Abstract
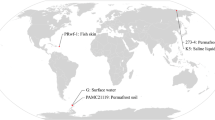
Subtercola boreus K300T is a novel psychrophilic strain that was isolated from permanently cold groundwater in Finland and has also been found in several places in Antarctica including lake, soil, and rocks. We performed genomic and transcriptomic analyses of 5 strains from Antarctica and a type strain to understand their adaptation to different environments. Interestingly, the isolates from rocks showed a low growth rate and smaller genome size than strains from the other isolation sources (lake, soil, and groundwater). Based on these habitat-dependent characteristics, the strains could be classified into two ecotypes, which showed differences in energy production, signal transduction, and transcription in the clusters of orthologous groups of proteins (COGs) functional category. In addition, expression pattern changes revealed differences in metabolic processes, including uric acid metabolism, DNA repair, major facilitator superfamily (MFS) transporters, and xylose degradation, depending on the nutritional status of their habitats. These findings provide crucial insights into the environmental adaptation of bacteria, highlighting genetic diversity and regulatory mechanisms that enable them to thrive in the cryosphere.






Similar content being viewed by others
Data availability
The sequences of 6 strains reported in this paper have been deposited in the GenBank database (BioProject no. PRJNA378458).
References
Aguilar, P. S., Cronan, J. E., Jr., & de Mendoza, D. (1998). A Bacillus subtilis gene induced by cold shock encodes a membrane phospholipid desaturase. Journal of Bacteriology, 180, 2194–2200.
Baker-Austin, C., & Dopson, M. (2007). Life in acid: PH homeostasis in acidophiles. Trends in Microbiology, 15, 165–171.
Bankevich, A., Nurk, S., Antipov, D., Gurevich, A. A., Dvorkin, M., Kulikov, A. S., Lesin, V. M., Nikolenko, S. I., Pham, S., Prjibelski, A. D., et al. (2012). SPAdes: A new genome assembly algorithm and its applications to single-cell sequencing. Journal of Computational Biology, 19, 455–477.
Baraúna, R. A., Freitas, D. Y., Pinheiro, J. C., Folador, A. R., & Silva, A. (2017). A proteomic perspective on the bacterial adaptation to cold: Integrating OMICs data of the psychrotrophic bacterium Exiguobacterium antarcticum B7. Proteomes, 5, 9.
Barria, C., Malecki, M., & Arraiano, C. M. (2013). Bacterial adaptation to cold. Microbiology, 159, 2437–2443.
Bolger, A. M., Lohse, M., & Usadel, B. (2014). Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics, 30, 2114–2120.
Burger, A. E., Lindeboom, H. J., & Williams, A. J. (1971). The mineral and energy contributions of guano of selected species of birds to the Marion Island terrestrial ecosystem. South African Journal of Antarctic Research, 8, 59–70.
Cary, S. C., McDonald, I. R., Barrett, J. E., & Cowan, D. A. (2010). On the rocks: The microbiology of Antarctic Dry Valley soils. Nature Reviews Microbiology, 8, 129–138.
Cordero, O. X., & Hogeweg, P. (2009). The impact of long-distance horizontal gene transfer on prokaryotic genome size. Proceedings of the National Academy of Sciences, 106, 21748–21753.
Deming, J. W. (2002). Psychrophiles and polar regions. Current Opinion in Microbiology, 5, 301–309.
Ding, W., Baumdicker, F., & Neher, R. A. (2018). panX: Pan-genome analysis and exploration. Nucleic Acids Research, 46, e5.
Emms, D. M., & Kelly, S. (2015). OrthoFinder: Solving fundamental biases in whole genome comparisons dramatically improves orthogroup inference accuracy. Genome Biology, 16, 157.
Garrity, G. (2007). Bergey’s manual® of Systematic bacteriology: volume 2: the proteobacteria, part B: the Gammaproteobacteria. New York: Springer.
Gianese, G., Argos, P., & Pascarella, S. (2001). Structural adaptation of enzymes to low temperatures. Protein Engneering, 14, 141–148.
Hammann, R., & Kutzner, H. J. (1998). Key enzymes for the degradation of benzoate, m- and p-hydroxybenzoate by some members of the order Actinomycetales. Journal of Basic Microbiology, 38, 207–220.
