Abstract
Objective
The COVID-19 pandemic has been recognized as severe acute respiratory syndrome, one of the worst and disastrous infectious diseases in human history. Until now, there is no cure to this contagious infection although some multinational pharmaceutical companies have synthesized the vaccines and injecting them into humans, but a drug treatment regimen is yet to come.
Aim
Among the multiple areas of SARS-CoV-2 that can be targeted, protease protein has significant values due to its essential role in viral replication and life. The repurposing of FDA-approved drugs for the treatment of COVID-19 has been a critical strategy during the pandemic due to the urgency of effective therapies. The novelty in this work refers to the innovative use of existing drugs with greater safety, speed, cost-effectiveness, broad availability, and diversity in the mechanism of action that have been approved and developed for other medical conditions.
Methods
In this research work, we have engaged drug reprofiling or drug repurposing to recognize possible inhibitors of protease protein 6M03 in an instantaneous approach through computational docking studies.
Results
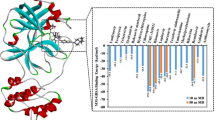
We screened 16 FDA-approved anti-viral drugs that were known for different viral infections to be tested against this contagious novel strain. Through these reprofiling studies, we come up with 5 drugs, namely, Delavirdine, Fosamprenavir, Imiquimod, Stavudine, and Zanamivir, showing excellent results with the negative binding energies in Kcal/mol as − 8.5, − 7.0, − 6.8, − 6.8, and − 6.6, respectively, in the best binding posture. In silico studies allowed us to demonstrate the potential role of these drugs against COVID-19.
Conclusion
In our study, we also observed the nucleotide sequence of protease protein consisting of 316 amino acid residues and the influence of these pronouncing drugs over these sequences. The outcome of this research work provides researchers with a track record for carrying out further investigational procedures by applying docking simulations and in vitro and in vivo experimentation with these reprofile drugs so that a better drug can be formulated against coronavirus.















Similar content being viewed by others
Data availability
The data associated with this work is included in the text of this article.
References
Wilson S, Steele S, Adeli K (2022) Innovative technological advancements in laboratory medicine: predicting the lab of the future. Biotechnol Biotechnol Equip 36:S9–S21
Yamini RS, Bhargavi RS, Bhuvaneswari K (2020) Drug repositioning in the treatment of Covid-19. J Med Pharm Allied Sci 9:2412–2417
Langley GR, Adcock IM, Busquet F et al (2017) Towards a 21st-century roadmap for biomedical research and drug discovery: consensus report and recommendations. Drug Discovery Today 22:327–339
Hughes JP, Rees S, Kalindjian SB, Philpott KL (2011) Principles of early drug discovery. Br J Pharmacol 162:1239–1249
Duarte RR, Copertino DC Jr, Iñiguez LP et al (2020) Repurposing FDA-approved drugs for covid-19 using a data-driven approach
Meng F, Liang Z, Zhao K, Luo C (2020) Drug design targeting active posttranslational modification protein isoforms. Med Res Rev
Sahoo BM, Ravi Kumar B, Sruti J et al (2021) Drug repurposing strategy (DRS): emerging approach to identify potential therapeutics for treatment of novel coronavirus infection. Front Mol Biosci 8:628144
Krishnamurthy N, Grimshaw AA, Axson SA et al (2022) Drug repurposing: a systematic review on root causes, barriers and facilitators. BMC Health Serv Res 22:1–17
Zhang Y-F, Wang X, Kaushik AC et al (2020) SPVec: a Word2vec-inspired feature representation method for drug-target interaction prediction. Front Chem 7:895
Sindhu S, Murugan S (2020) Challenges and opportunities in repurposing of drugs: mini review. Bangladesh J Med Sci 19:365–371
Lee HS, Bae T, Lee J-H et al (2012) Rational drug repositioning guided by an integrated pharmacological network of protein, disease and drug. BMC Syst Biol 6:1–10
Cheng F, Lu W, Liu C et al (2019) A genome-wide positioning systems network algorithm for in silico drug repurposing. Nat Commun 10:1–14
Meng Y, Jin M, Tang X, Xu J (2021) Drug repositioning based on similarity constrained probabilistic matrix factorization: COVID-19 as a case study. Appl Soft Comput 107135
Su L, Ma X, Yu H et al (2020) The different clinical characteristics of corona virus disease cases between children and their families in China–the character of children with COVID-19. Emerg Microbes Infect 9:707–713
Subbarao K, Mahanty S (2020) Respiratory virus infections: understanding COVID-19. Immunity 52:905–909
Correia G, Rodrigues L, Da Silva MG, Gonçalves T (2020) Airborne route and bad use of ventilation systems as non-negligible factors in SARS-CoV-2 transmission. Med Hypotheses 141:109781
