Abstract
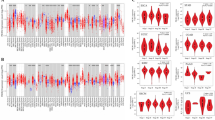
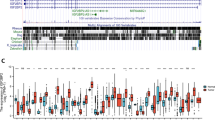
Prolyl 3-hydroxylase 1 (P3H1) has been implicated in cancer development, but no pan-cancer analysis has been conducted on P3H1. In this study, for the first time, aspects associated with P3H1, such as the mRNA expression, any mutation, promoter methylation, and prognostic significance, the relationship between P3H1 and clinicopathological parameters, drug sensitivity, and immune cell infiltration were investigated by searching several databases including The Cancer Genome Atlas (TCGA), Genotype-Tissue Expression (GTEx), cBioPortal, and The Tumor Immune Evaluation Resource (TIMER2.0) using bioinformatics tools. The findings indicate significant differential expression of P3H1 in most tumors when compared to normal tissues, with a strong association with clinical prognosis. A pan-cancer Cox regression analysis revealed that high P3H1 expression is significantly associated with low overall survival in patients with brain lower grade glioma, kidney clear cell carcinoma, adrenocortical cancer, liver hepatocellular carcinoma, mesothelioma, sarcoma, uveal melanoma, bladder urothelial carcinoma, kidney papillary cell carcinoma, kidney chromophobe, thymoma, and thyroid carcinoma. A negative correlation was observed between P3H1 DNA methylation and its expression. P3H1 is significantly associated with infiltrating cells, immune-related genes, tumor mutation burden, microsatellite instability, and mismatch repair. Finally, A significant correlation was found between P3H1 expression and sensitivity to nine drugs. Thus, enhanced P3H1 expression is associated with poor prognosis in a variety of tumors, which may be due to its role in tumor immune regulation and tumor microenvironment. This pan-cancer analysis provides insight into the function of P3H1 in tumorigenesis of different cancers and provides a theoretical basis for further in-depth studies to follow.












Similar content being viewed by others
Data Availability
The data sets used in this research are publicly available online.
References
Sung, H., Ferlay, J., Siegel, R. L., Laversanne, M., Soerjomataram, I., et al. (2021). Global cancer statistics 2020: Globocan estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA: A Cancer Journal for Clinicians, 71, 209–249. https://doi.org/10.3322/caac.21660
Yang, M., Oh, I. Y., Mahanty, A., Jin, W. L., & Yoo, J. S. (2020). Immunotherapy for glioblastoma: current state, challenges, and future perspectives, Cancers (Basel), 12. https://doi.org/10.3390/cancers12092334
Bisht, D., Arora, A., & Sachan, M. (2022). Role of dna de-methylation intermediate “5-hydroxymethylcytosine” in ovarian cancer management: A comprehensive review. Biomedicine & Pharmacotherapy, 155, 113674. https://doi.org/10.1016/j.biopha.2022.113674
Liu, X., Chen, L., & Wang, T. (2022). Overcoming cisplatin resistance of human lung cancer by sinomenine through targeting the mir-200a-3p-gls axis, J Chemother, 1–10. https://doi.org/10.1080/1120009X.2022.2111490.
Vranka, J. A., Pokidysheva, E., Hayashi, L., Zientek, K., Mizuno, K., et al. (2010). Prolyl 3-hydroxylase 1 null mice display abnormalities in fibrillar collagen-rich tissues such as tendons, skin, and bones. Journal of Biological Chemistry, 285, 17253–17262. https://doi.org/10.1074/jbc.M110.102228
Wu, J., Zhang, W., Xia, L., Feng, L., Shu, Z., et al. (2019). Characterization of ppib interaction in the p3h1 ternary complex and implications for its pathological mutations. Cellular and Molecular Life Sciences, 76, 3899–3914. https://doi.org/10.1007/s00018-019-03102-8
Hudson, D. M., Weis, M., Rai, J., Joeng, K. S., Dimori, M., et al. (2017). P3h3-null and sc65-null mice phenocopy the collagen lysine under-hydroxylation and cross-linking abnormality of ehlers-danlos syndrome type via. Journal of Biological Chemistry, 292, 3877–3887. https://doi.org/10.1074/jbc.M116.762245
Cabral, W. A., Fratzl-Zelman, N., Weis, M., Perosky, J. E., Alimasa, A., et al. (2020). Substitution of murine type i collagen a1 3-hydroxylation site alters matrix structure but does not recapitulate osteogenesis imperfecta bone dysplasia. Matrix Biology, 90, 20–39. https://doi.org/10.1016/j.matbio.2020.02.003
Tan, W., Ji, Y., Qian, Y., Lin, Y., Ye, R., et al. (2022). Mutational screening of skeletal genes in 14 chinese children with osteogenesis imperfecta using targeted sequencing. Journal of Immunology Research, 2022, 5068523. https://doi.org/10.1155/2022/5068523
