Abstract
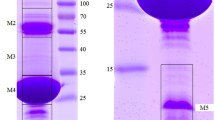
Phage (viral) tailspike proteins (TSP) are used in many translation applications such as in pathogenic bacteria detection and therapeutics. Especially, bacterial virus (or bacteriophage) P22 TSP is one of the most popular viral proteins in molecular virology and is very well-studied in terms of structure, interactions with O-antigen (a component of bacterial cell wall), and various aspects of its functions using X-ray crystallography and in vitro and in vivo studies. Although there are many papers on the virological and structural aspects of this protein, basic lab-based methodological studies and its solution-based characterizations using an in-house affordable experimental method which can be easily followed in a laboratory at an undergraduate or graduate level are missing in the literature. Our idea here is to provide a detailed method for preparing P22 TSP in a laboratory environment and studying its basic characterizations in the solution state using the available methods in our laboratory. These methods include SDS-PAGE and Native-PAGE for molecular weight determination, as well as specificity and affinity testing against its host bacterium. Our studies using SDS-PAGE show a band for TSP in the monomeric state at around 72 kDa, and the Native-PAGE proves the trimeric state with a size of around 250 kDa, thereby confirming the information regarding molar mass. On the other side, UV-Vis spectroscopy verifies absorption properties that lie around 250–300 nm. Furthermore, the solution structures of proteins in the native state, determined using dynamic light scattering (DLS), exhibit various size distributions around 10 nm, 100 nm, and 1000 nm, indicating the coexistence of trimeric and multimeric states, along with a small number of aggregates. Finally, the affinity of the TSP with the O-antigen of the host bacterium has been confirmed through western blotting studies, and the results are discussed in the context of the role of different intermolecular interactions involved in the binding of the TSP and the O-antigen group using molecular docking approach. We strongly believe our basic studies will be very helpful for beginners and graduate students in the related field.















Similar content being viewed by others
Data Availability
On request, the experimental data related to experiments can be provided.
References
Ackermann, H. W. (2009). Phage classification and characterization. Methods in Molecular Biology, 501, 127–40.
Seckler, R. (1998). Folding and function of repetitive structure in the homotrimeric phage P22 tailspike protein. Journal of Structural Biology, 122(1–2), 216–22.
Steinbacher, S., Baxa, U., Miller, S., Weintraub, A., Seckler, R., & Huber, R. (1996). Crystal structure of phage P22 tailspike protein complexed with Salmonella sp. O-antigen receptors. Proceedings of the National Academy of Sciences, 93(20), 10584–8.
Carbonell, X., & Villaverde, A. (1998). Unfolding of bacteriophage P22 tailspike protein: Enhanced thermal stability of an N-terminal fusion mutant. FEBS Letters, 432(3), 228–30.
Chen, B. L., & King, J. (1991). Thermal unfolding pathway for the thermostable P22 tailspike endorhamnosidase. Biochemistry, 30(25), 6260–9.
Maurides, P. A., Schwarz, J. J., & Berget, P. B. (1990). Intragenic suppression of a capsid assembly-defective P22 tailspike mutation. Genetics, 125(4), 673–81.
Dhakal, J., Sharma, C. S., Nannapaneni, R., McDANIEL, C. D., Kim, T., & Kiess, A. (2019). Effect of chlorine-induced sublethal oxidative stress on the biofilm-forming ability of Salmonella at different temperatures, nutrient conditions, and substrates. Journal of Food Protection, 82(1), 78–92.
Jajere, S. M. (2019). A review of Salmonella enterica with particular focus on the pathogenicity and virulence factors, host specificity and antimicrobial resistance including multidrug resistance. Veterinary World, 12(4), 504.
Balasubramanian, R., Im, J., Lee, J.-S., Jeon, H. J., Mogeni, O. D., Kim, J. H., et al. (2019). The global burden and epidemiology of invasive non-typhoidal salmonella infections. Human Vaccines & Immunotherapeutics, 15(6), 1421–6.