Brito IL, Gurry T, Zhao S et al (2019) Transmission of human-associated microbiota along family and social networks. Nat Microbiol 4:964–971
Wong NA, Saier MH (2021) The SARS-coronavirus infection cycle: a survey of viral membrane proteins, their functional interactions and pathogenesis. Int J Mol Sci 22:1308
Giannis D, Ziogas IA, Gianni P (2020) Coagulation disorders in coronavirus infected patients: COVID-19, SARS-CoV-1, MERS-CoV and lessons from the past. J Clin Virol 127:104362
Chen B, Tian E-K, He B et al (2020) Overview of lethal human coronaviruses. Signal Transduct Target Ther 5:1–16
Hafeez A, Ahmad S, Siddqui SA et al (2020) A review of COVID-19 (coronavirus disease-2019) diagnosis, treatments and prevention. EJMO 4:116–125
Ramadan N, Shaib H (2019) Middle East respiratory syndrome coronavirus (MERS-CoV): a review. Germs 9:35
Ding Q, Lu P, Fan Y et al (2020) The clinical characteristics of pneumonia patients coinfected with 2019 novel coronavirus and influenza virus in Wuhan, China. J Med Virol 92:1549–1555
Bun SS, Taghji P, Courjon J et al (2020) QT interval prolongation under hydroxychloroquine/azithromycin association for inpatients with SARS-CoV-2 lower respiratory tract infection. Clin Pharmacol Ther 108:1090–1097
Holmes EA, O'Connor RC, Perry VH et al (2020) Multidisciplinary research priorities for the COVID-19 pandemic: a call for action for mental health science. The Lancet Psychiatr
Lobo-Galo N, Gálvez-Ruíz J-C, Balderrama-Carmona AP et al (2021) Recent biotechnological advances as potential intervention strategies against COVID-19. 3 Biotech 11:1–21
Bolarin JA, Oluwatoyosi MA, Orege JI et al (2021) Therapeutic drugs for SARS-CoV-2 treatment: current state and perspective. Int Immunopharmacol 90
Bhatti JS, Bhatti GK, Khullar N et al (2020) Therapeutic strategies in the development of anti-viral drugs and vaccines against SARS-CoV-2 infection. Mol Neurobiol 57:4856–4877
Mittal L, Kumari A, Srivastava M et al (2020) Identification of potential molecules against COVID-19 main protease through structure-guided virtual screening approach. J Biomol Struct Dynamics 1–19
Zafar R, Zubair M, Ali S et al (2020) Zinc metal carboxylates as potential anti-Alzheimer’s candidate: in vitro anticholinesterase, antioxidant and molecular docking studies. J Biomol Struct Dynamics 1–11
Chandel V, Raj S, Rathi B, Kumar D (2020) In silico identification of potent COVID-19 main protease inhibitors from FDA approved antiviral compounds and active phytochemicals through molecular docking: a drug repurposing approach
Ibrahim IM, Abdelmalek DH, Elshahat ME, Elfiky AA (2020) COVID-19 spike-host cell receptor GRP78 binding site prediction. J Infect 80:554–562
Elmezayen AD, Al-Obaidi A, Şahin AT, Yelekçi K (2020) Drug repurposing for coronavirus (COVID-19): in silico screening of known drugs against coronavirus 3CL hydrolase and protease enzymes. J Biomol Struct Dynamics 1–13
Xue X, Yu H, Yang H et al (2008) Structures of two coronavirus main proteases: implications for substrate binding and antiviral drug design. J Virol 82:2515–2527
Cannalire R, Cerchia C, Beccari AR et al (2020) Targeting SARS-CoV-2 proteases and polymerase for COVID-19 treatment: state of the art and future opportunities. J Med Chem
Barbu MG, Condrat CE, Thompson DC et al (2020) MicroRNA involvement in signaling pathways during viral infection. Front Cell Develop Biol 8:143
Bartas M, Brázda V, Bohálová N et al (2020) In-depth bioinformatic analyses of nidovirales including human SARS-CoV-2, SARS-CoV, MERS-CoV viruses suggest important roles of non-canonical nucleic acid structures in their lifecycles. Front Microbiol 11:1583
Singh TU, Parida S, Lingaraju MC et al (2020) Drug repurposing approach to fight COVID-19. Pharmacol Rep 1–30
Acknowledgements
We are thankful to Princess Nourah bint Abdularahman University Researchers Supporting Project number (PNURSP2023R47), Princess Nourah bint Abdularahman University, Riyadh, Saudi Arabia.
Funding
Princess Nourah bint Abdularahman University Researchers Supporting Project number (PNURSP2023R47), Princess Nourah bint Abdularahman University, Riyadh, Saudi Arabia
Author information
Authors and Affiliations
Contributions
WW, RZ, and MSJ carried out the computational studies. TSA, NA, and AR involved in data collection, analysis, and editing of text. HK involved in analysis and helped in the drafting of research papers. MSJ and AR performed the refinement of the manuscript to be prepared for publication.
Corresponding authors
Ethics declarations
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Waseem, W., Zafar, R., Jan, M.S. et al. Drug repurposing of FDA-approved anti-viral drugs via computational screening against novel 6M03 SARS-COVID-19. Ir J Med Sci 193, 73–83 (2024). https://doi.org/10.1007/s11845-023-03473-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11845-023-03473-9