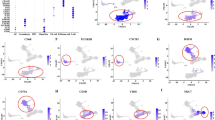
Chromosomal rearrangement involving 14q32 region that results in TNF receptor associated factor 3 (TRAF3) dysfunctional mutation is the most frequent NF-κB pathway mutation in multiple myeloma (MM). Subsequent NF-κB inducing Kinase (NIK) stabilization plays a critical role in alternative NF-κB activation. However, disease progression resulting from TRAF3 dysregulation has not been well understood. In this study, we identified lymphocyte cellular protein 1 (LCP1) as a novel NIK-driven alternative NF-κB target in TRAF3 dysfunctional mutation using RNA-seq, ChIP-seq (RelA/p65 and p52 NF-κB) and other validation methods. LCP1 is exclusively activated in MM cells with TRAF3 loss-of-function mutation. In MM patients, higher LCP1 expression was significantly pronounced in poor prognosis groups such as 4p16 and MAF. CD138 negative MM patient cells showed elevated LCP1 expression and inhibition of LCP1 can sensitize proteasome inhibitor bortezomib in TRAF3 mutant MM cells in vitro. We report that LCP1 is a NIK-driven biomarker in TRAF3 dysfunctional MM and targeting LCP1 can provide a valuable therapeutic intervention in TRAF3 mutated MM.







Similar content being viewed by others
References
Akincilar, S. C., Low, K. C., Liu, C. Y., Yan, T. D., Oji, A., Ikawa, M., et al. (2015). Quantitative assessment of telomerase components in cancer cell lines. FEBS Letters, 589, 974–984.
Annunziata, C. M., Davis, R. E., Demchenko, Y., Bellamy, W., Gabrea, A., Zhan, F., et al. (2007). Frequent engagement of the classical and alternative NF-kappaB pathways by diverse genetic abnormalities in multiple myeloma. Cancer Cell, 12, 115–130.
Ashburner, M., Ball, C. A., Blake, J. A., Botstein, D., Butler, H., Cherry, J. M., et al. (2000). Gene ontology: tool for the unification of biology. The gene ontology consortium. Nature Genetics, 25, 25–29.
Bray, N. L., Pimentel, H., Melsted, P., & Pachter, L. (2016). Near-optimal probabilistic RNA-seq quantification. Nature Biotechnology, 34, 525–527.
Chew, C. L., Conos, S. A., Unal, B., & Tergaonkar, V. (2018). Noncoding RNAs: master regulators of inflammatory signaling. Trends in Molecular Medicine, 24, 66–84.
Cildir, G., Akincilar, S. C., & Tergaonkar, V. (2013). Chronic adipose tissue inflammation: all immune cells on the stage. Trends in Molecular Medicine, 19, 487–500.
Claudio, E., Brown, K., Park, S., Wang, H., & Siebenlist, U. (2002). BAFF-induced NEMO-independent processing of NF-kappa B2 in maturing B cells. Nature immunology, 3, 958–965.
Colaprico, A., Silva, T. C., Olsen, C., Garofano, L., Cava, C., Garolini, D., et al. (2016). TCGAbiolinks: an R/Bioconductor package for integrative analysis of TCGA data. Nucleic Acids Research, 44, e71.
Demchenko, Y. N., Brents, L. A., Li, Z., Bergsagel, L. P., McGee, L. R., & Kuehl, M. W. (2014). Novel inhibitors are cytotoxic for myeloma cells with NFkB inducing kinase-dependent activation of NFkB. Oncotarget, 5, 4554–4566.
Demchenko, Y. N., Glebov, O. K., Zingone, A., Keats, J. J., Bergsagel, P. L., & Kuehl, W. M. (2010). Classical and/or alternative NF-kappaB pathway activation in multiple myeloma. Blood, 115, 3541–3552.
Dubovsky, J. A., Chappell, D. L., Harrington, B. K., Agrawal, K., Andritsos, L. A., Flynn, J. M., et al. (2013). Lymphocyte cytosolic protein 1 is a chronic lymphocytic leukemia membrane-associated antigen critical to niche homing. Blood, 122, 3308–3316.