Stanaway, J. D., Parisi, A., Sarkar, K., Blacker, B. F., Reiner, R. C., Hay, S. I., et al. (2019). The global burden of non-typhoidal salmonella invasive disease: A systematic analysis for the Global Burden of Disease Study 2017. The Lancet Infectious Diseases, 19(12), 1312–24.
Lin, Derek M., & KoskellaLin, B. H. C. (2017). Phage therapy: An alternative to antibiotics in the age of multi-drug resistance. World Journal of Gastrointestinal Pharmacology and Therapeutics, 8, 162.
Steinbacher, S., Miller, S., Baxa, U., Budisa, N., Weintraub, A., Seckler, R., et al. (1997). Phage P22 tailspike protein: Crystal structure of the head-binding domain at 2.3 Å, fully refined structure of the endorhamnosidase at 1.56 Å resolution, and the molecular basis of O-antigen recognition and cleavage. Journal of Molecular Biology, 267(4), 865–80.
Steinbacher, S., Miller, S., Baxa, U., Weintraub, A., & Seckler, R. (1997). Interaction of Salmonella phage P22 with its O-antigen receptor studied by X-ray crystallography. Biological Chemistry, 378(3–4), 337–44.
Freiberg, A., Morona, R., Van Den Bosch, L., Jung, C., Behlke, J., Carlin, N., et al. (2003). The tailspike protein of Shigella phage Sf6 a structural homolog of Salmonella phage P22 tailspike protein without sequence similarity in the β-helix domain. Journal of Biological Chemistry, 278(3), 1542–8.
Barbirz, S., Müller, J. J., Uetrecht, C., Clark, A. J., Heinemann, U., & Seckler, R. (2008). Crystal structure of Escherichia coli phage HK620 tailspike: Podoviral tailspike endoglycosidase modules are evolutionarily related. Molecular Microbiology, 69(2), 303–16.
Walter, M., Fiedler, C., Grassl, R., Biebl, M., Rachel, R., Hermo-Parrado, X. L., et al. (2008). Structure of the receptor-binding protein of bacteriophage Det7: A podoviral tail spike in a myovirus. Journal of Virology, 82(5), 2265–73.
Müller, J. J., Barbirz, S., Heinle, K., Freiberg, A., Seckler, R., & Heinemann, U. (2008). An intersubunit active site between supercoiled parallel β helices in the trimeric tailspike endorhamnosidase of Shigella flexneri Phage Sf6. Structure, 16(5), 766–75.
Barbirz, S., Becker, M., Freiberg, A., & Seckler, R. (2009). Phage tailspike proteins with β-solenoid fold as thermostable carbohydrate binding materials. Macromolecular Bioscience, 9(2), 169–73.
Bacteriophages. (1959). von M. H. Adams. Interscience Publishers Inc.
Vinga, I., São-José, C., Tavares, P., & Santos, M. (2006). Bacteriophage entry in the host cell. Modern Bacteriophage Biology and Biotechnology, Research Signpost, 81-308-0033-0. ⟨hal-02814363⟩
Vinga, I., Baptista, C., Auzat, I., Petipas, I., Lurz, R., Tavares, P., et al. (2012). Role of bacteriophage SPP1 tail spike protein gp21 on host cell receptor binding and trigger of phage DNA ejection. Molecular Microbiology, 83(2), 289–303.
Gildea, L., Ayariga, J. A., Robertson, B. K., & Villafane, R. (2022). P22 phage shows promising antibacterial activity under pathophysiological conditions. Archives of Microbiology & Immunology, 6, 81.
Waseh, S., Hanifi-Moghaddam, P., Coleman, R., Masotti, M., Ryan, S., Foss, M., MacKenzie, R., Henry, M., Szymanski, C. M., & Tanha, J. (2010). Orally administered p22 phage tailspike protein reduces salmonella colonization in chickens: Prospects of a novel therapy against bacterial infections. PLOS ONE, 5, e13904.
González-Mora, A., Hernández-Pérez, J., Iqbal, Hafiz M. N., Rito-Palomares, M., & Benavides, J. (2020). Bacteriophage-based vaccines: A potent approach for antigen delivery. Vaccines (Basel), 8, 504.
