Abstract
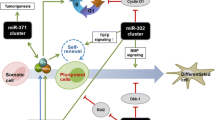
Cells of the inner cell mass (ICM) acquire a unique ability for unlimited self-renewal during transition into embryonic stem cells (ESCs) in vitro, while preserving their natural multi-lineage differentiation potential. Several different pathways have been identified to play roles in ESC formation but the function of non-coding RNAs in this process is poorly understood. Here, we describe several microRNAs (miRNAs) that are crucial for efficient generation of mouse ESCs from ICMs. Using small-RNA sequencing, we characterize dynamic changes in miRNA expression profiles during outgrowth of ICMs in a high-resolution, time-course dependent manner. We report several waves of miRNA transcription during ESC formation, to which miRNAs from the imprinted Dlk1-Dio3 locus contribute extensively. In silico analyses followed by functional investigations reveal that Dlk1-Dio3 locus-embedded miRNAs (miR-541-5p, miR-410-3p, and miR-381-3p), miR-183-5p, and miR-302b-3p promote, while miR-212-5p and let-7d-3p inhibit ESC formation. Collectively, these findings offer new mechanistic insights into the role of miRNAs during ESC derivation.
Graphical Abstract








Similar content being viewed by others
Data Availability
Small RNA sequencing data have been deposited in the Gene Expression Omnibus [46] database under GSE205361 [https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE205361]. All the other data supporting the findings of the present study are available in the article and its supplementary files, and from the lead contact (Hossein Baharvand: h.baharvand@royan-rc.ac.ir) upon request.
References
Choi, J., Huebner, A. J., Clement, K., Walsh, R. M., Savol, A., Lin, K., Gu, H., Di Stefano, B., Brumbaugh, J., Kim, S. Y., Sharif, J., Rose, C. M., Mohammad, A., Odajima, J., Charron, J., Shioda, T., Gnirke, A., Gygi, S., Koseki, H., Sadreyev, R. I., Xiao, A., Meissner, A., & Hochedlinger, K. (2017). Prolonged Mek1/2 suppression impairs the developmental potential of embryonic stem cells. Nature, 548(7666), 219–223. https://doi.org/10.1038/nature23274
Czechanski, A., Byers, C., Greenstein, I., Schrode, N., Donahue, L. R., Hadjantonakis, A. K., & Reinholdt, L. G. (2014). Derivation and characterization of mouse embryonic stem cells from permissive and nonpermissive strains. Nature protocols, 9(3), 559–574. https://doi.org/10.1038/nprot.2014.030
van Oosten, A. L., Costa, Y., Smith, A., & Silva, J. C. (2012). JAK/STAT3 signalling is sufficient and dominant over antagonistic cues for the establishment of naive pluripotency. Nature communications, 3, 817. https://doi.org/10.1038/ncomms1822
Ying, Q. L., Wray, J., Nichols, J., Batlle-Morera, L., Doble, B., Woodgett, J., Cohen, P., & Smith, A. (2008). The ground state of embryonic stem cell self-renewal. Nature, 453(7194), 519–523. https://doi.org/10.1038/nature06968
Hassani, S. N., Moradi, S., Taleahmad, S., Braun, T., & Baharvand, H. (2019). Transition of inner cell mass to embryonic stem cells: Mechanisms, facts, and hypotheses. Cellular and molecular life sciences: CMLS, 76(5), 873–892. https://doi.org/10.1007/s00018-018-2965-y