Dun, M. D., Chalkley, R. J., Faulkner, S., Keene, S., Avery-Kiejda, K. A., Scott, R. J., et al. (2015). Proteotranscriptomic profiling of 231-BR breast cancer cells: identification of potential biomarkers and therapeutic targets for brain metastasis. Molecular and Cellular Proteomics, 14, 2316–2330.
Fang, Z. Q., Zang, W. D., Chen, R., Ye, B. W., Wang, X. W., Yi, S. H., et al. (2013). Gene expression profile and enrichment pathways in different stages of bladder cancer. Genetics and Molecular Research: GMR, 12, 1479–1489.
Foran, E., McWilliam, P., Kelleher, D., Croke, D. T., & Long, A. (2006). The leukocyte protein l-plastin induces proliferation, invasion and loss of E-cadherin expression in colon cancer cells. International Journal of Cancer, 118, 2098–2104.
Grossman, R. L., Heath, A. P., Ferretti, V., Varmus, H. E., Lowy, D. R., Kibbe, W. A., et al. (2016). Toward a shared vision for cancer genomic data. New England Journal of Medicine, 375, 1109–1112.
Hacker, H., Redecke, V., Blagoev, B., Kratchmarova, I., Hsu, L. C., Wang, G. G., et al. (2006). Specificity in Toll-like receptor signalling through distinct effector functions of TRAF3 and TRAF6. Nature, 439, 204–207.
Hacker, H., Tseng, P. H., & Karin, M. (2011). Expanding TRAF function: TRAF3 as a tri-faced immune regulator. Nature REVIEWS Immunology, 11, 457–468.
Hayden, M. S., & Ghosh, S. (2008). Shared principles in NF-kappaB signaling. Cell, 132, 344–362.
Inaguma, S., Riku, M., Ito, H., Tsunoda, T., Ikeda, H., & Kasai, K. (2015). GLI1 orchestrates CXCR4/CXCR7 signaling to enhance migration and metastasis of breast cancer cells. Oncotarget, 6, 33648–33657.
Kawano, Y., Fujiwara, S., Wada, N., Izaki, M., Yuki, H., Okuno, Y., et al. (2012). Multiple myeloma cells expressing low levels of CD138 have an immature phenotype and reduced sensitivity to lenalidomide. International Journal of Oncology, 41, 876–884.
Keats, J. J., Fonseca, R., Chesi, M., Schop, R., Baker, A., Chng, W. J., et al. (2007). Promiscuous mutations activate the noncanonical NF-kappaB pathway in multiple myeloma. Cancer Cell, 12, 131–144.
Khattar, E., Kumar, P., Liu, C. Y., Akincilar, S. C., Raju, A., Lakshmanan, M., et al. (2016). Telomerase reverse transcriptase promotes cancer cell proliferation by augmenting tRNA expression. The Journal of Clinical Investigation, 126, 4045–4060.
Khattar, E., Maung, K. Z. Y., Chew, C. L., Ghosh, A., Mok, M. M. H., Lee, P., et al. (2019). Rap1 regulates hematopoietic stem cell survival and affects oncogenesis and response to chemotherapy. Nature Communications, 10, 5349.
Li, Y., Cheng, H. S., Chng, W. J., & Tergaonkar, V. (2016). Activation of mutant TERT promoter by RAS-ERK signaling is a key step in malignant progression of BRAF-mutant human melanomas. Proceedings of the National Academy Sciences USA, 113, 14402–14407.
Li, Y., Zhou, Q. L., Sun, W., Chandrasekharan, P., Cheng, H. S., Ying, Z., et al. (2015). Non-canonical NF-kappaB signalling and ETS1/2 cooperatively drive C250T mutant TERT promoter activation. Nature Cell Biology, 17, 1327–1338.
Lohr, J. G., Stojanov, P., Carter, S. L., Cruz-Gordillo, P., Lawrence, M. S., Auclair, D., et al. (2014). Widespread genetic heterogeneity in multiple myeloma: implications for targeted therapy. Cancer Cell, 25, 91–101.
Manojlovic, Z., Christofferson, A., Liang, W. S., Aldrich, J., Washington, M., Wong, S., et al. (2017). Comprehensive molecular profiling of 718 multiple myelomas reveals significant differences in mutation frequencies between African and European descent cases. PLoS Genetics, 13, e1007087.