Mahari, S., & Gandhi, S. (2022). Recent advances in electrochemical biosensors for the detection of Salmonellosis: Current prospective and challenges. Biosensors (Basel), 12, 365.
Simmons, M., Donovan, D. M., Siragusa, G. R., & Seal, B. S. (2010). Recombinant expression of two bacteriophage proteins that lyse clostridium perfringens and share identical sequences in the c-terminal cell wall binding domain of the molecules but are dissimilar in their n-terminal active domains. Journal of Agricultural and Food Chemistry, 58(19), 10330–7.
Lee, I.-M., Tu, I.-F., Yang, F.-L., Ko, T.-P., Liao, J.-H., Lin, N.-T., et al. (2017). Structural basis for fragmenting the exopolysaccharide of Acinetobacter baumannii by bacteriophage ΦAB6 tailspike protein. Scientific Reports, 7, 42711.
Kruger, N. J. (2009). The Bradford method for protein quantitation. The protein protocols handbook (pp. 17–24). Springer.
Sambrook, J., & Russell, D. W. (2006). SDS-polyacrylamide gel electrophoresis of proteins. CSH Protocols, 2006(4), pdb-prot4540.
Arndt, C., Koristka, S., Bartsch, H., & Bachmann, M. (2012). Native polyacrylamide gels. Protein electrophoresis (pp. 49–53). Springer.
Biswal, S., Rizwan, H., Pal, S., Sabnam, S., Parida, P., & Pal, A. (2019). Oxidative stress, antioxidant capacity, biomolecule damage, and inflammation symptoms of sickle cell disease in children. Hematology, 24(1), 1–9.
Origin(Pro), Version 8.5, OriginLab Corporation, Northampton, MA, USA.
Rosano, G. L., & Ceccarelli, E. A. (2014). Recombinant protein expression in Escherichia coli: Advances and challenges. Frontiers in Microbiology, 5, 172.
Pan, S., & Malcolm, B. A. (2000). Reduced background expression and improved plasmid stability with pET vectors in BL21 (DE3). Biotechniques, 29(6), 1234–8.
Sudha, A., Srinivasan, P., Thamilarasan, V., & Sengottuvelan, N. (2016). Exploring the binding mechanism of 5-hydroxy-3′,4′,7-trimethoxyflavone with bovine serum albumin: Spectroscopic and computational approach. Spectrochimica Acta Part A: Molecular and Biomolecular Spectroscopy, 157, 170.
Rial, R., González-Durruthy, M., Somoza, M., Liu, Z., & Ruso, J. M. (2021). Unraveling the compositional and molecular features involved in lysozyme-benzothiazole derivative interactions. Molecules, 26, 5855.
He, M., & Herr, A. E. (2010). Polyacrylamide gel photopatterning enables automated protein immunoblotting in a two-dimensional microdevice. Journal of the American Chemical Society, 132(8), 2512–3.
Murphy, R. M., & Lamb, G. D. (2013). Important considerations for protein analyses using antibody based techniques: Down-sizing Western blotting up-sizes outcomes. The Journal of Physiology, 591(23), 5823–31.
Kurien, B. T., & Scofield, R. H. (2015). Other notable protein blotting methods: A brief review. Western Blotting (pp. 487–503). Springer.
Mishra, M., Tiwari, S., & Gomes, A. V. (2017). Protein purification and analysis: Next generation Western blotting techniques. Expert Review of Proteomics, 14(11), 1037–53.
Counts, S. (2010). Western Blot. In Elsevier eBooks (pp. 323–326).
Jensen, E. C. (2012). The basics of western blotting. The Anatomical Record: Advances in Integrative Anatomy and Evolutionary Biology, 295(3), 369–71.
Lorber, B., Fischer, F., Bailly, M., Roy, H., & Kern, D. (2012). Protein analysis by dynamic light scattering: Methods and techniques for students. Biochemistry and Molecular Biology Education, 40(6), 372–82.
