Abstract
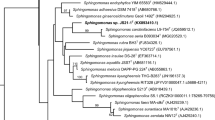
Sphingomonads are taxonomically heterogeneous bacteria possessing biologically diverse features. In the present study, the taxonomic status and some biological and phenotypic characteristics of sphingomonad strains isolated from stone and pome fruit trees in Mazandaran province were determined. The 115 sphingomonad-like strains isolated formed 13 phenotypic groups among which eight were identified as sphingomonads based on their 16S rDNA sequences. A representative from each phylogroup was used in multilocus sequence analysis (MLSA) employing atpD, dnaK, gap, recA, and rpoB housekeeping genes. MLSA analysis indicated that strains HM4, HM7, HM9, HM12, and HM20 belong to Sphingomonas, and the remaining strains HM5, HM16, and HM23 were members of the genus Novosphingobium. The strain HM7 was identified as Sphingomonas melonis and was capable of producing brown spot symptoms on melon fruit. The precise taxonomic position of the remaining strains, at the species level, remained unresolved. Nonetheless, strains HM4 and HM12 clustered with the group consisting of Sphingomonas sp. LK11 and Sphingomonas paucimobilis AIMST S2 at 100% bootstrap value. In the Novosphingobium clade, strains HM5 and HM23 clustered close to each other along with the possibly misidentified Sphingomonas sp. strain IC081 with high bootstrap supports. All strains produced siderophore and all but members of group HM20 induced a hypersensitive reaction on geranium leaves. However, the antibiotic sensitivity, biofilm formation, ice nucleation activity, and resistance to freezing/thawing cycles varied among the strains. This is the first comprehensive study on the taxonomy and some biological characteristics of sphingomonads associated with fruit trees in Iran.



Similar content being viewed by others
Abbreviations
- atpD :
-
ATP synthase subunit beta
- CAS–agar:
-
Chrome azorol S agar
- CTAB:
-
Cetyl trimethylammonium bromide
- dnaK :
-
heat-shock protein (molecular chaperone)
- gap :
-
Glyceraldehyde-3-phosphate dehydrogenase
- HR:
-
Hypersensitive reaction
- INA:
-
ice nucleation activity
- ML:
-
Maximum Likelihood
- MLSA:
-
Multi locus sequence analysis
- NA:
-
Nutrient agar
- OD:
-
Optical density
- ODc:
-
OD of the control
- PBS:
-
Phosphate buffer saline
- PCR:
-
Polymerase chain reaction
- Pss :
-
Pseudomonas syringae pv. Syringae
- recA :
-
DNA recombination/repair protein RecA
- SDW:
-
Sterile distilled water
- SNA:
-
Sucrose nutrient agar
- t:
-
tonnes
- TCP:
-
Tissue culture plate
- TM:
-
Tube method
- rpoB :
-
the beta subunit of RNA polymerase
References
Agrios GN (2005) Plant Pathology, 5th edn. Academic press, San Diego, CA. 992p
Albala K (2011) Food Cultures of the World Encyclopedia. ABC-CLIO. 240 p
Alexander DB, Zuberer DA (1991) Use of chrome azurol S reagents to evaluate siderophore production by rhizosphere bacteria. Biol Fert Soils 12:39–45. https://doi.org/10.1007/BF00369386
Asaf S, Khan AL, Khan MA, Al-Harrasi A, Lee IJ (2018) Complete genome sequencing and analysis of endophytic Sphingomonas sp. LK11 and its potential in plant growth. 3. Biotech 8:389. https://doi.org/10.1007/s13205-018-1403-z
Buonaurio R, Stravato VM, Kosako Y, Fujiwara N, Naka T, Kobayashi K, Cappelli C, Yabuuchi E (2002) Sphingomonas melonis sp. nov., a novel pathogen that causes brown spots on yellow Spanish melon fruits. Int J Syst Evol Microbiol 52:2081–2087. https://doi.org/10.1099/00207713-52-6-2081
Chen H (2012) Population structure and species description of aquatic Sphingomonadaceae. Dissertation at the Faculty of Biology Ludwig- Maximilians- University Munich, 122 p
