Abstract
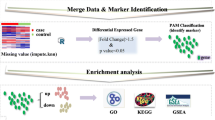
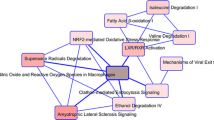
In pituitary adenoma (PA), invasiveness is the main cause of recurrence and poor prognosis. Thus, identifying specific biomarkers for diagnosis and effective treatment of invasive PAs is of great clinical significance. In this study, from the Gene Expression Omnibus database, we obtained and combined several microarrays of PA by the “sva” R package. Weighted gene co-expression network analysis was performed to construct a scale-free topology model and analyze the relationships between the modules and clinical traits. Our analysis results indicated that three key modules (dark turquoise, saddle brown, and steel blue) were associated with the invasiveness of PA. Kyoto Encyclopedia of Genes and Genomes pathway enrichment analysis and Gene Ontology analysis were performed for the functional annotation of the key modules. In addition, the hub genes in the three modules were identified and screened by differential expression analysis between normal samples and PA samples. Three upregulated differentially expressed genes (DGAT2, PIGZ, and DHRS2) were identified. The Fisher’s exact test and receiver operating characteristic curve were used to validate the capability of these genes to distinguish invasive traits, and transcription factor interaction networks were used to further explore the underlying mechanisms of the three genes. Moreover, a lower expression level of DGAT2 in invasive PA tissue than in noninvasive PA tissue was validated by quantitative reverse transcription-polymerase chain reaction. In general, this study contributes to potential molecular biomarkers of invasive PAs and provides a broader perspective for diagnosis and new therapeutic targets for the invasive PAs.








Similar content being viewed by others
References
J.S. Barnholtz-Sloan, Q.T. Ostrom, D. Cote, Epidemiology of brain tumors. Neurol. Clin. 36(3), 395–419 (2018). https://doi.org/10.1016/j.ncl.2018.04.001
E.D. Aflorei, M. Korbonits, Epidemiology and etiopathogenesis of pituitary adenomas. J. Neurooncol 117(3), 379–394 (2014). https://doi.org/10.1007/s11060-013-1354-5
B.W. Scheithauer, K.T. Kovacs, E.R. Laws Jr., R.V. Randall, Pathology of invasive pituitary tumors with special reference to functional classification. J. Neurosurg. 65(6), 733–744 (1986). https://doi.org/10.3171/jns.1986.65.6.0733
K. Thapar, K. Kovacs, B.W. Scheithauer, L. Stefaneanu, E. Horvath, P.J. Pernicone, D. Murray, E.R. Laws Jr., Proliferative activity and invasiveness among pituitary adenomas and carcinomas: an analysis using the MIB-1 antibody. Neurosurgery 38(1), 99–106 (1996). https://doi.org/10.1097/00006123-199601000-00024
B.P. Meij, M.B. Lopes, D.B. Ellegala, T.D. Alden, E.R. Laws Jr, The long-term significance of microscopic dural invasion in 354 patients with pituitary adenomas treated with transsphenoidal surgery. J. Neurosurg. 96(2), 195–208 (2002). https://doi.org/10.3171/jns.2002.96.2.0195
G.A. Kaltsas, P. Nomikos, G. Kontogeorgos, M. Buchfelder, A.B. Grossman, Clinical review: diagnosis and management of pituitary carcinomas. J. Clin. Endocrinol. Metab. 90(5), 3089–3099 (2005). https://doi.org/10.1210/jc.2004-2231
M. Buchfelder, Management of aggressive pituitary adenomas: current treatment strategies. Pituitary 12(3), 256–260 (2009). https://doi.org/10.1007/s11102-008-0153-z