Hassani, S. N., Totonchi, M., Sharifi-Zarchi, A., Mollamohammadi, S., Pakzad, M., Moradi, S., Samadian, A., Masoudi, N., Mirshahvaladi, S., Farrokhi, A., Greber, B., Arauzo-Bravo, M. J., Sabour, D., Sadeghi, M., Salekdeh, G. H., Gourabi, H., Scholer, H. R., & Baharvand, H. (2014). Inhibition of TGFbeta signaling promotes ground state pluripotency. Stem cell reviews, 10(1), 16–30. https://doi.org/10.1007/s12015-013-9473-0
Marks, H., Kalkan, T., Menafra, R., Denissov, S., Jones, K., Hofemeister, H., Nichols, J., Kranz, A., Stewart, A. F., Smith, A., & Stunnenberg, H. G. (2012). The transcriptional and epigenomic foundations of ground state pluripotency. Cell, 149(3), 590–604. https://doi.org/10.1016/j.cell.2012.03.026
Hackett, J. A., & Surani, M. A. (2014). Regulatory principles of pluripotency: From the ground state up. Cell Stem Cell, 15(4), 416–430. https://doi.org/10.1016/j.stem.2014.09.015
Moradi, S., Sharifi-Zarchi, A., Ahmadi, A., Mollamohammadi, S., Stubenvoll, A., Gunther, S., Salekdeh, G. H., Asgari, S., Braun, T., & Baharvand, H. (2017). Small RNA sequencing reveals Dlk1-Dio3 locus-embedded MicroRNAs as Major Drivers of Ground-State Pluripotency. Stem cell reports, 9(6), 2081–2096. https://doi.org/10.1016/j.stemcr.2017.10.009
Yan, Y., Yang, X., Li, T. T., Gu, K. L., Hao, J., Zhang, Q., & Wang, Y. (2017). Significant differences of function and expression of microRNAs between ground state and serum-cultured pluripotent stem cells. Journal of Genetics and Genomics, 44(4), 179–189. https://doi.org/10.1016/j.jgg.2017.01.005
Tang, F., Barbacioru, C., Bao, S., Lee, C., Nordman, E., Wang, X., Lao, K., & Surani, M. A. (2010). Tracing the derivation of embryonic stem cells from the inner cell mass by single-cell RNA-Seq analysis. Cell stem cell, 6(5), 468–478. https://doi.org/10.1016/j.stem.2010.03.015
Totonchi, M., Hassani, S. N., Sharifi-Zarchi, A., Tapia, N., Adachi, K., Arand, J., Greber, B., Sabour, D., Arauzo-Bravo, M. J., Walter, J., Pakzad, M., Gourabi, H., Scholer, H. R., & Baharvand, H. (2017). Blockage of the epithelial-to-mesenchymal transition is required for embryonic stem cell derivation. Stem cell reports, 9(4), 1275–1290. https://doi.org/10.1016/j.stemcr.2017.08.006
Li, N., Long, B., Han, W., Yuan, S., & Wang, K. (2017). microRNAs: Important regulators of stem cells. Stem cell research & therapy, 8(1), 110. https://doi.org/10.1186/s13287-017-0551-0
Gross, N., Kropp, J., & Khatib, H. (2017). MicroRNA signaling in embryo development. Biology, 6(3), https://doi.org/10.3390/biology6030034
Moradi, S., Asgari, S., & Baharvand, H. (2014). Concise review: Harmonies played by microRNAs in cell fate reprogramming. Stem cells, 32(1), 3–15. https://doi.org/10.1002/stem.1576
Bernstein, E., Kim, S. Y., Carmell, M. A., Murchison, E. P., Alcorn, H., Li, M. Z., Mills, A. A., Elledge, S. J., Anderson, K. V., & Hannon, G. J. (2003). Dicer is essential for mouse development. Nature genetics, 35(3), 215–217. https://doi.org/10.1038/ng1253
Solter, D., & Knowles, B. B. (1975). Immunosurgery of mouse blastocyst. Proceedings of the National Academy of Sciences of the United States of America, 72(12), 5099–5102. https://doi.org/10.1073/pnas.72.12.5099
Mitsui, K., Tokuzawa, Y., Itoh, H., Segawa, K., Murakami, M., Takahashi, K., Maruyama, M., Maeda, M., & Yamanaka, S. (2003). The homeoprotein nanog is required for maintenance of pluripotency in mouse epiblast and ES cells. Cell, 113(5), 631–642.