Schmidt, R. (2010). Dynamic light scattering for protein characterization. R. A. Meyers (Eds.), In Encyclopedia of Analytical Chemistry. https://doi.org/10.1002/9780470027318.a9092.
Venkatesan, K. Williams, J. & Villafane, R. (2014) Non-stem amino acids are involved in the phage P22 TSP NTD stability. Advances in Microbiology, 4, 521–526.
Kim, J., & Robinson, A. S. (2006). Dissociation of intermolecular disulfide bonds in P22 tailspike protein intermediates in the presence of SDS. Protein Science, 15(7), 1791–3.
Webber, T., Gurung, S., Saul, J., Baker, T., Spatara, M., Freyer, M., et al. (2009). The C-terminus of the P22 tailspike protein acts as an independent oligomerization domain for monomeric proteins. The Biochemical Journal, 419(3), 595–602.
Fischer, H., Polikarpov, I., & Craievich, A. F. (2004). Average protein density is a molecular-weight-dependent function. Protein Science, 13, 2825.
Dutt, S., Tanha, J., Evoy, S., & Singh, A. (2013). Immobilization of P22 bacteriophage Tailspike protein on Si surface for optimized Salmonella capture. Journal of Analytical & Bioanalytical Techniques Science, 7, 7.
Landström, J., Nordmark, E.-L., Eklund, R., Weintraub, A., Seckler, R., & Widmalm, G. (2008). Interaction of a Salmonella enteritidis O-antigen octasaccharide with the phage P22 tailspike protein by NMR spectroscopy and docking studies. Glycoconjugate Journal, 25(2), 137–43.
Handa, H., Gurczynski, S., Jackson, M. P., Auner, G., Walker, J., & Mao, G. (2008). Recognition of Salmonella typhimurium by immobilized phage P22 monolayers. Surface Science, 602(7), 1392–400.
Hyeon, S. H., Lim, W. K., & Shin, H. J. (2021). Novel surface plasmon resonance biosensor that uses full-length Det7 phage tail protein for rapid and selective detection of salmonella enterica serovar typhimurium. Biotechnology and applied biochemistry, 68(1), 5–12.
Singh, A., Arya, S. K., Glass, N., Hanifi-Moghaddam, P., Naidoo, R., Szymanski, C. M., et al. (2010). Bacteriophage tailspike proteins as molecular probes for sensitive and selective bacterial detection. Biosens Bioelectron [Internet], 26(1), 131–8. https://doi.org/10.1016/j.bios.2010.05.024. Available from.
Doulah, M. S. (1977). Mechanism of disintegration of biological cells in ultrasonic cavitation. Biotechnology and Bioengineering, 19(5), 649–60.
Feliu, J. X., Cubarsi, R., & Villaverde, A. (1998). Optimized release of recombinant proteins by ultrasonication of E. coli cells. Biotechnology and Bioengineering, 58(5), 536–40.
Steinbacher, S., Seckler, R., Miller, S., Steipe, B., Huber, R., & Reinemer, P. (1994). Crystal structure of P22 tailspike protein: Interdigitated subunits in a thermostable trimer. Science, 265(5170), 383–6.
Andres, D., Baxa, U., Hanke, C., Seckler, R., & Barbirz, S. (2010). Carbohydrate binding of Salmonella phage P22 tailspike protein and its role during host cell infection. Biochemical Society Transactions, 38(5), 1386–9.
Seul, A., Müller, J. J., Andres, D., Stettner, E., Heinemann, U., & Seckler, R. (2014). Bacteriophage P22 tailspike: Structure of the complete protein and function of the interdomain linker. Acta Crystallographica. Section D, Biological Crystallography, 70(Pt 5), 1336–45.
Waseh, S., Hanifi-Moghaddam, P., Coleman, R., Masotti, M., Ryan, S., Foss, M., MacKenzie, R., Henry, M., Szymanski, C. M., & Tanha, J. (2010). Orally administered P22 phage tailspike protein reduces salmonella colonization in chickens: Prospects of a novel therapy against bacterial infections. PLoS One, 5(11), e13904.