Hessen, D. O., Jeyasingh, P. D., Neiman, M., & Weider, L. J. (2010). Genome streamlining and the elemental costs of growth. Trends in Ecology & Evolution, 25, 75–80.
Horikoshi, K. (1999). Alkaliphiles: Some applications of their products for biotechnology. Microbiology and Molecular Biology Reviews, 63, 735–750.
Hunt, M., Silva, N. D., Otto, T. D., Parkhill, J., Keane, J. A., & Harris, S. R. (2015). Circlator: Automated circularization of genome assemblies using long sequencing reads. Genome Biology, 16, 294.
Jaenicke, R., & Bohm, G. (1998). The stability of proteins in extreme environments. Current Opinion in Structural Biology, 8, 738–748.
Keto-Timonen, R., Hietala, N., Palonen, E., Hakakorpi, A., Lindstrom, M., & Korkeala, H. (2016). Cold shock proteins: A minireview with special emphasis on Csp-family of enteropathogenic Yersinia. Frontiers in Microbiology, 7, 1151.
Kim, H., Lee, J. Y., & Lee, K. K. (2014). Thermal characteristics and bacterial diversity of forest soil in the Haean basin of Korea. Scientific World Journal, 2014, 247401.
Kim, J. H., Ahn, I. Y., Lee, K. S., Chung, H. S., & Choi, H. G. (2007). Vegetation of Barton Peninsula in the neighbourhood of King Sejong Station (King George Island, maritime Antarctic). Polar Biology, 30, 903–916.
Kim, O. S., Chae, N., Lim, H. S., Cho, A., Kim, J. H., Hong, S. G., & Oh, J. (2012a). Bacterial diversity in ornithogenic soils compared to mineral soils on King George Island, Antarctica. Journal of Microbiology, 50, 1081–1085.
Kim, O. S., Cho, Y. J., Lee, K., Yoon, S. H., Kim, M., Na, H., Park, S. C., Jeon, Y. S., Lee, J. H., Yi, H., et al. (2012b). Introducing EzTaxon-e: A prokaryotic 16S rRNA gene sequence database with phylotypes that represent uncultured species. International Journal of Systematic and Evolutionary Microbiology, 62, 716–721.
Kumar, A., Alam, A., Tripathi, D., Rani, M., Khatoon, H., Pandey, S., Ehtesham, N. Z., & Hasnain, S. E. (2018). Protein adaptations in extremophiles: An insight into extremophilic connection of mycobacterial proteome. Seminars in Cell & Developmental Biology, 84, 147–157.
Kumar, S., Stecher, G., & Tamura, K. (2016). MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Molecular Biology and Evolution, 33, 1870–1874.
Kurokawa, M., Seno, S., Matsuda, H., & Ying, B. W. (2016). Correlation between genome reduction and bacterial growth. DNA Research, 23, 517–525.
Langmead, B., & Salzberg, S. L. (2012). Fast gapped-read alignment with Bowtie 2. Nature Methods, 9, 357–359.
Lee, J., Cho, J., Cho, Y. J., Cho, A., Woo, J., Lee, J., Hong, S. G., Sul, W. J., & Kim, O. S. (2019). The latitudinal gradient in rock-inhabiting bacterial community compositions in Victoria Land, Antarctica. Science of the Total Environment, 657, 731–738.
Lee, Y. M., Kim, G., Jung, Y. J., Choe, C. D., Yim, J. H., Lee, H. K., & Hong, S. G. (2012). Polar and alpine microbial collection (PAMC): A culture collection dedicated to polar and alpine microorganisms. Polar Biology, 35, 1433–1438.
Liao, Y., Smyth, G. K., & Shi, W. (2013). The Subread aligner: Fast, accurate and scalable read mapping by seed-and-vote. Nucleic Acids Research, 41, e108.
Love, M. I., Huber, W., & Anders, S. (2014). Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biology, 15, 550.
Männistö, M., Schumann, P., Rainey, F., Kämpfer, P., Tsitko, I., Tiirola, M., & Salkinoja-Salonen, M. (2000). Subtercola boreus gen. nov., sp. nov. and Subtercola frigoramans sp. nov., two new psychrophilic actinobacteria isolated from boreal groundwater. International Journal of Systematic and Evolutionary Microbiology, 50, 1731–1739.