A.I. McCormack, J.A. Wass, A.B. Grossman, Aggressive pituitary tumours: the role of temozolomide and the assessment of MGMT status. Eur. J. Clin. Invest 41(10), 1133–1148 (2011). https://doi.org/10.1111/j.1365-2362.2011.02520.x
G. Raverot, F. Castinetti, E. Jouanneau, I. Morange, D. Figarella-Branger, H. Dufour, J. Trouillas, T. Brue, Pituitary carcinomas and aggressive pituitary tumours: merits and pitfalls of temozolomide treatment. Clin. Endocrinol. 76(6), 769–775 (2012). https://doi.org/10.1111/j.1365-2265.2012.04381.x
M.B.S. Lopes, The 2017 World Health Organization classification of tumors of the pituitary gland: a summary. Acta Neuropathol. 134(4), 521–535 (2017). https://doi.org/10.1007/s00401-017-1769-8
C. Dai, X. Liu, W. Ma, R. Wang, The treatment of refractory pituitary adenomas. Front. Endocrinol. 10, 334 (2019). https://doi.org/10.3389/fendo.2019.00334
Q. Yang, X. Li, Molecular network basis of invasive pituitary adenoma: a review. Front. Endocrinol. 10, 7 (2019). https://doi.org/10.3389/fendo.2019.00007
S. Chiloiro, F. Doglietto, B. Trapasso, D. Iacovazzo, A. Giampietro, F. Di Nardo, C. de Waure, L. Lauriola, A. Mangiola, C. Anile, G. Maira, L. De Marinis, A. Bianchi, Typical and atypical pituitary adenomas: a single-center analysis of outcome and prognosis. Neuroendocrinology 101(2), 143–150 (2015). https://doi.org/10.1159/000375448
C.P. Miermeister, S. Petersenn, M. Buchfelder, R. Fahlbusch, D.K. Ludecke, A. Holsken, M. Bergmann, U.J. Knappe, V.H. Hans, J. Flitsch, W. Saeger, R. Buslei, Erratum: histological criteria for atypical pituitary adenomas–data from the German pituitary adenoma registry suggests modifications. Acta Neuropathol. Commun. 4, 21 (2016). https://doi.org/10.1186/s40478-016-0290-y
A. Di Ieva, F. Rotondo, L.V. Syro, M.D. Cusimano, K. Kovacs, Aggressive pituitary adenomas–diagnosis and emerging treatments. Nat. Rev. Endocrinol. 10(7), 423–435 (2014). https://doi.org/10.1038/nrendo.2014.64
Y. Yang, L. Han, Y. Yuan, J. Li, N. Hei, H. Liang, Gene co-expression network analysis reveals common system-level properties of prognostic genes across cancer types. Nat. Commun. 5, 3231 (2014). https://doi.org/10.1038/ncomms4231
G. Fiscon, F. Conte, V. Licursi, S. Nasi, P. Paci, Computational identification of specific genes for glioblastoma stem-like cells identity. Sci. Rep. 8(1), 7769 (2018). https://doi.org/10.1038/s41598-018-26081-5
R. Falcone, F. Conte, G. Fiscon, V. Pecce, M. Sponziello, C. Durante, L. Farina, S. Filetti, P. Paci, A. Verrienti, BRAF(V600E)-mutant cancers display a variety of networks by SWIM analysis: prediction of vemurafenib clinical response. Endocrine 64(2), 406–413 (2019). https://doi.org/10.1007/s12020-019-01890-4
S. van Dam, U. Vosa, A. van der Graaf, L. Franke, J.P. de Magalhaes, Gene co-expression analysis for functional classification and gene-disease predictions. Brief. Bioinform. 19(4), 575–592 (2018). https://doi.org/10.1093/bib/bbw139
P. Langfelder, S. Horvath, WGCNA: an R package for weighted correlation network analysis. BMC Bioinform. 9, 559 (2008). https://doi.org/10.1186/1471-2105-9-559
T. Zhai, D. Muhanhali, X. Jia, Z. Wu, Z. Cai, Y. Ling, Identification of gene co-expression modules and hub genes associated with lymph node metastasis of papillary thyroid cancer. Endocrine 66(3), 573–584 (2019). https://doi.org/10.1007/s12020-019-02021-9
