Abstract
Background
The causal relationship and the direction of the effect between depression and aging remain controversial.
Methods
We used a bidirectional two-sample Mendelian randomization analysis to examine the relationship between depression and age proxy indicators. We obtained pooled statistics from genome-wide association studies (GWAS) on depression and the age proxy indicators. We employed five MR analysis methods to address potential biases and ensure robustness of our results, with the inverse variance weighted (IVW) method being the primary outcome. We also conducted outlier exclusion using Radial MR, MRPRESSO, and MR Steiger filters. Additionally, sensitivity analyses were performed to assess heterogeneity and pleiotropy.
Results
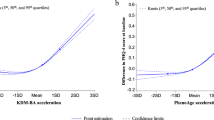
Our MR analysis revealed that depression causally leads to shortened telomere length (β = − 0.014; P = 0.038), increased frailty index (β = 0.076; P = 0.000), and accelerated GrimAge (β = 0.249; P = 0.024). Furthermore, our findings showed that the frailty index (OR = 1.679; P = 0.001) was causally associated with an increased risk of depression. Additionally, we found that appendicular lean mass (OR = 0.929; P = 0.000) and left-hand grip strength (OR = 0.836; P = 0.014) were causally associated with a reduced risk of depression. Sensitivity analyses demonstrated the robustness of our findings.
Conclusions
Our study provides evidence that depression contributes to the accelerated aging process, resulting in decreased telomere length, increased frailty index, and accelerated GrimAge. Additionally, we found that the frailty index increases the risk of depression, while appendicular lean mass and left-handed grip strength reduce the risk of depression.




Similar content being viewed by others
Availability of data and materials
Summary statistics for TL, FI, ALM, usual walking pace, MVPA, handed grip strength, and cognitive performance were derived from the IEU OpenGWAS Project at https://gwas.mrcieu.ac.uk/.Summary statistics for FA were downloaded from: http://fastgwa.info/.Summary statistics for depression were downloaded from FinnGen Consortium at https://www.finngen.fi/en/access_results.Summary statistics for epigenetic age acceleration measures of HannumAge, Intrinsic HorvathAge, PhenoAge, and GrimAge were downloaded from: https://datashare.is.ed.ac.uk/handle/10283/3645.
References
Beard JR, Officer A, de Carvalho IA et al (2016) The World report on ageing and health: a policy framework for healthy ageing. Lancet 387:2145–2154. https://doi.org/10.1016/S0140-6736(15)00516-4
Partridge L, Deelen J, Slagboom PE (2018) Facing up to the global challenges of ageing. Nature 561:45–56. https://doi.org/10.1038/s41586-018-0457-8
Cruz-Jentoft AJ, Baeyens JP, Bauer JM et al (2010) Sarcopenia: European consensus on definition and diagnosis: report of the European Working Group on sarcopenia in older people. Age Ageing 39:412–423. https://doi.org/10.1093/ageing/afq034
Wang K, Liu H, Hu Q et al (2022) Epigenetic regulation of aging: implications for interventions of aging and diseases. Sig Transduct Target Ther 7:1–22. https://doi.org/10.1038/s41392-022-01211-8
Brunet A, Berger SL (2014) Epigenetics of aging and aging-related disease. J Gerontol A Biol Sci Med Sci 69:S17–S20. https://doi.org/10.1093/gerona/glu042
López-Otín C, Blasco MA, Partridge L et al (2013) The hallmarks of aging. Cell 153:1194–1217. https://doi.org/10.1016/j.cell.2013.05.039
Chakravarti D, LaBella KA, DePinho RA (2021) Telomeres: history, health, and hallmarks of aging. Cell 184:306–322. https://doi.org/10.1016/j.cell.2020.12.028
Jylhävä J, Pedersen NL, Hägg S (2017) Biological age predictors. EBioMedicine 21:29–36. https://doi.org/10.1016/j.ebiom.2017.03.046
Rossiello F, Jurk D, Passos JF et al (2022) Telomere dysfunction in ageing and age-related diseases. Nat Cell Biol 24:135–147. https://doi.org/10.1038/s41556-022-00842-x
Gao X, Zhang Y, Mons U et al (2018) Leukocyte telomere length and epigenetic-based mortality risk score: associations with all-cause mortality among older adults. Epigenetics 13:846–857. https://doi.org/10.1080/15592294.2018.1514853
Hoogendijk EO, Afilalo J, Ensrud KE et al (2019) Frailty: implications for clinical practice and public health. Lancet 394:1365–1375. https://doi.org/10.1016/S0140-6736(19)31786-6