Ohnishi, Y., Totoki, Y., Toyoda, A., Watanabe, T., Yamamoto, Y., Tokunaga, K., Sakaki, Y., Sasaki, H., & Hohjoh, H. (2010). Small RNA class transition from siRNA/piRNA to miRNA during pre-implantation mouse development. Nucleic acids research, 38(15), 5141–5151. https://doi.org/10.1093/nar/gkq229
Kuenne, C., Preussner, J., Herzog, M., Braun, T., & Looso, M. (2014). MIRPIPE: Quantification of microRNAs in niche model organisms. Bioinformatics, 30(23), 3412–3413. https://doi.org/10.1093/bioinformatics/btu573
Fagnocchi, L., Cherubini, A., Hatsuda, H., Fasciani, A., Mazzoleni, S., Poli, V., Berno, V., Rossi, R. L., Reinbold, R., Endele, M., Schroeder, T., Rocchigiani, M., Szkarlat, Z., Oliviero, S., Dalton, S., & Zippo, A. (2016). A myc-driven self-reinforcing regulatory network maintains mouse embryonic stem cell identity. Nature communications, 7, 11903. https://doi.org/10.1038/ncomms11903
Huang, G., Ye, S., Zhou, X., Liu, D., & Ying, Q. L. (2015). Molecular basis of embryonic stem cell self-renewal: From signaling pathways to pluripotency network. Cellular and molecular life sciences: CMLS, 72(9), 1741–1757. https://doi.org/10.1007/s00018-015-1833-2
Chen, H., Guo, R., Zhang, Q., Guo, H., Yang, M., Wu, Z., Gao, S., Liu, L., & Chen, L. (2015). Erk signaling is indispensable for genomic stability and self-renewal of mouse embryonic stem cells. Proceedings of the National Academy of Sciences of the United States of America, 112(44), E5936–5943. https://doi.org/10.1073/pnas.1516319112
Judson, R. L., Babiarz, J. E., Venere, M., & Blelloch, R. (2009). Embryonic stem cell-specific microRNAs promote induced pluripotency. Nature biotechnology, 27(5), 459–461. https://doi.org/10.1038/nbt.1535
Wang, Y., Baskerville, S., Shenoy, A., Babiarz, J. E., Baehner, L., & Blelloch, R. (2008). Embryonic stem cell-specific microRNAs regulate the G1-S transition and promote rapid proliferation. Nature genetics, 40(12), 1478–1483. https://doi.org/10.1038/ng.250
Houbaviy, H. B., Murray, M. F., & Sharp, P. A. (2003). Embryonic stem cell-specific MicroRNAs. Developmental cell, 5(2), 351–358. https://doi.org/10.1016/s1534-5807(03)00227-2
He, L., He, X., Lim, L. P., de Stanchina, E., Xuan, Z., Liang, Y., Xue, W., Zender, L., Magnus, J., Ridzon, D., Jackson, A. L., Linsley, P. S., Chen, C., Lowe, S. W., Cleary, M. A., & Hannon, G. J. (2007). A microRNA component of the p53 tumour suppressor network. Nature, 447(7148), 1130–1134. https://doi.org/10.1038/nature05939
Choi, Y. J., Lin, C. P., Ho, J. J., He, X., Okada, N., Bu, P., Zhong, Y., Kim, S. Y., Bennett, M. J., Chen, C., Ozturk, A., Hicks, G. G., Hannon, G. J., & He, L. (2011). miR-34 miRNAs provide a barrier for somatic cell reprogramming. Nature cell biology, 13(11), 1353–1360. https://doi.org/10.1038/ncb2366
Liu, L., Luo, G. Z., Yang, W., Zhao, X., Zheng, Q., Lv, Z., Li, W., Wu, H. J., Wang, L., Wang, X. J., & Zhou, Q. (2010). Activation of the imprinted Dlk1-Dio3 region correlates with pluripotency levels of mouse stem cells. The Journal of biological chemistry, 285(25), 19483–19490. https://doi.org/10.1074/jbc.M110.131995
Stadtfeld, M., Apostolou, E., Akutsu, H., Fukuda, A., Follett, P., Natesan, S., Kono, T., Shioda, T., & Hochedlinger, K. (2010). Aberrant silencing of imprinted genes on chromosome 12qF1 in mouse induced pluripotent stem cells. Nature, 465(7295), 175–181. https://doi.org/10.1038/nature09017
Benetatos, L., Vartholomatos, G., & Hatzimichael, E. (2014). DLK1-DIO3 imprinted cluster in induced pluripotency: Landscape in the mist. Cellular and molecular life sciences: CMLS, 71(22), 4421–4430. https://doi.org/10.1007/s00018-014-1698-9
