Abstract
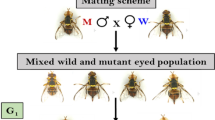
Heterosis has been exploited to enhance the yield and adaptability in various shellfish species; however, the molecular basis of it remains unclear. The Pacific oyster Crassostrea gigas is one of the most economically important aquaculture species, and its productive traits can be improved by hybridization. Here, an intraspecific cross between orange shell (O, 10th generation) and ‘Haida No. 1’ (H, 13th generation) of C. gigas was performed to assess the heterosis of survival trait. Survival rates of hybrid family (OH) and inbred families (HH and OO) were compared at larval stage, and eyed-pediveliger larvae of three families were subjected to transcriptome analysis. The analysis results of best-parent heterosis and mid-parent heterosis showed that the hybrid family exhibited a high heterosis in survival relative to the parental families. The OH-M (OH vs. OO) and OH-P (OH vs. HH) had 425 and 512 differentially expressed genes (DEGs), respectively. Functional enrichment analysis of these DEGs revealed that the significantly enriched genes function in virion binding, C-type lectin receptor signaling pathway, cellular defense response and other immune-related processes, which involves perlucin-like protein, CD209 antigen-like protein, ZNFX1, caspase-3 and acan genes. These differentially expressed genes in OH-M and OH-P, together with the immune-related processes mentioned above may play an important role in the larval survival of C. gigas. In addition, three genes (CYP450, fucolectin and perlucin-like) are associated with the orange shell and low survival of maternal oyster OO. These findings provide support for the application of hybrid with superior survival and will facilitate the understanding of heterosis formation in the Pacific oyster.
Similar content being viewed by others
References
Agnew, M. V, Friedman, C. S., Langdon, C., Divilov, K., School-field, B., Morga, B., et al., 2020. Differential mortality and high viral load in naive pacific oyster families exposed to OsHV-1 suggests tolerance rather than resistance to infection. Pathogens, 9 (12): 1057, DOI: https://doi.org/10.3390/pathogens9121057.
Amraei, R., Yin, W., Napoleon, M. A., Suder, E. L., Berrigan, J., Zhao, Q., et al., 2021. CD209L/L-SIGN and CD209/DC-SIGN act as receptors for SARS-CoV-2. ACS Central Science, 7 (7): 1156–1165, DOI: https://doi.org/10.1021/acscentsci.0c01537.
Birchler, J. A., Yao, H., Chudalayandi, S., Vaiman, D., and Veitia, R. A., 2010. Heterosis. The Plant Cell, 22 (7): 2105–2112, DOI: https://doi.org/10.1105/tpc.110.076133.
Bruce, A. B., 1910. The Mendelian theory of heredity and the augmentation of vigor. Science, 32 (827): 627–628, DOI: https://doi.org/10.1126/science.32.827.627-a.
Burdon, D., Callaway, R., Elliott, M., Smith, T., and Wither, A., 2014. Mass mortalities in bivalve populations: A review of the edible cockle Cerastoderma edule (L.). Estuarine Coastal and Shelf Science, 150: 271–280, DOI: https://doi.org/10.1016/j.ecss.2014.04.011.
Chen, S. Y., Zhang, Z. Y., Ji, H. J., Xu, S. X., Yang, Y. X., Jia, C. F., et al., 2020. Transcriptome profiles of F1 hybrids (Acanthopagrus schlegelii ♂ × Pagrus major ♀) and parents reveal hybrid effects on individual development. Aquaculture Research, 51 (10): 4011–4021, DOI: https://doi.org/10.1111/are.14744.
Du, X., Wang, G. H., Su, Y. L., Zhang, M., and Hu, Y. H., 2018. Black rockfish C-type lectin, SsCTL4: A pattern recognition receptor that promotes bactericidal activity and virus escape from host immune defense. Fish & Shellfish Immunology, 79: 340–350, DOI: https://doi.org/10.1016/j.fsi.2018.05.033.
Garcia, C., Thebault, A., Degremont, L., Arzul, I., Miossec, L., Robert, M., et al., 2011. Ostreid herpesvirus 1 detection and relationship with Crassostrea gigas spat mortality in France between 1998 and 2006. Veterinary Research, 42: 73, DOI: https://doi.org/10.1186/1297-9716-42-73.
Ge, X. M., Chen, W. H., Song, S. H., Wang, W. W., Hu, S. N., and Yu, J., 2008. Transcriptomic profiling of mature embryo from an elite super-hybrid rice LYP9 and its parental lines. BMC Plant Biology, 8: 114, DOI: https://doi.org/10.1186/1471-2229-8-114.