N. Li, X. Zhan, Identification of clinical trait-related lncRNA and mRNA biomarkers with weighted gene co-expression network analysis as useful tool for personalized medicine in ovarian cancer. EPMA J. 10(3), 273–290 (2019). https://doi.org/10.1007/s13167-019-00175-0
P. Shannon, A. Markiel, O. Ozier, N.S. Baliga, J.T. Wang, D. Ramage, N. Amin, B. Schwikowski, T. Ideker, Cytoscape: a software environment for integrated models of biomolecular interaction networks. Genome Res. 13(11), 2498–2504 (2003). https://doi.org/10.1101/gr.1239303
E. Knosp, E. Steiner, K. Kitz, C. Matula, Pituitary adenomas with invasion of the cavernous sinus space: a magnetic resonance imaging classification compared with surgical findings. Neurosurgery 33(4), 610–617 (1993). https://doi.org/10.1227/00006123-199310000-00008
O. Mete, S. Ezzat, S.L. Asa, Biomarkers of aggressive pituitary adenomas. J. Mol. Endocrinol. 49(2), R69–R78 (2012). https://doi.org/10.1530/JME-12-0113
F. Conte, G. Fiscon, V. Licursi, D. Bizzarri, T. D’Anto, L. Farina, P. Paci, A paradigm shift in medicine: a comprehensive review of network-based approaches. Biochim. Biophys. Acta Gene Regul. Mech., 194416 (2019). https://doi.org/10.1016/j.bbagrm.2019.194416
A.L. Barabasi, N. Gulbahce, J. Loscalzo, Network medicine: a network-based approach to human disease. Nat. Rev. Genet. 12(1), 56–68 (2011). https://doi.org/10.1038/nrg2918
B. Aydin, K.Y. Arga, Co-expression network analysis elucidated a core module in association with prognosis of non-functioning non-invasive human pituitary adenoma. Front. Endocrinol. 10, 361 (2019). https://doi.org/10.3389/fendo.2019.00361
W. Xing, Z. Qi, C. Huang, N. Zhang, W. Zhang, Y. Li, M. Qiu, Q. Fang, G. Hui, Genome-wide identification of lncRNAs and mRNAs differentially expressed in non-functioning pituitary adenoma and construction of an lncRNA-mRNA co-expression network. Biol. Open 8(1), (2019). https://doi.org/10.1242/bio.037127
H. Joshi, B. Vastrad, C. Vastrad, Identification of important invasion-related genes in non-functional pituitary adenomas. J. Mol. Neurosci. 68(4), 565–589 (2019). https://doi.org/10.1007/s12031-019-01318-8
S.J. Thomas, J.A. Snowden, M.P. Zeidler, S.J. Danson, The role of JAK/STAT signalling in the pathogenesis, prognosis and treatment of solid tumours. Br. J. Cancer 113(3), 365–371 (2015). https://doi.org/10.1038/bjc.2015.233
J.J. O’Shea, D.M. Schwartz, A.V. Villarino, M. Gadina, I.B. McInnes, A. Laurence, The JAK-STAT pathway: impact on human disease and therapeutic intervention. Annu. Rev. Med. 66, 311–328 (2015). https://doi.org/10.1146/annurev-med-051113-024537
Y. Asari, K. Kageyama, Y. Nakada, M. Tasso, S. Takayasu, K. Niioka, N. Ishigame, M. Daimon, Inhibitory effects of a selective Jak2 inhibitor on adrenocorticotropic hormone production and proliferation of corticotroph tumor AtT20 cells. Onco Targets Ther. 10, 4329–4338 (2017). https://doi.org/10.2147/OTT.S141345
R. van der Pas, J.H. van Esch, C. de Bruin, A.H. Danser, A.M. Pereira, P.M. Zelissen, R. Netea-Maier, D.M. Sprij-Mooij, I.M. van den Berg-Garrelds, R.H. van Schaik, S.W. Lamberts, A.H. van den Meiracker, L.J. Hofland, R.A. Feelders, Cushing’s disease and hypertension: in vivo and in vitro study of the role of the renin-angiotensin-aldosterone system and effects of medical therapy. Eur. J. Endocrinol. 170(2), 181–191 (2014). https://doi.org/10.1530/EJE-13-0477