Abdelalim, E. M., & Tooyama, I. (2012). The p53 inhibitor, pifithrin-alpha, suppresses self-renewal of embryonic stem cells. Biochemical and biophysical research communications, 420(3), 605–610. https://doi.org/10.1016/j.bbrc.2012.03.041
Brosh, R., Assia-Alroy, Y., Molchadsky, A., Bornstein, C., Dekel, E., Madar, S., Shetzer, Y., Rivlin, N., Goldfinger, N., Sarig, R., & Rotter, V. (2013). p53 counteracts reprogramming by inhibiting mesenchymal-to-epithelial transition. Cell death and differentiation, 20(2), 312–320. https://doi.org/10.1038/cdd.2012.125
Jain, A. K., Allton, K., Iacovino, M., Mahen, E., Milczarek, R. J., Zwaka, T. P., Kyba, M., & Barton, M. C. (2012). p53 regulates cell cycle and microRNAs to promote differentiation of human embryonic stem cells. PLoS biology, 10(2), e1001268. https://doi.org/10.1371/journal.pbio.1001268
Hassani, S. N., Totonchi, M., Farrokhi, A., Taei, A., Larijani, M. R., Gourabi, H., & Baharvand, H. (2012). Simultaneous suppression of TGF-beta and ERK signaling contributes to the highly efficient and reproducible generation of mouse embryonic stem cells from previously considered refractory and non-permissive strains. Stem cell reviews, 8(2), 472–481. https://doi.org/10.1007/s12015-011-9306-y
Hassani, S. N., Totonchi, M., Sharifi-Zarchi, A., Mollamohammadi, S., Pakzad, M., Moradi, S., Samadian, A., Masoudi, N., Mirshahvaladi, S., Farrokhi, A., Greber, B., Arauzo-Bravo, M. J., Sabour, D., Sadeghi, M., Salekdeh, G. H., Gourabi, H., Scholer, H. R., & Baharvand, H. (2013). Inhibition of TGFbeta Signaling promotes Ground State Pluripotency. Stem cell reviews. https://doi.org/10.1007/s12015-013-9473-0
Frum, T., Murphy, T. M., & Ralston, A. (2018). HIPPO signaling resolves embryonic cell fate conflicts during establishment of pluripotency in vivo. Elife, 7, https://doi.org/10.7554/eLife.42298
Sun, X., Ren, Z., Cun, Y., Zhao, C., Huang, X., Zhou, J., Hu, R., Su, X., Ji, L., Li, P., Mak, K. L. K., Gao, F., Yang, Y., Xu, H., Ding, J., Cao, N., Li, S., Zhang, W., Lan, P., Sun, H., Wang, J., & Yuan, P. (2020). Hippo-YAP signaling controls lineage differentiation of mouse embryonic stem cells through modulating the formation of super-enhancers. Nucleic acids research, 48(13), 7182–7196. https://doi.org/10.1093/nar/gkaa482
Pfaff, N., Liebhaber, S., Mobus, S., Beh-Pajooh, A., Fiedler, J., Pfanne, A., Schambach, A., Thum, T., Cantz, T., & Moritz, T. (2017). Inhibition of miRNA-212/132 improves the reprogramming of fibroblasts into induced pluripotent stem cells by de-repressing important epigenetic remodelling factors. Stem cell research, 20, 70–75. https://doi.org/10.1016/j.scr.2017.03.003
Yu, B., Zhou, S., Wang, Y., Qian, T., Ding, G., Ding, F., & Gu, X. (2012). miR-221 and miR-222 promote Schwann cell proliferation and migration by targeting LASS2 after sciatic nerve injury. Journal of cell science, 125(Pt 11), 2675–2683. https://doi.org/10.1242/jcs.098996
Schee, K., Lorenz, S., Worren, M. M., Gunther, C. C., Holden, M., Hovig, E., Fodstad, O., Meza-Zepeda, L. A., & Flatmark, K. (2013). Deep sequencing the MicroRNA Transcriptome in Colorectal Cancer. PloS one, 8(6), e66165. https://doi.org/10.1371/journal.pone.0066165
Clancy, J. L., Patel, H. R., Hussein, S. M., Tonge, P. D., Cloonan, N., Corso, A. J., Li, M., Lee, D. S., Shin, J. Y., Wong, J. J., Bailey, C. G., Benevento, M., Munoz, J., Chuah, A., Wood, D., Rasko, J. E., Heck, A. J., Grimmond, S. M., Rogers, I. M., Seo, J. S., Wells, C. A., Puri, M. C., Nagy, A., & Preiss, T. (2014). Small RNA changes en route to distinct cellular states of induced pluripotency. Nature communications, 5, 5522. https://doi.org/10.1038/ncomms6522