Guo, H. B., Mendrikahy, J. N., Xie, L., Deng, J. F., Lu, Z. J., Wu, J. W., et al., 2017. Transcriptome analysis of neo-tetraploid rice reveals specific differential gene expressions associated with fertility and heterosis. Scientific Reports, 7: 40139, DOI: https://doi.org/10.1038/srep40139.
Han, Z. Q., Li, Q., Liu, S. K., Yu, H., and Kong, L. F., 2019. Genetic variability of an orange-shell line of the Pacific oyster Crassostrea gigas during artificial selection inferred from microsatellites and mitochondrial COI sequences. Aquaculture, 508: 159–166, DOI: https://doi.org/10.1016/j.aquaculture.2019.04.074.
Jiang, G. W., Zhou, J. M., Cheng, G., Meng, L. G., Chi, Y., Xu, C. X., et al., 2022. Examination of survival, physiological parameters and immune response in relation to the thermo-resistant heterosis of hybrid oysters derived from Crassostrea gigas and C. angulata. Aquaculture, 559: 738454, DOI: https://doi.org/10.1016/j.aquaculture.2022.738454.
Kong, L. F., Song, S. L., and Li, Q., 2017. The effect of inter-strain hybridization on the production performance in the Pacific oyster Crassostrea gigas. Aquaculture, 472 (S1): 44–49, DOI: https://doi.org/10.1016/j.aquaculture.2016.07.018.
Li, Q., Wang, Q. Z., Liu, S. K., and Kong, L. F., 2011. Selection response and realized heritability for growth in three stocks of the Pacific oyster Crassostrea gigas. Fisheries Science, 77 (4): 643–648, DOI: https://doi.org/10.1007/s12562-011-0369-0.
Li, Z. Z., Li, Q., Liu, S. K., Han, Z. Q., Kong, L. F., and Yu, H., 2021. Integrated analysis of coding genes and non-coding RNAs associated with shell color in the Pacific oyster (Crassostrea gigas). Marine Biotechnology, 23 (3): 417–429, DOI: https://doi.org/10.1007/s10126-021-10034-7.
Liang, Y. X., Zhang, G. H., Jiang, G. W., Hu, Y. M., Fang, J. F., Chi, Y., et al., 2022. Hybridization between ‘Haida No. 1’ and Orange-shell line of the Pacific oyster reveals high heterosis in survival. Aquaculture, 551: 737945, DOI: https://doi.org/10.1016/j.aquaculture.2022.737945.
Liao, Y., Smyth, G. K., and Shi, W., 2014. FeatureCounts: An efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics, 30 (7): 923–930, DOI: https://doi.org/10.1093/bioinformatics/btt656.
Lippman, Z. B., and Zamir, D., 2007. Heterosis: Revisiting the magic. Trends in Genetics, 23 (2): 60–66, DOI: https://doi.org/10.1016/j.tig.2006.12.006.
Martins, E., Figueras, A., Novoa, B., Santos, R. S., Moreira, R., and Bettencourt, R., 2014. Comparative study of immune responses in the deep-sea hydrothermal vent mussel Bathymodiolus azoricus and the shallow-water mussel Mytilus galloprovincialis challenged with Vibrio bacteria. Fish & Shellfish Immunology, 40 (2): 485–499, DOI: https://doi.org/10.1016/j.fsi.2014.07.018.
Meng, L. X., Li, Q., Xu, C. X., Liu, S. K., Kong, L. F., and Yu, H., 2021. Hybridization improved stress resistance in the Pacific oyster: Evidence from physiological and immune responses. Aquaculture, 545: 737227, DOI: https://doi.org/10.1016/j.aquaculture.2021.737227.
Mohd-Shamsudin, M. I., Kang, Y., Zhao, L. L., Tan, T. T., Kwong, Q. B., Liu, H., et al., 2013. In-depth tanscriptomic analysis on giant freshwater prawns. PLoS One, 8 (5): e60839, DOI: https://doi.org/10.1371/journal.pone.0060839.
Moreira, R., Milan, M., Balseiro, P., Romero, A., Babbucci, M., Figueras, A., et al., 2014. Gene expression profile analysis of Manila clam (Ruditapes philippinarum) hemocytes after a Vibrio alginolyticus challenge using an immune-enriched oligo-microarray. BMC Genomics, 15: 267, DOI: https://doi.org/10.1186/1471-2164-15-267.
Mundy, N. I., Stapley, J., Bennison, C., Tucker, R., Twyman, H., Kim, K. W., et al., 2016. Red carotenoid coloration in the Zebra Finch is controlled by a cytochrome P450 gene cluster. Current Biology, 26 (11): 1435–1440, DOI: https://doi.org/10.1016/j.cub.2016.04.047.
Pfaffl, M. W., Horgan, G. W., and Dempfle, L., 2002. Relative expression software tool (REST) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Research, 30 (9): e36, DOI: https://doi.org/10.1093/nar/30.9.e36.
