Abstract
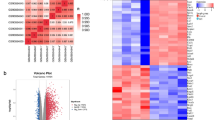
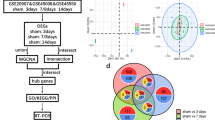
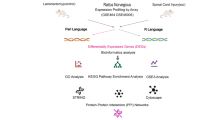
Spinal cord injury (SCI) is a serious disease without effective therapeutic strategies. To identify the potential treatments for SCI, it is extremely important to explore the underlying mechanism. Current studies demonstrate that anoikis might play an important role in SCI. In this study, we aimed to identify the key anoikis-related genes (ARGs) providing therapeutic targets for SCI. The mRNA expression matrix of GSE45006 was downloaded from the Gene Expression Omnibus (GEO) database, and the ARGs were downloaded from the Molecular Signatures Database (MSigDB database). Then, the potential differentially expressed ARGs were identified. Next, correlation analysis, gene ontology (GO) enrichment analysis and Kyoto Encyclopedia of Genes and Genomes (KEGG) pathway enrichment analysis, and protein–protein interaction (PPI) analysis were employed for the differentially expressed ARGs. Moreover, miRNA-gene networks were constructed by the hub ARGs. Finally, RNA expression of the top ten hub ARGs was validated in the SCI cell model and rat SCI model. A total of 27 common differentially expressed ARGs were identified at different time points (1, 3, 7, and 14 days) following SCI. The GO and KEGG enrichment analysis of these ARGs indicated several enriched terms related to proliferation, cell cycle, and apoptotic process. The PPI results revealed that most of the ARGs interacted with each other. Ten hub ARGs were further screened, and all the 10 genes were validated in the SCI cell model. In the rat model, only seven genes were validated eventually. We identified 27 differentially expressed ARGs of the SCI through bioinformatic analysis. Seven real hub ARGs (CCND1, FN1, IGF1, MYC, STAT3, TGFB1, and TP53) were identified eventually. These results may expand our understanding of SCI and contribute to the exploration of potential SCI targets.









Similar content being viewed by others
Data Availability
The datasets used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
References
Mark AA, Jordan WS, Matthieu G, Thomas HH, Claudia K, Quentin B, Jocelyne B, Grégoire C (2022) Natural and targeted circuit reorganization after spinal cord injury[J]. Nat Neurosci 25(12):1584–1596
Griffin JM, Bradke F (2020) Therapeutic repair for spinal cord injury: combinatory approaches to address a multifaceted problem[J]. EMBO Mol Med 12(3):e11505
Ahuja CS, Wilson JR, Nori S, Mark RNK, Claudia D, Armin C, Michael GF (2017) Traumatic spinal cord injury[J]. Nat Rev Dis Primers 3:17018
Kwon BK, Tetzlaff W, Grauer JN et al (2004) Pathophysiology and pharmacologic treatment of acute spinal cord injury[J]. Spine J 4(4):451–464
Huang H, Young W, Skaper S, Lin C, Gustavo M, Hooshang S, Ziad AZ, Hari SS et al (2020) Clinical neurorestorative therapeutic guidelines for spinal cord injury (IANR/CANR version 2019)[J]. J Orthop Translat 20:14–24
Arnold PM, Anderson PA, Chi JH, Dailey TA, Dhall SS, Eichholz KM, Harrop JS, Hoh DJ et al (2019) Congress of Neurological surgeons systematic review and evidence-based guidelines on the evaluation and treatment of patients with thoracolumbar spine trauma: pharmacological treatment[J]. Neurosurgery 84(1):E36–E38
O’Toole JE, Kaiser MG, Anderson PA, Arnold PM, Chi JH, Dailey AT, Dhall SS, Eichholz KM, Harrop JS, Hoh DJ, Qureshi S et al (2019) Congress of neurological surgeons systematic review and evidence-based guidelines on the evaluation and treatment of patients with thoracolumbar spine trauma: executive summary[J]. Neurosurgery 84(1):2–6