Che-Noraini CH, Morad N, Norli I, Teng TT, Ogugbue CJ (2012) Methylene blue degradation by Sphingomonas paucimobilis under aerobic conditions. Water Air Soil Pollut 223:5131–5142. https://doi.org/10.1007/s11270-012-1264-8
Christensen GD, Simpson WA, Bisno AL, Beachy EH (1982) Adherence of slime producing strains of Staphylococcus epidermis to smooth surfaces. Infect Immun 37:318–326. https://doi.org/10.1128/IAI.37.1.318-326.1982
Deldavleh M, Bahmani K, Harighi B (2013) Bacterial leaf blight of Christ's thorn in Iran: a new disease caused by Sphingomonas sp. J Plant Pathol 95:75–78. https://doi.org/10.4454/JPP.V95I1.018
Dworkin M, Falkow S, Rosenberg E, Schleifer KH, Stackebrandt F (2006) The Prokaryotes, A Handbook on the Biology of Bacteria, Symbiotic Associations, Biotechnology, Applied Microbiology,3rd edn, vol 1. Springer Science + Business Media, Inc., New York
FAOSTAT (2018) Food and Agriculture Organization of the United Nations, FAOSTAT database, available athttp://faostat.fao.org/
Fernandez-Sanz AM, Rodicio MR, Gonzalez AJ (2016)Pseudomonas syringae pv. phaseolicola isolated from weeds in bean crop fields. Lett Appl Microbiol 62:344–348. https://doi.org/10.1111/lam.12556
Glaeser SP, Kampfer P (2014) The family Sphingomonadaceae. In: Rosenberg E, DeLong EF, Lory S, Stackebrandt E (eds) The prokaryotes- Alphaproteobacteria and Betaproteobacteria. Springer, Berlin, pp 641–707. https://doi.org/10.1007/978-3-642-30197-1_302
Gulati P, Ghosh M (2017) Biofilm forming ability of Sphingomonas paucimobilis isolated from community drinking water systems on plumbing materials used in water distribution. J Water Health 15:942–954. https://doi.org/10.2166/wh.2017.294
Guo C, Dang Z, Wong Y, Tam NF (2010) Biodegradation Ability and Dioxygenase Genes of PAH-DegradingSphingomonas and Mycobacterium strains isolated from mangrove sediments. Int Biodeterior Biodegradation 64:419–426. https://doi.org/10.1016/j.ibiod.2010.04.008
Hall TA (1999) BioEdit: A User-Friendly Biological Sequence Alignment Editor and Analysis Program for Windows 95/98/NT. Nucl Acids Symp Ser 41:95-98. https://doi.org/10.14601/PHYTOPATHOL_MEDITERR-14998U1.29
Halo BA, Khan AL, Waqas M, Al-Harrasi A, Hussain J, Ali L, Adnan M, Lee IJ (2015) Endophytic bacteria (Sphingomonas sp. LK11) and gibberellin can improve Solanum lycopersicum growth and oxidative stress under salinity. J Plant Interact 10:117–125. https://doi.org/10.1080/17429145.2015.1033659
Hirano SS, Upper CD (1983) Ecology and epidemiology of foliar bacterial plant pathogens. Annu Rev Phytopathol 21:243–269. https://doi.org/10.1146/annurev.py.21.090183.001331
Holmes B, Owen RJ, Evans A, Malnick H, Wilcox WR (1977)Pseudomonas paucimobilis, a new species isolated from human clinical specimens, the hospital environment, and other sources. Int J Syst Bacteriol 27:133–146. https://doi.org/10.1099/00207713-27-2-133
Huang Y, Feng H, Lu H, Zeng Y (2017) Novel 16S rDNA primers revealed the diversity and habitats-related community structure of sphingomonads in 10 different niches. Antonie Van Leeuwenhoek 110:877–889. https://doi.org/10.1007/s10482-017-0860-4
Huson DH, Bryant D (2006) Application of phylogenetic networks in evolutionary studies. Mol Biol Evol 23:254–267. https://doi.org/10.1093/molbev/msj030
Innerebner G, Knief C, Vorholt JA (2011) Protection of Arabidopsis thaliana against Leaf-PathogenicPseudomonas syringae by Sphingomonas Strains in a Controlled Model System. Appl Environ Microbiol 77:3202–3210. https://doi.org/10.1128/AEM.00133-11
Jacques MA, Arlat M, Boulanger A, Boureau T, Carrère S, Cesbron S, Chen NW, Cociancich S, Darrasse A, Denancé N, Fischer-Le Saux M, Gagnevin L, Koebnik R, Lauber E, Noël LD, Pieretti I, Portier P, Pruvost O, Rieux A et al (2016) Using Ecology, Physiology, and Genomics to Understand Host Specificity in Xanthomonas. Annu Rev Phytopathol 54:163–187. https://doi.org/10.1146/annurev-phyto-080615-100147