Pomin, V. H., and Mulloy, B., 2018. Glycosaminoglycans and Proteoglycans. Pharmaceuticals, 11 (1): 27, DOI: https://doi.org/10.3390/ph11010027.
Qin, Y. P., Liao, Q. L., Shi, G. P. Y., Yang, Y., Zhou, Y. Y., Li, J., et al., 2022. Comparison of growth, survival and fertility of the southern and northern populations of Crassostrea ariakensis and their hybrids in southern China. Aquaculture, 549: 737744, DOI: https://doi.org/10.1016/j.aquaculture.2021.737744.
Sekino, Y., Han, X. R., Kawaguchi, T., Babasaki, T., Goto, K., Inoue, S., et al., 2019. TUBB3 reverses resistance to docetaxel and cabazitaxel in prostate cancer. International Journal of Molecular Sciences, 20 (16): 3936, DOI: https://doi.org/10.3390/ijms20163936.
Shahzad, K., Zhang, X. X., Guo, L. P., Qi, T. X., Bao, L. S., Zhang, M., et al., 2020. Comparative transcriptome analysis between inbred and hybrids reveals molecular insights into yield heterosis of upland cotton. BMC Plant Biology, 20 (1): 239, DOI: https://doi.org/10.1186/s12870-020-02442-z.
Shang, L. G., Liang, Q. Z., Wang, Y. M., Zhao, Y. P., Wang, K. B., and Hua, J. P., 2016. Epistasis together with partial dominance, over-dominance and QTL by environment interactions contribute to yield heterosis in upland cotton. Theoretical and Applied Genetics, 129 (7): 1429–1446, DOI: https://doi.org/10.1007/s00122-016-2714-2.
Shao, Y. N., Che, Z. J., Xing, R. L., Wang, Z. D., Zhang, W. W., Zhao, X. L., et al., 2018. Divergent immune roles of two fucolectin isoforms in Apostichopus japonicus. Developmental & Comparative Immunology, 89: 1–6, DOI: https://doi.org/10.1016/j.dci.2018.07.028.
Solomieu, V. B., Renault, T., and Travers, M. A., 2015. Mass mortality in bivalves and the intricate case of the Pacific oyster, Crassostrea gigas. Journal of Invertebrate Pathology, 131: 2–10, DOI: https://doi.org/10.1016/j.jip.2015.07.011.
Song, G. S., Zhai, H. L., Peng, Y. G., Zhang, L., Wei, G., Chen, X. Y., et al., 2010. Comparative transcriptional profiling and preliminary study on heterosis mechanism of super-hybrid rice. Molecular Plant, 3 (6): 1012–1025, DOI: https://doi.org/10.1093/mp/ssq046.
Song, J. L., and Wang, C. D., 2019. Transcriptomic and proteomic analyses of genetic factors influencing adductor muscle coloration in QN Orange scallops. BMC Genomics, 20 (1): 363, DOI: https://doi.org/10.1186/s12864-019-5717-y.
Sun, B. J., Dissel, D. V., Mo, I., Boysen, P., Haslene-Hox, H., and Lund, H., 2022. Identification of novel biomarkers of inflammation in Atlantic salmon (Salmo salar L.) by a plasma proteomic approach. Developmental & Comparative Immunology, 127: 104268, DOI: https://doi.org/10.1016/j.dci.2021.104268.
Tan, K., Liu, H. X., Ye, T., Ma, H. Y., Li, S. K., and Zheng, H. P., 2020. Growth, survival and lipid composition of Crassostrea gigas, C. angulata and their reciprocal hybrids cultured in southern China. Aquaculture, 516: 734524, DOI: https://doi.org/10.1016/j.aquaculture.2019.734524.
Ummanni, R., Lehnigk, U., Zimmermann, U., Woenckhaus, C., Walther, R., and Giebel, J., 2010. Immunohistochemical expression of caspase-1 and -9, uncleaved caspase-3 and -6, cleaved caspase-3 and -6 as well as Bcl-2 in benign epithelium and cancer of the prostate. Experimental and Therapeutic Medicine, 1 (1): 47–52, DOI: https://doi.org/10.3892/etm_00000008.
Wang, Y., Xue, Z., Yi, Q. L., Wang, H., Wang, L. L., Lu, G. X., et al., 2018a. A novel fucolectin from Apostichopus japonicus with broad PAMP recognition pattern. Fish & Shellfish Immunology, 77: 402–409, DOI: https://doi.org/10.1016/j.fsi.2018.04.013.