Zhytnik, L., Duy, B. H., Eekhoff, M., Wisse, L., & Pals, G., et al. (2022) Phenotypic variation in vietnamese osteogenesis imperfecta patients sharing a recessive p3h1 pathogenic variant, Genes (Basel), 13. https://doi.org/10.3390/genes13030407
Nadyrshina, D., Zaripova, A., Tyurin, A., Minniakhmetov, I., Zakharova, E., & Khusainova, R. (2022). Osteogenesis imperfecta: search for mutations in patients from the republic of bashkortostan (russia), Genes (Basel), 13. https://doi.org/10.3390/genes13010124
Tuysuz, B., Elkanova, L., Uludag, A. D., Gulec, C., Toksoy, G., et al. (2022). Osteogenesis imperfecta in 140 turkish families: Molecular spectrum and comparison of long-term clinical outcome of those with col1a1/a2 and biallelic variants. Bone, 155, 116293. https://doi.org/10.1016/j.bone.2021.116293
Scollo, P., Snead, M. P., Richards, A. J., Pollitt, R., & Devile, C. (2018). Bilateral giant retinal tears in osteogenesis imperfecta. BMC Medical Genetics, 19, 8. https://doi.org/10.1186/s12881-018-0521-0
Pokidysheva, E., Tufa, S., Bresee, C., Brigande, J. V., & Bachinger, H. P. (2013). Prolyl 3-hydroxylase-1 null mice exhibit hearing impairment and abnormal morphology of the middle ear bone joints. Matrix Biology, 32, 39–44. https://doi.org/10.1016/j.matbio.2012.11.006
Chen, Z., Liu, G., Hossain, A., Danilova, I. G., Bolkov, M. A., et al. (2019). A co-expression network for differentially expressed genes in bladder cancer and a risk score model for predicting survival. Hereditas, 156, 24. https://doi.org/10.1186/s41065-019-0100-1
Li, Y., Chen, Y., Ma, Y., Nenkov, M., Haase, D., & Petersen, I. (2018). Collagen prolyl hydroxylase 3 has a tumor suppressive activity in human lung cancer. Experimental Cell Research, 363, 121–128. https://doi.org/10.1016/j.yexcr.2017.12.020
Xue, W., Sun, C., Yuan, H., Yang, X., Zhang, Q., et al. (2022). Establishment and analysis of an individualized emt-related gene signature for the prognosis of breast cancer in female patients. Disease Markers, 2022, 1289445. https://doi.org/10.1155/2022/1289445
Zhou, P., Liu, Z., Hu, H., Lu, Y., Xiao, J., et al. (2022). Comprehensive analysis of senescence characteristics defines a novel prognostic signature to guide personalized treatment for clear cell renal cell carcinoma. Frontiers in Immunology, 13, 901671. https://doi.org/10.3389/fimmu.2022.901671
Consortium G. (2013). The genotype-tissue expression (gtex) project. Nature Genetics, 45, 580-585.https://doi.org/10.1038/ng.2653
Nusinow, D. P., Szpyt, J., Ghandi, M., Rose, C. M., Mcdonald, E. R., et al. (2020). Quantitative proteomics of the cancer cell line encyclopedia. Cell, 180, 387–402. https://doi.org/10.1016/j.cell.2019.12.023
Tomczak, K., Czerwinska, P., & Wiznerowicz, M. (2015). The cancer genome atlas (tcga): An immeasurable source of knowledge. Contemp Oncol (Pozn), 19, A68–A77. https://doi.org/10.5114/wo.2014.47136
Goldman, M. J., Craft, B., Hastie, M., Repecka, K., Mcdade, F., et al. (2020). Visualizing and interpreting cancer genomics data via the xena platform. Nature Biotechnology, 38, 675–678. https://doi.org/10.1038/s41587-020-0546-8
Cerami, E., Gao, J., Dogrusoz, U., Gross, B. E., Sumer, S. O., et al. (2012). The cbio cancer genomics portal: An open platform for exploring multidimensional cancer genomics data. Cancer Discovery, 2, 401–404. https://doi.org/10.1158/2159-8290.CD-12-0095
Zeng, D., Li, M., Zhou, R., Zhang, J., Sun, H., et al. (2019). Tumor microenvironment characterization in gastric cancer identifies prognostic and immunotherapeutically relevant gene signatures. Cancer Immunology Research, 7, 737–750. https://doi.org/10.1158/2326-6066.CIR-18-0436
Li, T., Fu, J., Zeng, Z., Cohen, D., Li, J., et al. (2020). Timer2.0 for analysis of tumor-infiltrating immune cells. Nucleic Acids Research, 48, W509–W514. https://doi.org/10.1093/nar/gkaa407
Yang, W., Soares, J., Greninger, P., Edelman, E. J., Lightfoot, H., et al. (2013). Genomics of drug sensitivity in cancer (gdsc): A resource for therapeutic biomarker discovery in cancer cells. Nucleic Acids Research, 41, D955–D961. https://doi.org/10.1093/nar/gks1111
Lv, W., Shi, L., Pan, J., & Wang, S. (2022). Comprehensive prognostic and immunological analysis of cct2 in pan-cancer. Frontiers in Oncology, 12, 986990. https://doi.org/10.3389/fonc.2022.986990