Goldenberg, D. P., & Creighton, T. E. (1994). Phage tailspike protein. A fishy tale of protein folding. Current Biology, 4(11), 1026–9.
Andres, D., Gohlke, U., Broeker, N. K., Schulze, S., Rabsch, W., Heinemann, U., Barbirz, S., & Seckler, R. (2013). An essential serotype recognition pocket on phage P22 tailspike protein forces Salmonella enterica serovar Paratyphi A O-antigen fragments to bind as nonsolution conformers. Glycobiology, 23(4), 486–94.
Morris, G. M., Huey, R., Lindstrom, W., Sanner, M. F., Belew, R. K., Goodsell, D. S., et al. (2009). AutoDock4 and AutoDockTools4: Automated docking with selective receptor flexibility. Journal of Computational Chemistry, 30(16), 2785–91.
Forli, S., Huey, R., Pique, M. E., Sanner, M. F., Goodsell, D. S., & Olson, A. J. (2016). Computational protein–ligand docking and virtual drug screening with the AutoDock suite. Nature Protocols, 11(5), 905–19.
Goodsell, D. S., & Olson, A. J. (1990). Automated docking of substrates to proteins by simulated annealing. Proteins: Structure, Function, and Bioinformatics, 8(3), 195–202.
Morris, G. M., Goodsell, D. S., Halliday, R. S., Huey, R., Hart, W. E., Belew, R. K., et al. (1998). Automated docking using a Lamarckian genetic algorithm and an empirical binding free energy function. Journal of Computational Chemistry, 19(14), 1639–62.
Goodford, P. J. (1985). A computational procedure for determining energetically favorable binding sites on biologically important macromolecules. Journal of Medicinal Chemistry, 28(7), 849–57.
Mehler, E. L., & Solmajer, T. (1991). Electrostatic effects in proteins: Comparison of dielectric and charge models. Protein Engineering, Design & Selection, 4(8), 903–10.
Morris, G. M., Goodsell, D. S., Huey, R., & Olson, A. J. (1996). Distributed automated docking of flexible ligands to proteins: Parallel applications of AutoDock 2.4. Journal of Computer-Aided Molecular Design, 10(4), 293–304.
Böhm, H.-J. (1994). The development of a simple empirical scoring function to estimate the binding constant for a protein-ligand complex of known three-dimensional structure. Journal of Computer-Aided Molecular Design, 8(3), 243–56.
Mohanty, M., & Mohanty, P. S. (2023). Molecular docking in organic, inorganic, and hybrid systems: a tutorial review. Monatshefte fur chemie, 1–25. https://doi.org/10.1007/s00706-023-03076-1
DeLano, W. L. (2002). The PyMOL molecular graphics system. Delano Scientific.
Acknowledgements
PSM acknowledges BIRAC, Center of Excellence project, Department of Biotechnology, Govt. Of India , Depart of Biotechnology and ICMR (Indian Council of Medical Research), Government of India, for support in the current work.
Author information
Authors and Affiliations
Contributions
PSM: principal investigator of the work, conceptualization of the problem, formulation of paper, methodology, carried out some parts of experiments, critical analysis and interpretation of all experiments.
Manoswini: experimental work, synthesized the proteins, characterized and prepared the original draft in the protein study.
MM: theoretical understanding and in silico molecular docking simulation study
AGM: referencing, correction, editing,
BRS: supervision and useful suggestion in the discussion.
The entire manuscript has undergone a comprehensive revision by MM. AGM assisted in referencing and some editing work in the revised manuscript. This revision included the correction of English grammar throughout the manuscript, restructuring of sentences and paragraphs, detailed docking methodology, and studies. Additionally, figures were revised to enhance resolution, and references were updated
Corresponding author
Ethics declarations
Ethical Approval
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Manoswini, M., Mohanty, M., Majumdar, A.G. et al. Extraction and Characterizations of Viral Protein Particles: A Methodological Study. BioNanoSci. (2024). https://doi.org/10.1007/s12668-023-01289-6
Accepted:
Published:
DOI: https://doi.org/10.1007/s12668-023-01289-6