Wang, Y., Yuan, S. C., Jia, X., Ge, Y., Ling, T., Nie, M., et al., 2019. Mitochondria-localised ZNFX1 functions as a dsRNA sensor to initiate antiviral responses through MAVS. Nature Cell Biology, 21 (11): 1346–1356, DOI: https://doi.org/10.1038/s41556-019-0416-0.
Wang, Z. C., Cui, J., Song, J., Wang, H. Z., Gao, K. L., Qiu, X. M., et al., 2018b. Comparative transcriptome analysis reveals growth-related genes in juvenile Chinese sea cucumber, Russian sea cucumber, and their hybrids. Marine Biotechnology, 20 (2): 193–205, DOI: https://doi.org/10.1007/s10126-018-9796-6.
Whitlock, M. C., Ingvarsson, P. K., and Hatfield, T., 2000. Local drift load and the heterosis of interconnected populations. Heredity (Edinb), 84 (Pt 4): 452–457, DOI: https://doi.org/10.1046/j.1365-2540.2000.00693.x.
Xiao, Q. Z., Huang, Z. K., Shen, Y. W., Gan, Y., Wang, Y., Gong, S. H., et al., 2021. Transcriptome analysis reveals the molecular mechanisms of heterosis on thermal resistance in hybrid abalone. BMC Genomics, 22 (1): 650, DOI: https://doi.org/10.1186/s12864-021-07954-y.
Xu, J. C., Jiang, S., Li, Y. Q., Li, M. J., Cheng, Q., Zhao, D. P., et al., 2016. Caspase-3 serves as an intracellular immune receptor specific for lipopolysaccharide in oyster Crassostrea gigas. Developmental & Comparative Immunology, 61: 1–12, DOI: https://doi.org/10.1016/j.dci.2016.03.015.
Yang, J. M., Luo, S. J., Li, J. H., Zheng, Z., Du, X. D., and Deng, Y. W., 2018. Transcriptome analysis of growth heterosis in pearl oyster Pinctada fucata martensii. FEBS Open Bio, 8 (11): 1794–1803, DOI: https://doi.org/10.1002/2211-5463.12502.
Yin, X. S., and Hedgecock, D., 2021. Overt and concealed genetic loads revealed by QTL mapping of genotype-dependent viability in the Pacific oyster Crassostrea gigas. Genetics, 219 (4): iyab165, DOI: https://doi.org/10.1093/genetics/iyab165.
Zhang, G. S., Li, J., Zhang, J. J., Liang, X., Zhang, X. Y., Wang, T., et al., 2019. Integrated analysis of transcriptomic, miRNA and proteomic changes of a novel hybrid yellow catfish uncovers key roles for miRNAs in heterosis. Molecular & Cellular Proteomics, 18 (7): 1437–1453, DOI: https://doi.org/10.1074/mcp.RA118.001297.
Zhang, H., Jia, H. X., Xiong, P. P., Yao, G. Y., and He, M. X., 2022a. Transcriptome and enzyme activity analyses of tolerance mechanisms in pearl oyster (Pinctada fucata) under high-temperature stress. Aquaculture, 550: 737888, DOI: https://doi.org/10.1016/j.aquaculture.2022.737888.
Zhang, X. K., Fan, C., Li, J. L., Zhang, X. Z., Li, Q., and Wang, Z. P., 2022b. Transcriptome analysis of Crassostrea sikamea (♀) × Crassostrea gigas (♂) hybrids under hypoxia in occluded water. Frontiers in Marine Science, 9: 851098, DOI: https://doi.org/10.3389/fmars.2022.851098.
Zhang, X. Y., Wen, H. S., Wang, H. L., Ren, Y. Y., Zhao, J., and Li, Y., 2017. RNA-Seq analysis of salinity stress-responsive transcriptome in the liver of spotted sea bass (Lateolabrax maculatus). PLoS One, 12 (3): e0173238, DOI: https://doi.org/10.1371/journal.pone.0173238.
Zhao, Y., Hu, F. X., Zhang, X. G., Wei, Q. Y., Dong, J. L., Bo, C., et al., 2019. Comparative transcriptome analysis reveals important roles of nonadditive genes in maize hybrid An’nong 591 under heat stress. BMC Plant Biology, 19 (1): 273, DOI: https://doi.org/10.1186/s12870-019-1878-8.
Acknowledgements
This research was supported by the grants from the China Agriculture Research System Project (No. CARS-49), and the Earmarked Fund for Agriculture Seed Improvement Project of Shandong Province (No. 2020LZGC016).
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Yang, H., Li, Q. Transcriptome Analysis of Heterosis in Survival in the Hybrid Progenies of ‘Haida No. 1’ and Orange-Shelled Lines of the Pacific Oyster Crassostrea gigas. J. Ocean Univ. China 23, 199–208 (2024). https://doi.org/10.1007/s11802-024-5643-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11802-024-5643-8