Abstract
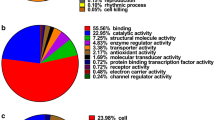
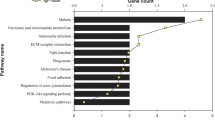
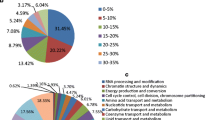
Identifying proteins associated with the onset of early intestinal-type gastric cancer (EIGC) can yield valuable insights into the pathogenesis of this specific subtype of gastric cancer. Data-independent acquisition mass spectroscopy (DIA-MS) was utilized to identify the differential protein between 10 cases of EIGC and atrophic gastritis with intestinal metaplasia (NGC). The expressions of IPO4, TBL1XR1, p62/SQSTM1, PKP3, and CRTAP were verified by immunohistochemistry (IHC) in 20 EIGC samples, 17 gastric low-grade intraepithelial neoplasia (LGIN) samples, and 21 healthy controls. The prognostic values of the five genes were validated in the transcriptome data by survival analysis. A total of 4,028 proteins were identified using DIA-MS and a total of 177 differential proteins were screened with log2(fold change) > 1.5. Among them, 113 proteins were significantly up-regulated, and 64 proteins were significantly down-regulated in EIGC tissues. IHC results showed that proteins IPO4, TBL1XR1, p62/SQSTM1, PKP3, and CRTAP were highly expressed in the cytoplasm of EIGC and LGIN, which was consistent with the results of DIA-MS. Among them, p62/SQSTM1 may undergo nuclear-cytoplasmic transfer. The five protein-coding genes were associated with intestinal-type gastric cancer survival and exhibited differential expression across various disease stages. The study successfully identified differentially expressed proteins between EIGC and NGC, providing potential biomarkers and valuable insights into the mechanism underlying intestinal-type gastric cancer.






Similar content being viewed by others
Data availability
The datasets analyzed in the study are available in public repositories at NCBI GEO (https://www.ncbi.nlm.nih.gov/geo/).
References
Ajani JA, Bentrem DJ, Besh S, D’Amico TA, Das P, Denlinger C, et al. Gastric cancer, version 2.2013: featured updates to the NCCN Guidelines. J Natl Compr Canc Netw. 2013;11:531–46. https://doi.org/10.6004/jnccn.2013.0070.
Aquino PF, Fischer JS, Neves-Ferreira AG, Perales J, Domont GB, Araujo GD, et al. Are gastric cancer resection margin proteomic profiles more similar to those from controls or tumors? J Proteome Res. 2012;11:5836–42. https://doi.org/10.1021/pr300612x.
Ashburner M, Ball CA, Blake JA, Botstein D, Butler H, Cherry JM, et al. Gene ontology: tool for the unification of biology The Gene Ontology Consortium. Nat Gene. 2000;25:25–9. https://doi.org/10.1038/75556.
Balluff B, Rauser S, Meding S, Elsner M, Schone C, Feuchtinger A, et al. MALDI imaging identifies prognostic seven-protein signature of novel tissue markers in intestinal-type gastric cancer. Am J Pathol. 2011;179:2720–9. https://doi.org/10.1016/j.ajpath.2011.08.032.
Chandrashekar DS, Karthikeyan SK, Korla PK, Patel H, Shovon AR, Athar M, et al. UALCAN: An update to the integrated cancer data analysis platform. Neoplasia. 2022;25:18–27. https://doi.org/10.1016/j.neo.2022.01.001.
Correa P. Gastric cancer: overview. Gastroenterol Clin North Am. 2013;42:211–7. https://doi.org/10.1016/j.gtc.2013.01.002.
Demirag GG, Sullu Y, Gurgenyatagi D, Okumus NO, Yucel I. Expression of plakophilins (PKP1, PKP2, and PKP3) in gastric cancers. Diagn Pathol. 2011;6:1. https://doi.org/10.1186/1746-1596-6-1.
Dixon MF, Genta RM, Yardley JH, Correa P. Classification and grading of gastritis. The updated Sydney System. International Workshop on the Histopathology of Gastritis, Houston 1994. Am J Surg Pathol. 1996;20:1161–81. https://doi.org/10.1097/00000478-199610000-00001.
Domon B, Aebersold R. Mass spectrometry and protein analysis. Science. 2006;312:212–7. https://doi.org/10.1126/science.1124619.
Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015;136:E359-386. https://doi.org/10.1002/ijc.29210.
Fujiyoshi MRA, Inoue H, Fujiyoshi Y, Nishikawa Y, Toshimori A, Shimamura Y, et al. Endoscopic classifications of early gastric cancer: a literature review. Cancers. 2021. https://doi.org/10.3390/cancers14010100.