Margesin, R., & Miteva, V. (2011). Diversity and ecology of psychrophilic microorganisms. Research in Microbiology, 162, 346–361.
Markkula, A., Lindström, M., Johansson, P., Björkroth, J., & Korkeala, H. (2012). Roles of four putative DEAD-box RNA helicase genes in growth of Listeria monocytogenes EGD-e under heat, pH, osmotic, ethanol, and oxidative stress conditions. Applied and Environmental Microbiology, 78, 6875–6882.
Mocali, S., Chiellini, C., Fabiani, A., Decuzzi, S., de Pascale, D., Parrilli, E., Tutino, M. L., Perrin, E., Bosi, E., Fondi, M., et al. (2017). Ecology of cold environments: New insights of bacterial metabolic adaptation through an integrated genomic-phenomic approach. Scientific Reports, 7, 839.
Qin, Q. L., Xie, B. B., Yu, Y., Shu, Y. L., Rong, J. C., Zhang, Y. J., Zhao, D. L., Chen, X. L., Zhang, X. Y., Chen, B., et al. (2014). Comparative genomics of the marine bacterial genus Glaciecola reveals the high degree of genomic diversity and genomic characteristic for cold adaptation. Environmental Microbiology, 16, 1642–1653.
Ribaudo, R., Gilman, M., Kingston, R. E., Choczynski, P., & Sacchi, N. (2001). Preparation of RNA from tissues and cells. Current Protocols in Immunology. https://doi.org/10.1002/0471142735.im1011s04
Richter, M., Rosselló-Móra, R., Oliver Glöckner, F., & Peplies, J. (2016). JSpeciesWS: A web server for prokaryotic species circumscription based on pairwise genome comparison. Bioinformatics, 32, 929–931.
Si, H. L., Shi, F. X., Zhang, L. L., Yue, H. S., Wang, H. Y., & Zhao, Z. T. (2017). Subtercola lobariae sp. nov., an actinobacterium of the family Microbacteriaceae isolated from the lichen Lobaria retigera. International Journal of Systematic and Evolutionary Microbiology, 67, 1516–1521.
Steven, B., Léveillé, R., Pollard, W. H., & Whyte, L. G. (2006). Microbial ecology and biodiversity in permafrost. Extremophiles, 10, 259–267.
Tamura, K., & Nei, M. (1993). Estimation of the number of nucleotide substitutions in the control region of mitochondrial DNA in humans and chimpanzees. Molecular Biology and Evolution, 10, 512–526.
Tatusova, T., DiCuccio, M., Badretdin, A., Chetvernin, V., Nawrocki, E. P., Zaslavsky, L., Lomsadze, A., Pruitt, K. D., Borodovsky, M., & Ostell, J. (2016). NCBI prokaryotic genome annotation pipeline. Nucleic Acids Research, 44, 6614–6624.
Treseder, K. K., & Lennonb, J. T. (2015). Fungal traits that drive ecosystem dynamics on land. Microbiology and Molecular Biology Reviews, 79, 243–262.
van de Vossenberg, J. L., Driessen, A. J., & Konings, W. N. (1998a). The essence of being extremophilic: The role of the unique archaeal membrane lipids. Extremophiles, 2, 163–170.
van de Vossenberg, J. L., Driessen, A. J., Zillig, W., & Konings, W. N. (1998b). Bioenergetics and cytoplasmic membrane stability of the extremely acidophilic, thermophilic archaeon Picrophilus oshimae. Extremophiles, 2, 67–74.
Wiegel, J., & Kevbrin, V. V. (2004). Alkalithermophiles. Biochemical Society Transactions, 32, 193–198.
Wiegeshoff, F., Beckering, C. L., Debarbouille, M., & Marahiel, M. A. (2006). Sigma L is important for cold shock adaptation of Bacillus subtilis. Journal of Bacteriology, 188, 3130–3133.
Acknowledgements
This work was supported by a Grant from the Korea Polar Research Institute (PE23130).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Lee, H., Cho, YJ., Cho, A. et al. Environmental Adaptation of Psychrophilic Bacteria Subtercola spp. Isolated from Various Cryospheric Habitats. J Microbiol. 61, 663–672 (2023). https://doi.org/10.1007/s12275-023-00068-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12275-023-00068-y