Vacca V, Madaro L, De Angelis F, Proietti D, Cobianchi S, Orsini T, Puri PL, Luvisetto S et al (2020) Revealing the therapeutic potential of botulinum neurotoxin type A in counteracting paralysis and neuropathic pain in spinally injured mice[J]. Toxins (Basel) 12(8):491
Hausmann ON (2003) Post-traumatic inflammation following spinal cord injury[J]. Spinal Cord 41(7):369–378
Mothe AJ, Tam RY, Zahir T, Tator CH, Shoichet MS (2013) Repair of the injured spinal cord by transplantation of neural stem cells in a hyaluronan-based hydrogel[J]. Biomaterials 34(15):3775–3783
Chen Y, Liu S, Li J et al (2020) The latest view on the mechanism of ferroptosis and its research progress in spinal cord injury[J]. Oxid Med Cell Longev 2020:6375938
Wang W, Huang X, Li J et al (2017) Methane suppresses microglial activation related to oxidative, inflammatory, and apoptotic injury during spinal cord injury in rats[J]. Oxid Med Cell Longev 2017:2190897
Fan H, Tang HB, Shan LQ, Liu SC, Huang DG, Chen X, Chen Z, Yang M et al (2019) Quercetin prevents necroptosis of oligodendrocytes by inhibiting macrophages/microglia polarization to M1 phenotype after spinal cord injury in rats[J]. J Neuroinflammation 16(1):206
Frisch SM, Francis H (1994) Disruption of epithelial cell-matrix interactions induces apoptosis[J]. J Cell Biol 124(4):619–626
Grossmann J (2002) Molecular mechanisms of “detachment-induced apoptosis–Anoikis”[J]. Apoptosis 7(3):247–260
Danial NN, Korsmeyer SJ (2004) Cell death: critical control points[J]. Cell 116(2):205–219
Gilmore AP (2005) Anoikis[J]. Cell Death Differ 12(Suppl 2):1473–1477
Paoli P, Giannoni E, Chiarugi P (2013) Anoikis molecular pathways and its role in cancer progression[J]. Biochim Biophys Acta 1833(12):3481–3498
Alvarez JI, Dodelet-Devillers A, Kebir H, Ifergan I, Fabre PJ, Terouz S, Sabbagh M, Wosik K et al (2011) The Hedgehog pathway promotes blood-brain barrier integrity and CNS immune quiescence[J]. Science 334(6063):1727–1731
Wang Y, Jin S, Sonobe Y et al (2014) Interleukin-1beta induces blood-brain barrier disruption by downregulating Sonic hedgehog in astrocytes[J]. PLoS ONE 9(10):e110024
Kapadia R, Yi JH, Vemuganti R (2008) Mechanisms of anti-inflammatory and neuroprotective actions of PPAR-gamma agonists[J]. Front Biosci 13:1813–1826
da Huang W, Sherman BT, Lempicki RA (2009) Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources[J]. Nat Protoc 4(1):44–57
Shannon P, Markiel A, Ozier O, Baliga NS, Wang JT, Ramage D, Amin N, Schwikowski B et al (2003) Cytoscape: a software environment for integrated models of biomolecular interaction networks[J]. Genome Res 13(11):2498–2504
Sticht C, De La Torre C, Parveen A et al (2018) miRWalk: an online resource for prediction of microRNA binding sites[J]. PLoS ONE 13(10):e0206239
Zhou J, Li Z, Zhao Q, Wu T, Zhao Q, Cao Y (2021) Knockdown of SNHG1 alleviates autophagy and apoptosis by regulating miR-362-3p/Jak2/stat3 pathway in LPS-injured PC12 cells[J]. Neurochem Res 46(4):945–956
Boyd JG, Lee J, Skihar V, Doucette R, Kawaja MD (2004) LacZ-expressing olfactory ensheathing cells do not associate with myelinated axons after implantation into the compressed spinal cord[J]. Proc Natl Acad Sci U S A 101(7):2162–2166
Rossignol S, Frigon A (2011) Recovery of locomotion after spinal cord injury: some facts and mechanisms[J]. Annu Rev Neurosci 34:413–440
Ray SK, Samntaray S, Banik NL (2016) Future directions for using estrogen receptor agonists in the treatment of acute and chronic spinal cord injury[J]. Neural Regen Res 11(9):1418–1419