Gravina GL, Senapedis W, McCauley D, Baloglu E, Shacham S, Festuccia C. Nucleo-cytoplasmic transport as a therapeutic target of cancer. J Hematol Oncol. 2014;7:85. https://doi.org/10.1186/s13045-014-0085-1.
Ivanova OM, Anufrieva KS, Kazakova AN, Malyants IK, Shnaider PV, Lukina MM, et al. Non-canonical functions of spliceosome components in cancer progression. Cell Death Dis. 2023;14:77. https://doi.org/10.1038/s41419-022-05470-9.
Japanese Gastric Cancer A. (2011) Japanese classification of gastric carcinoma: 3rd English edition. Gastric cancer : official journal of the International Gastric Cancer Association and the Japanese Gastric Cancer Association 14:101–112. doi: https://doi.org/10.1007/s10120-011-0041-5
Kaise M, Kato M, Urashima M, Arai Y, Kaneyama H, Kanzazawa Y, et al. Magnifying endoscopy combined with narrow-band imaging for differential diagnosis of superficial depressed gastric lesions. Endoscopy. 2009;41:310–5. https://doi.org/10.1055/s-0028-1119639.
Kim HK, Reyzer ML, Choi IJ, Kim CG, Kim HS, Oshima A, et al. Gastric cancer-specific protein profile identified using endoscopic biopsy samples via MALDI mass spectrometry. J Proteome Res. 2010;9:4123–30. https://doi.org/10.1021/pr100302b.
Kim JS, Bae GE, Kim KH, Lee SI, Chung C, Lee D, et al. Prognostic significance of LC3B and p62/SQSTM1 expression in gastric adenocarcinoma. Anticancer Res. 2019;39:6711–22. https://doi.org/10.21873/anticanres.13886.
Kitata RB, Yang JC, Chen YJ. Advances in data-independent acquisition mass spectrometry towards comprehensive digital proteome landscape. Mass Spectrometry Rev. 2022. https://doi.org/10.1002/mas.21781.
Komatsu M, Kageyama S, Ichimura Y. p62/SQSTM1/A170: physiology and pathology. Pharmacol Res. 2012;66:457–62. https://doi.org/10.1016/j.phrs.2012.07.004.
Kuramitsu Y, Baron B, Yoshino S, Zhang X, Tanaka T, Yashiro M, et al. Proteomic differential display analysis shows up-regulation of 14-3-3 sigma protein in human scirrhous-type gastric carcinoma cells. Anticancer Res. 2010;30:4459–65.
Lauren P. The two histological main types of gastric carcinoma: diffuse and so-called intestinal-type carcinoma. an attempt at a histo-clinical classification. Acta Pathol Microbiol Scand. 1965;64:31–49. https://doi.org/10.1111/apm.1965.64.1.31.
Liu W, Yang Q, Liu B, Zhu Z. Serum proteomics for gastric cancer. Clin Chim Acta. 2014;431:179–84. https://doi.org/10.1016/j.cca.2014.02.001.
Lordick F, Carneiro F, Cascinu S, Fleitas T, Haustermans K, Piessen G, et al. Gastric cancer: ESMO Clinical Practice Guideline for diagnosis, treatment and follow-up. Ann Oncol. 2022;33:1005–20. https://doi.org/10.1016/j.annonc.2022.07.004.
Lu J, Bang H, Kim SM, Cho SJ, Ashktorab H, Smoot DT, et al. Lymphatic metastasis-related TBL1XR1 enhances stemness and metastasis in gastric cancer stem-like cells by activating ERK1/2-SOX2 signaling. Oncogene. 2021;40:922–36. https://doi.org/10.1038/s41388-020-01571-x.
Ludwig C, Gillet L, Rosenberger G, Amon S, Collins BC, Aebersold R. Data-independent acquisition-based SWATH-MS for quantitative proteomics: a tutorial. Mol Syst Biol. 2018;14: e8126. https://doi.org/10.15252/msb.20178126.
Mao D, Xu R, Chen H, Chen X, Li D, Song S, et al. Cross-talk of focal adhesion-related gene defines prognosis and the immune microenvironment in gastric cancer. Front Cell Develop Biol. 2021;9: 716461. https://doi.org/10.3389/fcell.2021.716461.
Park S, Karalis JD, Hong C, Clemenceau JR, Porembka MR, Kim IH, et al. ACTA2 expression predicts survival and is associated with response to immune checkpoint inhibitors in gastric cancer. Clin Cancer Res. 2023;29:1077–85. https://doi.org/10.1158/1078-0432.CCR-22-1897.
Perdigao-Henriques R, Petrocca F, Altschuler G, Thomas MP, Le MT, Tan SM, et al. miR-200 promotes the mesenchymal to epithelial transition by suppressing multiple members of the Zeb2 and Snail1 transcriptional repressor complexes. Oncogene. 2016;35:158–72. https://doi.org/10.1038/onc.2015.69.