L. Faggi, A. Giustina, G. Tulipano, Effects of metformin on cell growth and AMPK activity in pituitary adenoma cell cultures, focusing on the interaction with adenylyl cyclase activating signals. Mol. Cell Endocrinol. 470, 60–74 (2018). https://doi.org/10.1016/j.mce.2017.09.030
A.B. Grossman, The molecular biology of pituitary tumors: a personal perspective. Pituitary 12(3), 265–270 (2009). https://doi.org/10.1007/s11102-008-0158-7
Y. Jin Kim, C. Hyun Kim, J. Hwan Cheong, J. Min Kim, Relationship between expression of vascular endothelial growth factor and intratumoral hemorrhage in human pituitary adenomas. Tumori 97(5), 639–646 (2011). https://doi.org/10.1700/989.10725
R. Sanchez-Ortiga, L. Sanchez-Tejada, O. Moreno-Perez, P. Riesgo, M. Niveiro, A.M. Pico Alfonso, Over-expression of vascular endothelial growth factor in pituitary adenomas is associated with extrasellar growth and recurrence. Pituitary 16(3), 370–377 (2013). https://doi.org/10.1007/s11102-012-0434-4
C. Zhao, M. Zhang, W. Liu, C. Wang, Q. Zhang, W. Li, Beta-catenin knockdown inhibits pituitary adenoma cell proliferation and invasion via interfering with AKT and gelatinases expression. Int. J. Oncol. 46(4), 1643–1650 (2015). https://doi.org/10.3892/ijo.2015.2862
X. Zhan, D.M. Desiderio, Signaling pathway networks mined from human pituitary adenoma proteomics data. BMC Med. Genomics 3, 13 (2010). https://doi.org/10.1186/1755-8794-3-13
C. Onofri, M. Theodoropoulou, M. Losa, E. Uhl, M. Lange, E. Arzt, G.K. Stalla, U. Renner, Localization of vascular endothelial growth factor (VEGF) receptors in normal and adenomatous pituitaries: detection of a non-endothelial function of VEGF in pituitary tumours. J. Endocrinol. 191(1), 249–261 (2006). https://doi.org/10.1677/joe.1.06992
Y. Li, T. Li, Y. Jin, J. Shen, Dgat2 reduces hepatocellular carcinoma malignancy via downregulation of cell cycle-related gene expression. Biomed. Pharmacother. 115, 108950 (2019). https://doi.org/10.1016/j.biopha.2019.108950
R. Nurminen, T. Rantapero, S.C. Wong, D. Fischer, R. Lehtonen, T.L. Tammela, M. Nykter, T. Visakorpi, T. Wahlfors, J. Schleutker, Expressional profiling of prostate cancer risk SNPs at 11q13.5 identifies DGAT2 as a new target gene. Genes Chromosomes Cancer 55(8), 661–673 (2016). https://doi.org/10.1002/gcc.22368
Y. Han, Z. Wang, S. Sun, Z. Zhang, J. Liu, X. Jin, P. Wu, T. Ji, W. Ding, B. Wang, Q. Gao, Decreased DHRS2 expression is associated with HDACi resistance and poor prognosis in ovarian cancer. Epigenetics 15(1–2), 122–133 (2020). https://doi.org/10.1080/15592294.2019.1656155
Y. Zhou, L. Wang, X. Ban, T. Zeng, Y. Zhu, M. Li, X.Y. Guan, Y. Li, DHRS2 inhibits cell growth and motility in esophageal squamous cell carcinoma. Oncogene 37(8), 1086–1094 (2018). https://doi.org/10.1038/onc.2017.383
B.W. Taron, P.A. Colussi, J.M. Wiedman, P. Orlean, C.H. Taron, Human Smp3p adds a fourth mannose to yeast and human glycosylphosphatidylinositol precursors in vivo. J. Biol. Chem. 279(34), 36083–36092 (2004). https://doi.org/10.1074/jbc.M405081200
S.L. Asa, Practical pituitary pathology: what does the pathologist need to know? Arch. Pathol. Lab. Med. 132(8), 1231–1240 (2008). https://doi.org/10.1043/1543-2165(2008)132[1231:PPPWDT]2.0.CO;2