Medzhitov, R. (2008). Origin and physiological roles of inflammation. Nature, 454, 428–435.
Mi, H., Muruganujan, A., Ebert, D., Huang, X., & Thomas, P. D. (2019). PANTHER version 14: more genomes, a new PANTHER GO-slim and improvements in enrichment analysis tools. Nucleic Acids Research, 47, D419–D426.
Ning, Y., Gerger, A., Zhang, W., Hanna, D. L., Yang, D., Winder, T., et al. (2014). Plastin polymorphisms predict gender- and stage-specific colon cancer recurrence after adjuvant chemotherapy. Molecular Cancer Therapeutics, 13, 528–539.
Oganesyan, G., Saha, S. K., Guo, B., He, J. Q., Shahangian, A., Zarnegar, B., et al. (2006). Critical role of TRAF3 in the Toll-like receptor-dependent and -independent antiviral response. Nature, 439, 208–211.
Ooi, M. G., de Mel, S., & Chng, W. J. (2016). Risk stratification in multiple myeloma. Current Hematologic Malignancy Reports, 11, 137–147.
Ozturk, M. B., Li, Y., & Tergaonkar, V. (2017). Current insights to regulation and role of telomerase in human diseases. Antioxidants (Basel), 6(1), 17. https://doi.org/10.3390/antiox6010017.
Puar, Y. R., Shanmugam, M. K., Fan, L., Arfuso, F., Sethi, G., & Tergaonkar, V. (2018). Evidence for the involvement of the master transcription factor NF-kappaB in cancer initiation and progression. Biomedicines, 6(3), 82. https://doi.org/10.3390/biomedicines6030082.
Ranuncolo, S. M., Pittaluga, S., Evbuomwan, M. O., Jaffe, E. S., & Lewis, B. A. (2012). Hodgkin lymphoma requires stabilized NIK and constitutive RelB expression for survival. Blood, 120, 3756–3763.
Ryu, D., Kim, S. J., Hong, Y., Jo, A., Kim, N., Kim, H. J., et al. (2020). Alterations in the transcriptional programs of myeloma cells and the microenvironment during extramedullary progression affect proliferation and immune evasion. Clinical Cancer Research, 26, 935–944.
Shin, E. M., Hay, H. S., Lee, M. H., Goh, J. N., Tan, T. Z., Sen, Y. P., et al. (2014). DEAD-box helicase DP103 defines metastatic potential of human breast cancers. The Journal of Clinical Investigation, 124, 3807–3824.
Stuhmer, T., Chatterjee, M., Hildebrandt, M., Herrmann, P., Gollasch, H., Gerecke, C., et al. (2005). Nongenotoxic activation of the p53 pathway as a therapeutic strategy for multiple myeloma. Blood, 106, 3609–3617.
Su Kim, D., Choi, Y. D., Moon, M., Kang, S., Lim, J. B., Kim, K. M., et al. (2013). Composite three-marker assay for early detection of kidney cancer. Cancer Epidemiology, Biomarkers and Prevention: a Publication of the American Association for Cancer Research, Cosponsored by the American Society of Preventive Oncology, 22, 390–398.
Sun, S. C. (2011). Non-canonical NF-kappaB signaling pathway. Cell Research, 21, 71–85.
Takeuchi, O., & Akira, S. (2010). Pattern recognition receptors and inflammation. Cell, 140, 805–820.
Tang, Z., Li, C., Kang, B., Gao, G., Li, C., & Zhang, Z. (2017). GEPIA: a web server for cancer and normal gene expression profiling and interactive analyses. Nucleic Acids Research, 45, W98–W102.
The Gene Ontology, Consortium. (2019). The gene ontology resource: 20 years and still going strong. Nucleic Acids Research, 47, D330–D338.
Torre, D., Lachmann, A., & Ma'ayan, A. (2018). BioJupies: automated generation of interactive notebooks for RNA-seq data analysis in the cloud. Cell Systems, 7(556–61), e3.
Vallabhapurapu, S., & Karin, M. (2009). Regulation and function of NF-kappaB transcription factors in the immune system. Annual Review of Immunology, 27, 693–733.