Raftopoulos SC, Kumarasinghe P, de Boer B, Iacobelli J, Kontorinis N, Fermoyle S, et al. Gastric intraepithelial neoplasia in a Western population. Eur J Gastroenterol Hepatol. 2012;24:48–54. https://doi.org/10.1097/MEG.0b013e32834dc1bb.
Raju D, Hussey S, Ang M, Terebiznik MR, Sibony M, Galindo-Mata E, et al. Vacuolating cytotoxin and variants in Atg16L1 that disrupt autophagy promote Helicobacter pylori infection in humans. Gastroenterology. 2012;142:1160–71. https://doi.org/10.1053/j.gastro.2012.01.043.
Schlemper RJ, Kato Y, Stolte M. Review of histological classifications of gastrointestinal epithelial neoplasia: differences in diagnosis of early carcinomas between Japanese and Western pathologists. J Gastroenterol. 2001;36:445–56. https://doi.org/10.1007/s005350170067.
Sekine S, Yoshida H, Jansen M, Kushima R. The Japanese viewpoint on the histopathology of early gastric cancer. Adv Exp Med Biol. 2016;908:331–46. https://doi.org/10.1007/978-3-319-41388-4_16.
Shen Q, Wang YE, Palazzo AF. Crosstalk between nucleocytoplasmic trafficking and the innate immune response to viral infection. J Biol Chem. 2021;297: 100856. https://doi.org/10.1016/j.jbc.2021.100856.
Szasz AM, Lanczky A, Nagy A, Forster S, Hark K, Green JE, et al. Cross-validation of survival associated biomarkers in gastric cancer using transcriptomic data of 1,065 patients. Oncotarget. 2016;7:49322–33. https://doi.org/10.18632/oncotarget.10337.
Tang B, Li N, Gu J, Zhuang Y, Li Q, Wang HG, et al. Compromised autophagy by MIR30B benefits the intracellular survival of Helicobacter pylori. Autophagy. 2012;8:1045–57. https://doi.org/10.4161/auto.20159.
Wagner AD, Syn NL, Moehler M, Grothe W, Yong WP, Tai BC, et al. Chemotherapy for advanced gastric cancer. Cochrane Database Syst Rev. 2017. https://doi.org/10.1002/14651858.CD004064.pub4.
Wang S, Han H, Hu Y, Yang W, Lv Y, Wang L, et al. MicroRNA-130a-3p suppresses cell migration and invasion by inhibition of TBL1XR1-mediated EMT in human gastric carcinoma. Mol Carcinog. 2018;57:383–92. https://doi.org/10.1002/mc.22762.
Xu X, Zhang X, Xing H, Liu Z, Jia J, Jin C, et al. Importin-4 functions as a driving force in human primary gastric cancer. J Cell Biochem. 2019;120:12638–46. https://doi.org/10.1002/jcb.28530.
Zhou X, Yao K, Zhang L, Zhang Y, Han Y, Liu HL, et al. Identification of differentiation-related proteins in gastric adenocarcinoma tissues by proteomics. Technol Cancer Res Treat. 2016;15:697–706. https://doi.org/10.1177/1533034615595792.
Acknowledgements
We are grateful for the technical support from Shanghai Applied Protein Technology Co. Ltd. This study was funded by the Zhejiang Provincial Natural Science Foundation of China (Nos. LGF19H030006 and LQ20H030001), Ningbo Science and Technology Project (No. 2019C50100), and Ningbo Clinical Medicine Research Center Project (No. 2019A21003).
Funding
This article was funded by Zhejiang Provincial Natural Science Foundation of China, LGF19H030006, Hua Ye, LQ20H030001, Hua Ye, Ningbo Science and Technology Project, 2019C50100, Hua Ye,Ningbo Clinical Medicine Research Center Project, 2019A21003, Hong Li.
Author information
Authors and Affiliations
Contributions
Hua Ye and Hong Li: conceived and designed the experiments. Liangshun Zhang and Feng Xu: collected samples, performed the proteome analysis, and analysed the data. Hongna Lu and Xianwen Dong: data analysis and plotting figures. Zhiqiang Gao and Qiaosu Zhao: performed the WGCNA and transcriptome data analysis. Ting Weng: survival analysis. Hong Li: drafting the manuscript. Hua Ye: project administration and revised the manuscript. All the authors participated in the manuscript revision.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no known competing financial interests or personal relationships that could have appeared to influence the work reported in this paper.
Ethical approval
The experimental protocol was approved by the Ethics Committee of Ningbo Medical Center Lihuili Hospital (KY2019PJ055). Informed consent was obtained from all participating patients or their family members.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhang, L., Xu, F., Lu, H. et al. Data-independent acquisition (DIA) mass spectrometry reveals related proteins involved in the occurrence of early intestinal-type gastric cancer. Med Oncol 41, 23 (2024). https://doi.org/10.1007/s12032-023-02241-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s12032-023-02241-0