Chen, F., Fan, Y., Cao, P., Liu, B., Hou, J., et al. (2021). Pan-cancer analysis of the prognostic and immunological role of hsf1: A potential target for survival and immunotherapy. Oxidative Medicine and Cellular Longevity, 2021, 5551036. https://doi.org/10.1155/2021/5551036
Erbas, I. M., Ilgun, G. D., Manav, K. Z., Koc, A., Unuvar, T., et al. (2022). Clinical, genetic characteristics and treatment outcomes of children and adolescents with osteogenesis imperfecta: A two-center experience. Connective Tissue Research, 63, 349–358. https://doi.org/10.1080/03008207.2021.1932853
de Souza, L. T., Nunes, R. R., de Azevedo, M. O., & Maria, F. T. (2021). A new case of osteogenesis imperfecta type viii and retinal detachment. American Journal of Medical Genetics. Part A, 185, 238–241. https://doi.org/10.1002/ajmg.a.61934
Tang, C., Fang, M., Tan, G., Zhang, S., Yang, B., et al. (2022). Discovery of novel circulating immune complexes in lupus nephritis using immunoproteomics. Frontiers in Immunology, 13, 850015. https://doi.org/10.3389/fimmu.2022.850015
Gawel, D. R., Lee, E. J., Li, X., Lilja, S., Matussek, A., et al. (2019). An algorithm-based meta-analysis of genome- and proteome-wide data identifies a combination of potential plasma biomarkers for colorectal cancer. Science and Reports, 9, 15575. https://doi.org/10.1038/s41598-019-51999-9
Xiong, C., Wang, G., & Bai, D. (2020). A novel prognostic models for identifying the risk of hepatocellular carcinoma based on epithelial-mesenchymal transition-associated genes. Bioengineered, 11, 1034–1046. https://doi.org/10.1080/21655979.2020.1822715
Zhang, Y., Li, C. Y., Pan, M., Li, J. Y., Ge, W., et al. (2021). Exploration of the key proteins of high-grade intraepithelial neoplasia to adenocarcinoma sequence using in-depth quantitative proteomics analysis. J Oncol, 2021, 5538756. https://doi.org/10.1155/2021/5538756
He, W., Lin, S., Guo, Y., Wu, Y., Zhang, L. L., et al. (2022). Targeted demethylation at znf154 promotor upregulates znf154 expression and inhibits the proliferation and migration of esophageal squamous carcinoma cells. Oncogene. https://doi.org/10.1038/s41388-022-02366-y
Avram, E. G., Moatar, I. A., Miok, V., Baderca, F., Samoila, C., et al. (2022). Gene network analysis of the transcriptome impact of methylated micrornas on oral squamous cell carcinoma. Advances in Clinical and Experimental Medicine. https://doi.org/10.17219/acem/151911
Xu, Q., Lan, X., Lin, H., Xi, Q., & Wang, M., et al. (2022). Tumor microenvironment-regulating nanomedicine design to fight multi-drug resistant tumors, Wiley Interdisciplinary Reviews Nanomedicine and Nanobiotechnology, e1842. https://doi.org/10.1002/wnan.1842.
Luo, J., Xie, Y., Zheng, Y., Wang, C., Qi, F., et al. (2020). Comprehensive insights on pivotal prognostic signature involved in clear cell renal cell carcinoma microenvironment using the estimate algorithm. Cancer Medicine, 9, 4310–4323. https://doi.org/10.1002/cam4.2983
Yoshihara, K., Shahmoradgoli, M., Martinez, E., Vegesna, R., Kim, H., et al. (2013). Inferring tumour purity and stromal and immune cell admixture from expression data. Nature Communications, 4, 2612. https://doi.org/10.1038/ncomms3612
Signorini, L., Delbue, S., Ferrante, P., & Bregni, M. (2016). Review on the immunotherapy strategies against metastatic colorectal carcinoma. Immunotherapy-Uk, 8, 1245–1261. https://doi.org/10.2217/imt-2016-0045
Acknowledgements
We acknowledge the TCGA, GTEx, CCLE, TIMER2.0, cBioPortal, and GDSC databases for free use.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
Conceptualization, Yongjie Li; Formal analysis, Yongjie Li and Ting Wang; Software analyses, Ting Wang; Visualization, Feng Jiang; Writing – original draft, Yongjie Li; Writing – review & editing, Yongjie Li.
Corresponding author
Ethics declarations
Ethics Statement
All the experiments were conducted in accordance with the ethical guidelines of Shaoyang University.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare that there are no conflict of interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, Y., Wang, T. & Jiang, F. Pan-Cancer Analysis of P3H1 and Experimental Validation in Renal Clear Cell Carcinoma. Appl Biochem Biotechnol (2024). https://doi.org/10.1007/s12010-023-04845-8
Accepted:
Published:
DOI: https://doi.org/10.1007/s12010-023-04845-8