Franco AC, Aveleira C, Cavadas C (2022) Skin senescence: mechanisms and impact on whole-body aging. Trends Mol Med 28:97–109. https://doi.org/10.1016/j.molmed.2021.12.003
Gonzales MM, Garbarino VR, Pollet E et al (2022) Biological aging processes underlying cognitive decline and neurodegenerative disease. J Clin Invest. https://doi.org/10.1172/JCI158453
Malhi GS, Mann JJ (2018) Depression. Lancet 392:2299–2312. https://doi.org/10.1016/S0140-6736(18)31948-2
GBD 2017 Disease and Injury Incidence and Prevalence Collaborators (2018) Global, regional, and national incidence, prevalence, and years lived with disability for 354 diseases and injuries for 195 countries and territories, 1990–2017: a systematic analysis for the Global Burden of Disease Study 2017. Lancet 392:1789–1858. https://doi.org/10.1016/S0140-6736(18)32279-7
Fiedorowicz JG (2014) Depression and cardiovascular disease: an update on how course of illness may influence risk. Curr Psychiatry Rep 16:492. https://doi.org/10.1007/s11920-014-0492-6
(2015) Depression, anxiety and 6-year risk of cardiovascular disease. J Psychosomatic Res 78:123–129. https://doi.org/10.1016/j.jpsychores.2014.10.007
Wolkowitz OM, Reus VI, Mellon SH (2011) Of sound mind and body: depression, disease, and accelerated aging. Dialog Clin Neurosci 13:25–39. https://doi.org/10.31887/DCNS.2011.13.1/owolkowitz
Protsenko E, Yang R, Nier B et al (2021) “GrimAge”, an epigenetic predictor of mortality, is accelerated in major depressive disorder. Transl Psychiatry 11:1–9. https://doi.org/10.1038/s41398-021-01302-0
Stanley K (2007) Design of randomized controlled trials. Circulation 115:1164–1169. https://doi.org/10.1161/CIRCULATIONAHA.105.594945
Davey Smith G, Hemani G (2014) Mendelian randomization: genetic anchors for causal inference in epidemiological studies. Hum Mol Genet 23:R89–R98. https://doi.org/10.1093/hmg/ddu328
Burgess S, Scott RA, Timpson NJ et al (2015) Using published data in Mendelian randomization: a blueprint for efficient identification of causal risk factors. Eur J Epidemiol 30:543–552. https://doi.org/10.1007/s10654-015-0011-z
Bohannon RW (2015) Muscle strength: clinical and prognostic value of hand-grip dynamometry. Curr Opin Clin Nutr Metab Care 18:465. https://doi.org/10.1097/MCO.0000000000000202
Pei Y-F, Liu Y-Z, Yang X-L et al (2020) The genetic architecture of appendicular lean mass characterized by association analysis in the UK Biobank study. Commun Biol 3:608. https://doi.org/10.1038/s42003-020-01334-0
Ruth Mitchell E (2019) MRC IEU UK Biobank GWAS pipeline version 2. In: data.bris. https://data.bris.ac.uk/data/dataset/pnoat8cxo0u52p6ynfaekeigi. Accessed 16 July 2023
Klimentidis YC, Raichlen DA, Bea J et al (2018) Genome-wide association study of habitual physical activity in over 377,000 UK Biobank participants identifies multiple variants including CADM2 and APOE. Int J Obes (Lond) 42:1161–1176. https://doi.org/10.1038/s41366-018-0120-3
Jiang L, Zheng Z, Qi T et al (2019) A resource-efficient tool for mixed model association analysis of large-scale data. Nat Genet 51:1749–1755. https://doi.org/10.1038/s41588-019-0530-8
Atkins JL, Jylhävä J, Pedersen NL et al (2021) A genome-wide association study of the frailty index highlights brain pathways in ageing. Aging Cell 20:e13459. https://doi.org/10.1111/acel.13459
Lee JJ, Wedow R, Okbay A et al (2018) Gene discovery and polygenic prediction from a genome-wide association study of educational attainment in 1.1 million individuals. Nat Genet 50:1112–1121. https://doi.org/10.1038/s41588-018-0147-3
Codd V, Wang Q, Allara E et al (2021) Polygenic basis and biomedical consequences of telomere length variation. Nat Genet 53:1425–1433. https://doi.org/10.1038/s41588-021-00944-6
McCartney DL, Min JL, Richmond RC et al (2021) Genome-wide association studies identify 137 genetic loci for DNA methylation biomarkers of aging. Genome Biol 22:194. https://doi.org/10.1186/s13059-021-02398-9
Hannum G, Guinney J, Zhao L et al (2013) Genome-wide methylation profiles reveal quantitative views of human aging rates. Mol Cell 49:359–367. https://doi.org/10.1016/j.molcel.2012.10.016
Horvath S (2013) DNA methylation age of human tissues and cell types. Genome Biol 14:R115. https://doi.org/10.1186/gb-2013-14-10-r115