Ahuja CS, Martin AR, Fehlings M (2016) Recent advances in managing a spinal cord injury secondary to trauma[J]. F1000Res 5:1017
Sater AP, Rael LT, Tanner AH et al (2018) Cell death after traumatic brain injury: detrimental role of anoikis in healing[J]. Clin Chim Acta 482:149–154
Warren KM, Reeves TM, Phillips LL (2012) MT5-MMP, ADAM-10, and N-cadherin act in concert to facilitate synapse reorganization after traumatic brain injury[J]. J Neurotrauma 29(10):1922–1940
Sandhir R, Gregory E, He YY et al (2011) Upregulation of inflammatory mediators in a model of chronic pain after spinal cord injury[J]. Neurochem Res 36(5):856–862
Moon C, Heo S, Sim KB, Shin T (2004) Upregulation of CD44 expression in the spinal cords of rats with clip compression injury[J]. Neurosci Lett 367(1):133–136
Juan Z, Dake C, Tanaka K, Shuixiang H (2021) EGFL7 as a novel therapeutic candidate regulates cell invasion and anoikis in colorectal cancer through PI3K/AKT signaling pathway[J]. Int J Clin Oncol 26(6):1099–1108
Gan L, Liu P, Lu H, Chen S, Yang J, McCarthy JB, Knudsen KE, Huang H (2009) Cyclin D1 promotes anchorage-independent cell survival by inhibiting FOXO-mediated anoikis[J]. Cell Death Differ 16(10):1408–1417
Pinchi E, Frati A, Cantatore S, D’Errico S, Russa RL, Maiese A, Palmieri M, Pesce A et al (2019) Acute spinal cord injury: a systematic review investigating miRNA families involved[J]. Int J Mol Sci 20(8):1841. https://doi.org/10.3390/ijms20081841
Zhang H, Wang D, Tong J et al (2022) MiR-30b-5p attenuates the inflammatory response and facilitates the functional recovery of spinal cord injury by targeting the NEFL/mTOR pathway[J]. Brain Behav 12(12):e2788
Malvandi AM, Rastegar-Moghaddam SH, Ebrahimzadeh-Bideskan S, Lombardi G, Ebrahimzadeh-Bideskan A, Mohammadipour A (2022) Targeting miR-21 in spinal cord injuries: a game-changer?[J]. Mol Med 28(1):118
Zhang Z, Wan F, Zhuang Q, Zhang Y, Xu Z (2017) Suppression of miR-127 protects PC-12 cells from LPS-induced inflammatory injury by downregulation of PDCD4[J]. Biomed Pharmacother 96:1154–1162
Wakulik K, Wiatrak B, Szczukowski L, Bodetko D, Szandruk-Bender M, Dobosz A, Świątek P, Gąsiorowski K (2020) Effect of novel pyrrolo[3,4-d]pyridazinone derivatives on lipopolysaccharide-induced neuroinflammation[J]. Int J Mol Sci 21(7):2575
Wu J, Stoica BA, Faden AI (2011) Cell cycle activation and spinal cord injury[J]. Neurotherapeutics 8(2):221–228
Byrnes KR, Stoica BA, Fricke S et al (2007) Cell cycle activation contributes to post-mitotic cell death and secondary damage after spinal cord injury[J]. Brain 130(Pt 11):2977–2992
Wu J, Stoica BA, Dinizo M, Pajoohesh-Ganji A, Piao C, Faden AI (2012) Delayed cell cycle pathway modulation facilitates recovery after spinal cord injury[J]. Cell Cycle 11(9):1782–1795
Di Giovanni S, Knoblach SM, Brandoli C, Aden SA, Hoffman EP, Faden AI (2003) Gene profiling in spinal cord injury shows role of cell cycle in neuronal death[J]. Ann Neurol 53(4):454–468
Singh P, Carraher C, Schwarzbauer JE (2010) Assembly of fibronectin extracellular matrix[J]. Annu Rev Cell Dev Biol 26:397–419
Frisch SM, Ruoslahti E (1997) Integrins and anoikis[J]. Curr Opin Cell Biol 9(5):701–706
Gonzalez-Perez F, Hernandez J, Heimann C, Phillips JB, Udina E, Navarro X (2018) Schwann cells and mesenchymal stem cells in laminin- or fibronectin-aligned matrices and regeneration across a critical size defect of 15 mm in the rat sciatic nerve[J]. J Neurosurg Spine 28(1):109–118