Wang, C. Q., Chin, D. W., Chooi, J. Y., Chng, W. J., Taniuchi, I., Tergaonkar, V., et al. (2015). Cbfb deficiency results in differentiation blocks and stem/progenitor cell expansion in hematopoiesis. Leukemia, 29, 753–757.
Wang, X., Spandidos, A., Wang, H., & Seed, B. (2012). PrimerBank: a PCR primer database for quantitative gene expression analysis, 2012 update. Nucleic Acids Research, 40, D1144–D1149.
Wen, D., Nong, Y., Morgan, J. G., Gangurde, P., Bielecki, A., Dasilva, J., et al. (2006). A selective small molecule IkappaB kinase beta inhibitor blocks nuclear factor kappaB-mediated inflammatory responses in human fibroblast-like synoviocytes, chondrocytes, and mast cells. The Journal of Pharmacology and Experimental Therapeutics, 317, 989–1001.
Xiao, G., Fong, A., & Sun, S. C. (2004). Induction of p100 processing by NF-kappaB-inducing kinase involves docking IkappaB kinase alpha (IKKalpha) to p100 and IKKalpha-mediated phosphorylation. The Journal of Biological Chemistry, 279, 30099–30105.
Xiao, G., Harhaj, E. W., & Sun, S. C. (2001a). NF-kappaB-inducing kinase regulates the processing of NF-kappaB2 p100. Molecular Cell, 7, 401–409.
Xiao, G., Harhaj, E. W., & Sun, S. C. (2001b). NF-kappaB-inducing kinase regulates the processing of NF-kappaB2 p100. Molecular Cell, 7, 401–409.
Xie, P., Stunz, L. L., Larison, K. D., Yang, B., & Bishop, G. A. (2007). Tumor necrosis factor receptor-associated factor 3 is a critical regulator of B cell homeostasis in secondary lymphoid organs. Immunity, 27, 253–267.
Xu, Y., Cheng, G., & Baltimore, D. (1996). Targeted disruption of TRAF3 leads to postnatal lethality and defective T-dependent immune responses. Immunity, 5, 407–415.
Xu, X., Li, Y., Bharath, S. R., Ozturk, M. B., Bowler, M. W., Loo, B. Z. L., et al. (2018). Structural basis for reactivating the mutant TERT promoter by cooperative binding of p52 and ETS1. Nature Communications, 9, 3183.
Yuregir, O. O., Sahin, F. I., Yilmaz, Z., Kizilkilic, E., Karakus, S., & Ozdogu, H. (2009). Fluorescent in situ hybridization studies in multiple myeloma. Hematology, 14, 90–94.
Zarnegar, B. J., Wang, Y., Mahoney, D. J., Dempsey, P. W., Cheung, H. H., He, J., et al. (2008). Noncanonical NF-kappaB activation requires coordinated assembly of a regulatory complex of the adaptors cIAP1, cIAP2, TRAF2 and TRAF3 and the kinase NIK. Nature Immunology, 9, 1371–1378.
Zhao, B., Barrera, L. A., Ersing, I., Willox, B., Schmidt, S. C., Greenfeld, H., et al. (2014). The NF-kappaB genomic landscape in lymphoblastoid B cells. Cell Reports, 8, 1595–1606.
Acknowledgements
We thank Dr. Manikandan Lakshmanan, Dr. Chew Chen Li, Dr. Lee Sook Yee, Dr. Gireedhar Venkatachalam for their productive discussions and suggestions for the manuscript.
Funding
This study was funded by Singapore Ministry of Health, National Medical Research Council Grant NMRC/CNIG/1170/2017.
Author information
Authors and Affiliations
Contributions
EMS performed the majority of experiments, SAN prepared RNA-seq samples, THC, NB, KF, JYHC performed bioinformatics studies. VT, WJC and MO designed the study and EMS and MO drafted the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interests.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Shin, E.M., Neja, S.A., Fidan, K. et al. Lymphocyte cytosolic protein 1 (LCP1) is a novel TRAF3 dysregulation biomarker with potential prognostic value in multiple myeloma. GENOME INSTAB. DIS. 1, 286–299 (2020). https://doi.org/10.1007/s42764-020-00014-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s42764-020-00014-x