Levine ME, Lu AT, Quach A et al (2018) An epigenetic biomarker of aging for lifespan and healthspan. Aging (Albany NY) 10:573–591. https://doi.org/10.18632/aging.101414
Lu AT, Quach A, Wilson JG et al (2019) DNA methylation GrimAge strongly predicts lifespan and healthspan. Aging (Albany NY) 11:303–327. https://doi.org/10.18632/aging.101684
Kurki MI, Karjalainen J, Palta P et al (2023) FinnGen provides genetic insights from a well-phenotyped isolated population. Nature 613:508–518. https://doi.org/10.1038/s41586-022-05473-8
Larsson SC (2021) Mendelian randomization as a tool for causal inference in human nutrition and metabolism. Curr Opin Lipidol 32:1–8. https://doi.org/10.1097/MOL.0000000000000721
Durbin RM, Altshuler D, Durbin RM et al (2010) A map of human genome variation from population-scale sequencing. Nature 467:1061–1073. https://doi.org/10.1038/nature09534
Staley JR, Blackshaw J, Kamat MA et al (2016) PhenoScanner: a database of human genotype-phenotype associations. Bioinformatics 32:3207–3209. https://doi.org/10.1093/bioinformatics/btw373
Kamat MA, Blackshaw JA, Young R et al (2019) PhenoScanner V2: an expanded tool for searching human genotype-phenotype associations. Bioinformatics 35:4851–4853. https://doi.org/10.1093/bioinformatics/btz469
Bowden J, Spiller W, Del Greco MF et al (2018) Improving the visualization, interpretation and analysis of two-sample summary data Mendelian randomization via the radial plot and radial regression. Int J Epidemiol 47:1264–1278. https://doi.org/10.1093/ije/dyy101
Verbanck M, Chen C-Y, Neale B et al (2018) Detection of widespread horizontal pleiotropy in causal relationships inferred from Mendelian randomization between complex traits and diseases. Nat Genet 50:693–698. https://doi.org/10.1038/s41588-018-0099-7
Zhao Q, Chen Y, Wang J et al (2019) Powerful three-sample genome-wide design and robust statistical inference in summary-data Mendelian randomization. Int J Epidemiol 48:1478–1492. https://doi.org/10.1093/ije/dyz142
Burgess S, Thompson SG (2017) Interpreting findings from Mendelian randomization using the MR-Egger method. Eur J Epidemiol 32:377–389. https://doi.org/10.1007/s10654-017-0255-x
Burgess S, Butterworth A, Thompson SG (2013) Mendelian randomization analysis with multiple genetic variants using summarized data. Genet Epidemiol 37:658–665. https://doi.org/10.1002/gepi.21758
Bowden J, Davey Smith G, Burgess S (2015) Mendelian randomization with invalid instruments: effect estimation and bias detection through Egger regression. Int J Epidemiol 44:512–525. https://doi.org/10.1093/ije/dyv080
Zhu Z, Zheng Z, Zhang F et al (2018) Causal associations between risk factors and common diseases inferred from GWAS summary data. Nat Commun 9:224. https://doi.org/10.1038/s41467-017-02317-2
Greco MFD, Minelli C, Sheehan NA et al (2015) Detecting pleiotropy in Mendelian randomisation studies with summary data and a continuous outcome. Stat Med 34:2926–2940. https://doi.org/10.1002/sim.6522
Pierce BL, Ahsan H, Vanderweele TJ (2011) Power and instrument strength requirements for Mendelian randomization studies using multiple genetic variants. Int J Epidemiol 40:740–752. https://doi.org/10.1093/ije/dyq151
Hemani G, Tilling K, Davey Smith G (2017) Orienting the causal relationship between imprecisely measured traits using GWAS summary data. PLoS Genet 13:e1007081. https://doi.org/10.1371/journal.pgen.1007081
Simon NM, Smoller JW, McNamara KL et al (2006) Telomere shortening and mood disorders: preliminary support for a chronic stress model of accelerated aging. Biol Psychiatry 60:432–435. https://doi.org/10.1016/j.biopsych.2006.02.004
Hartmann N, Boehner M, Groenen F et al (2010) Telomere length of patients with major depression is shortened but independent from therapy and severity of the disease. Depress Anxiety 27:1111–1116. https://doi.org/10.1002/da.20749
Lung F-W, Chen NC, Shu B-C (2007) Genetic pathway of major depressive disorder in shortening telomeric length. Psychiatr Genet 17:195–199. https://doi.org/10.1097/YPG.0b013e32808374f6
Wikgren M, Maripuu M, Karlsson T et al (2012) Short telomeres in depression and the general population are associated with a hypocortisolemic state. Biol Psychiatry 71:294–300. https://doi.org/10.1016/j.biopsych.2011.09.015