Wang, Y., Melton, C., Li, Y. P., Shenoy, A., Zhang, X. X., Subramanyam, D., & Blelloch, R. (2013). miR-294/miR-302 promotes proliferation, suppresses G1-S restriction point, and inhibits ESC differentiation through separable mechanisms. Cell reports, 4(1), 99–109. https://doi.org/10.1016/j.celrep.2013.05.027
Silva, J., Nichols, J., Theunissen, T. W., Guo, G., van Oosten, A. L., Barrandon, O., Wray, J., Yamanaka, S., Chambers, I., & Smith, A. (2009). Nanog is the gateway to the pluripotent ground state. Cell, 138(4), 722–737. https://doi.org/10.1016/j.cell.2009.07.039
Polo, J. M., Anderssen, E., Walsh, R. M., Schwarz, B. A., Nefzger, C. M., Lim, S. M., Borkent, M., Apostolou, E., Alaei, S., Cloutier, J., Bar-Nur, O., Cheloufi, S., Stadtfeld, M., Figueroa, M. E., Robinton, D., Natesan, S., Melnick, A., Zhu, J., Ramaswamy, S., & Hochedlinger, K. (2012). A molecular roadmap of reprogramming somatic cells into iPS cells. Cell, 151(7), 1617–1632. https://doi.org/10.1016/j.cell.2012.11.039
Edgar, R., Domrachev, M., & Lash, A. E. (2002). Gene expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic acids research, 30(1), 207–210. https://doi.org/10.1093/nar/30.1.207
Acknowledgements
We thank Behrouz Asgari and Zahrasadat Ghasemi for assistance with harvesting blastocyst embryos. We are also grateful to other colleagues at Royan Institute as well as at Max-Planck Institute for Heart & Lung Research for discussions.
Funding
Research in H.B. lab is funded by Royan Institute, the Iranian Council of Stem Cell Research and Technology, the Iran National Science Foundation (INSF), and Iran Science Elites Federation. T.B. was supported by grants from Max-Planck Society. SM was supported by a grant from Royan Institute.
Author information
Authors and Affiliations
Contributions
H.B. and S.M. conceived and designed the study. S.M., H.B., and T.B. designed the experiments, and analyzed and interpreted the data. S.M. performed most of the experiments and wrote the manuscript. S.G. performed small-RNA sequencing. S.S. contributed to ESC derivation and maintenance, miRNA transfection, and data analysis. H.B., S.M., and T.B. provided financial and administrative support, discussed the results, and approved the manuscript. C.K., A.S.-Z., and SM performed bioinformatics analysis of the small-RNA sequencing data and regulatory network construction. V.K. contributed to defining criteria for selecting candidate miRNAs and data analysis. P.T. and S.K. handled mice and harvested blastocyst-stage embryos. All authors reviewed and confirmed the manuscript before submission.
Corresponding authors
Ethics declarations
Ethics Approval
Animal procedures were conducted in accordance with Animal Research Guidelines of the Iran’s Ministry of Health and Education and Ethics Committee of the Royan Institute.
Consent for Publication
All authors approved the manuscript for integrity and publication.
Conflicts of Interest
The authors declare no conflicts of interests.
Consent to Participate
Not applicable.
Additional information
Publisher’s Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Moradi, S., Guenther, S., Soori, S. et al. Time-resolved Small-RNA Sequencing Identifies MicroRNAs Critical for Formation of Embryonic Stem Cells from the Inner Cell Mass of Mouse Embryos. Stem Cell Rev and Rep 19, 2361–2377 (2023). https://doi.org/10.1007/s12015-023-10582-6
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12015-023-10582-6