O. Mete, S.L. Asa, Clinicopathological correlations in pituitary adenomas. Brain Pathol. 22(4), 443–453 (2012). https://doi.org/10.1111/j.1750-3639.2012.00599.x
H. Nishioka, N. Inoshita, O. Mete, S.L. Asa, K. Hayashi, A. Takeshita, N. Fukuhara, M. Yamaguchi-Okada, Y. Takeuchi, S. Yamada, The complementary role of transcription factors in the accurate diagnosis of clinically nonfunctioning pituitary adenomas. Endocr. Pathol. 26(4), 349–355 (2015). https://doi.org/10.1007/s12022-015-9398-z
R.S. Viger, S.M. Guittot, M. Anttonen, D.B. Wilson, M. Heikinheimo, Role of the GATA family of transcription factors in endocrine development, function, and disease. Mol. Endocrinol. 22(4), 781–798 (2008). https://doi.org/10.1210/me.2007-0513
M. Pihlajoki, A. Farkkila, T. Soini, M. Heikinheimo, D.B. Wilson, GATA factors in endocrine neoplasia. Mol. Cell Endocrinol. 421, 2–17 (2016). https://doi.org/10.1016/j.mce.2015.05.027
J. He, J.J. Yu, Q. Xu, L. Wang, J.Z. Zheng, L.Z. Liu, B.H. Jiang, Downregulation of ATG14 by EGR1-MIR152 sensitizes ovarian cancer cells to cisplatin-induced apoptosis by inhibiting cyto-protective autophagy. Autophagy 11(2), 373–384 (2015). https://doi.org/10.1080/15548627.2015.1009781
H.T. Liu, S. Liu, L. Liu, R.R. Ma, P. Gao, EGR1-mediated transcription of lncRNA-HNF1A-AS1 promotes cell-cycle progression in gastric cancer. Cancer Res. 78(20), 5877–5890 (2018). https://doi.org/10.1158/0008-5472.CAN-18-1011
L. Li, A.H. Ameri, S. Wang, K.H. Jansson, O.M. Casey, Q. Yang, M.L. Beshiri, L. Fang, R.G. Lake, S. Agarwal, A.N. Alilin, W. Xu, J. Yin, K. Kelly, EGR1 regulates angiogenic and osteoclastogenic factors in prostate cancer and promotes metastasis. Oncogene 38(35), 6241–6255 (2019). https://doi.org/10.1038/s41388-019-0873-8
S.W. Sun, X.M. Fang, Y.F. Li, Q.B. Wang, Y.X. Li, Expression and clinical significance of EGR-1 and PTEN in the pituitary tumors of elderly patients. Oncol. Lett. 14(2), 2165–2169 (2017). https://doi.org/10.3892/ol.2017.6375
L. Xu, Y. Chen, M. Dutra-Clarke, A. Mayakonda, M. Hazawa, S.E. Savinoff, N. Doan, J.W. Said, W.H. Yong, A. Watkins, H. Yang, L.W. Ding, Y.Y. Jiang, J.W. Tyner, J. Ching, J.P. Kovalik, V. Madan, S.L. Chan, M. Muschen, J.J. Breunig, D.C. Lin, H.P. Koeffler, BCL6 promotes glioma and serves as a therapeutic target. Proc. Natl Acad. Sci. USA 114(15), 3981–3986 (2017). https://doi.org/10.1073/pnas.1609758114
Acknowledgements
The authors thank all the researchers and staff who supported The Gene Expression Omnibus database.
Funding
This work was funded by the General Program of National Natural Science Foundation of Jiangxi Province (grant number: 20192BAB205042) and the Health and Family Planning Commission of Jiangxi Province (grant number: 20195109).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of The First Affiliated Hospital of Nanchang University and with the 1964 Helsinki declaration and its later amendments or comparable ethical standards.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Zhou, Y., Fu, X., Zheng, Z. et al. Identification of gene co-expression modules and hub genes associated with the invasiveness of pituitary adenoma. Endocrine 68, 377–389 (2020). https://doi.org/10.1007/s12020-020-02316-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12020-020-02316-2