Phillips AC, Robertson T, Carroll D et al (2013) Do symptoms of depression predict telomere length? Evidence from the west of Scotland twenty-07 study. Psychosom Med 75:288–296. https://doi.org/10.1097/PSY.0b013e318289e6b5
Needham BL, Mezuk B, Bareis N et al (2015) Depression, anxiety and telomere length in young adults: evidence from the National Health and Nutrition Examination Survey. Mol Psychiatry 20:520–528. https://doi.org/10.1038/mp.2014.89
Shaffer JA, Epel E, Kang MS et al (2012) Depressive symptoms are not associated with leukocyte telomere length: findings from the Nova Scotia Health Survey (NSHS95), a population-based study. PLoS ONE 7:e48318. https://doi.org/10.1371/journal.pone.0048318
Liu Z, Leung D, Thrush K et al (2020) Underlying features of epigenetic aging clocks in vivo and in vitro. Aging Cell 19:e13229. https://doi.org/10.1111/acel.13229
Lu AT, Seeboth A, Tsai P-C et al (2019) DNA methylation-based estimator of telomere length. Aging (Albany NY) 11:5895–5923. https://doi.org/10.18632/aging.102173
Borges MK, Romanini CV, Lima NA et al (2021) Longitudinal association between late-life depression (LLD) and frailty: findings from a prospective cohort study (MiMiCS-FRAIL). J Nutr Health Aging 25:895–902. https://doi.org/10.1007/s12603-021-1639-x
Liu H, Li D, Zhao X et al (2021) Longitudinal impact of frailty states and sleep duration on subsequent depressive symptoms of older adults. J Am Geriatr Soc 69:1003–1011. https://doi.org/10.1111/jgs.16999
Borges MK, Aprahamian I, Romanini CV et al (2021) Depression as a determinant of frailty in late life. Aging Ment Health 25:2279–2285. https://doi.org/10.1080/13607863.2020.1857689
Dapp U, Minder CE, Golgert S et al (2021) The inter-relationship between depressed mood, functional decline and disability over a 10-year observational period within the Longitudinal Urban Cohort Ageing Study (LUCAS). J Epidemiol Community Health 75:450–457. https://doi.org/10.1136/jech-2020-214168
Oude Voshaar RC, Dimitriadis M, van den Brink RHS et al (2021) A 6-year prospective clinical cohort study on the bidirectional association between frailty and depressive disorder. Int J Geriatr Psychiatry 36:1699–1707. https://doi.org/10.1002/gps.5588
Marques A, Gaspar de Matos M, Henriques-Neto D et al (2020) Grip strength and depression symptoms among middle-age and older adults. Mayo Clin Proc 95:2134–2143. https://doi.org/10.1016/j.mayocp.2020.02.035
Cabanas-Sánchez V, Esteban-Cornejo I, Parra-Soto S et al (2022) Muscle strength and incidence of depression and anxiety: findings from the UK Biobank prospective cohort study. J Cachexia Sarcopenia Muscle 13:1983–1994. https://doi.org/10.1002/jcsm.12963
Gu Y, Li X, Zhang Q et al (2021) Grip strength and depressive symptoms in a large-scale adult population: The TCLSIH cohort study. J Affect Disord 279:222–228. https://doi.org/10.1016/j.jad.2020.08.023
Jiang R, Westwater ML, Noble S et al (2022) Associations between grip strength, brain structure, and mental health in > 40,000 participants from the UK Biobank. BMC Med 20:286. https://doi.org/10.1186/s12916-022-02490-2
Acknowledgements
The authors acknowledged The FinnGen consortium for contributing the data used in this work. We thank all the genetics consortiums for making the GWAS summary data publicly available.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
LL and ZR conceived, initiated, and supervised the project. XL collected and analyzed the data, and wrote a draft of the manuscript. JH assisted in reviewing the relevant manuscripts. The authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflicts of interest.
Ethics approval and consent to participate
We confirm that we have read the Journal’s position on issues involved in ethical publication and affirm that this report is consistent with those guidelines.
Statement of human and animal rights
This study uses publicly available data, therefore no additional ethical declaration is required. All of the studies and consortia accessed in the present study were approved by their respective ethics committee.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Luo, X., Ruan, Z. & Liu, L. Causal relationship between depression and aging: a bidirectional two-sample Mendelian randomization study. Aging Clin Exp Res 35, 3179–3187 (2023). https://doi.org/10.1007/s40520-023-02596-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s40520-023-02596-4