Faber-Elman A, Solomon A, Abraham JA, Marikovsky M, Schwartz M (1996) Involvement of wound-associated factors in rat brain astrocyte migratory response to axonal injury: in vitro simulation[J]. J Clin Invest 97(1):162–171
Bellver-Landete V, Bretheau F, Mailhot B, Vallières N, Lessard M, Janelle ME, Vernoux N, Tremblay MÈ (2019) Microglia are an essential component of the neuroprotective scar that forms after spinal cord injury[J]. Nat Commun 10(1):518
Zaytseva O, Quinn LM (2017) Controlling the master: chromatin dynamics at the MYC promoter integrate developmental signaling[J]. Genes (Basel) 8(4):118
Li N, Yao M, Liu J, Zhu Z, Lam TL, Zhang P, Kiang KMY, Leung GKK (2022) Vitamin D promotes remyelination by suppressing c-Myc and Inducing oligodendrocyte precursor cell differentiation after traumatic spinal cord injury[J]. Int J Biol Sci 18(14):5391–5404
Dominguez E, Rivat C, Pommier B, Mauborgne A, Pohl M (2008) JAK/STAT3 pathway is activated in spinal cord microglia after peripheral nerve injury and contributes to neuropathic pain development in rat[J]. J Neurochem 107(1):50–60
Ramsauer M, Krause D, Dermietzel R (2002) Angiogenesis of the blood-brain barrier in vitro and the function of cerebral pericytes[J]. FASEB J 16(10):1274–1276
Ferbert T, Child C, Graeser V, Swing T, Akbar M, Heller R, Biglari B, Moghaddam A (2017) Tracking spinal cord injury: differences in cytokine expression of IGF-1, TGF- B1, and sCD95l can be measured in blood samples and correspond to neurological remission in a 12-week follow-up[J]. J Neurotrauma 34(3):607–614
Powell E, Piwnica-Worms D, Piwnica-Worms H (2014) Contribution of p53 to metastasis[J]. Cancer Discov 4(4):405–414
Babaei G, Aliarab A, AsghariVostakolaei M, Hotelchi M, Neisari R, Aziz SGG, Bazl MR (2021) Crosslink between p53 and metastasis: focus on epithelial-mesenchymal transition, cancer stem cell, angiogenesis, autophagy, and anoikis[J]. Mol Biol Rep 48(11):7545–7557
Joshi Y, Soria MG, Quadrato G et al (2015) The MDM4/MDM2-p53-IGF1 axis controls axonal regeneration, sprouting and functional recovery after CNS injury[J]. Brain 138(Pt 7):1843–1862
Acknowledgements
We thank everyone who provided support for this study. Thank the SangerBox website (http://sangerbox.com/Tool) for the helpful information on this manuscript.
Funding
This study was supported by the National Natural Science Foundation of China [Grant No. 81472355 (XJ)] and The Strategic Priority Research Program of the Chinese Academy of Sciences (XDA16040601). We are sincerely grateful to those who created and maintained these public databases.
Author information
Authors and Affiliations
Contributions
XJJ and CPR were responsible for experimental design. WY, ZPJ, and YWG were responsible for experimental analysis and thesis writing. HCZ, YDC, ZPW, YZ, QC, WDL, and MZ were responsible for data screening, and collection. All authors contributed to the article and approved the submitted version.
Corresponding authors
Ethics declarations
Ethics Approval
All experimental protocols were approved by the National Institute of Health’s guidelines and following the policies established by the Animal Care and Use Committee of Xiangya Hospital.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Competing Interests
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Yin, W., Jiang, Z., Guo, Y. et al. Identification of Anoikis-Related Genes in Spinal Cord Injury: Bioinformatics and Experimental Validation. Mol Neurobiol (2024). https://doi.org/10.1007/s12035-024-04121-8
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12035-024-04121-8