Jogler M, Chen H, Simon J, Rohde M, Busse HJ, Klenk HP, Tindall BJ, Overmann J (2013) Description of Sphingorhabdus planktonica gen. nov., sp. nov. and reclassification of three related members of the genus Sphingopyxis in the genus Sphingorhabdus gen. nov. Int J Syst Evol Microbiol 63:1342–1349. https://doi.org/10.1099/ijs.0.043133-0
Kado CI (2010) Plant Bacteriology. St. Paul, MN, USA, 336 p
Khan AL, Waqas M, Asaf S, Kamran M, Shahzad R, Bilal S, Khan MA, Kang SM, Kim YH, Ran BW, Al-Rawahi A (2017) Plant growth-promoting endophyte Sphingomonas sp. LK11 alleviates salinity stress in Solanum pimpinellifolium. Environ Exp Bot 133:58–69. https://doi.org/10.1016/j.envexpbot.2016.09.009
Kim YJ, Park JY, Balusamy SR, Huo Y, Nong LK, Le HT, Yang DC, Kim D (2020) Comprehensive genome analysis on the novel species Sphingomonas panacis DCY99T reveals insights into Iron tolerance of Ginseng. Int J Mol Sci 21:2019. https://doi.org/10.3390/ijms21062019
Kini K, Agnimonhan R, Dossa R, Soglonou B, Gbogbo V, Ouedraogo I, Kpemoua K, Traoré M, Silue D (2017) First report of Sphingomonas sp. causing bacterial leaf blight of rice in Benin, Burkina Faso, The Gambia, Ivory Coast, Mali, Nigeria, Tanzania and Togo. New Dis Rep 35:32. https://doi.org/10.5197/j.2044-0588.2017.035.032
Klement Z (1963) Rapid detection of the pathogenicity of phytopathogenic Pseudomonads. Nature 199:299–300. https://doi.org/10.1038/199299b0
Krishnan R, Menon R, Likhith R, Busse H, Tanaka N, Krishnamurthi S, Rameshkumar N (2017)Novosphingobium pokkalii sp nov, a novel rhizosphere-associated bacterium with plant beneficial properties isolated from saline-tolerant pokkali rice. Res Microbiol 168:113–121. https://doi.org/10.1016/j.resmic.2016.09.001
Lee JC, Whang KS (2020)Sphingomonas segetis sp. nov., isolated from spinach farming field soil. Int J Syst Evol Microbiol 70:3905–3911. https://doi.org/10.1099/ijsem.0.004257
Li X, He J, Li S (2007) Isolation of a chlorpyrifos-degrading bacterium, Sphingomonas sp. strain Dsp-2, and cloning of the mpd gene. Res Microbiol 158:143–149. https://doi.org/10.1016/j.resmic.2006.11.007
Li Y, GuoLM CJP, Yang XQ, Xie SJ, Liu H, Piao CG (2019)Sphingobacterium corticibacter sp. nov., isolated from bark of Populus × euramericana. Int J Syst Evol Microbiol 69:1870–1874. https://doi.org/10.1099/ijsem.0.003148
Li Y, Wang S, Chang JP, Bian DR, Guo LM, Yang XQ (2020)Sphingomonas cortices sp. nov., and Sphingobacterium corticibacterium sp. nov., from bark canker. Int J Syst Evol Microbiol 70:5627–5633. https://doi.org/10.1099/ijsem.0.004451
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452. https://doi.org/10.1093/bioinformatics/btp187
Lindow SE, Arny DC, Upper CD (1978)Erwinia herbicola: a bacterial ice nucleus active in increasing frost injury to corn. Phytopathology 68:523–527. https://doi.org/10.1094/Phyto-68-523
Liu F, Zhan RL, He ZQ (2018) First report of bacterial dry rot of Mango caused by Sphingomonas sanguinis in China. Plant Dis 102:2632. https://doi.org/10.1094/PDIS-04-18-0589-PDN
Luo Y, Wang F, Huang Y, Zhou M, Gao J, Yan T, Sheng H, An L (2019)Sphingomonas sp. Cra20 Increases Plant Growth Rate and Alters Rhizosphere Microbial Community Structure of Arabidopsis thaliana Under Drought Stress. Front Microbiol 10:1221. https://doi.org/10.3389/fmicb.2019.01221
Matsumoto H, Fan X, Wang Y, Kusstatscher P, Duan J, Wu S, Chen S, Qiao K, Wang Y, Ma B, Zhu G, Hashidoko Y, Berg G, Cernava T, Wang M (2021) Bacterial seed endophyte shapes disease resistance in rice. Nat Plants 7:60–72. https://doi.org/10.1038/s41477-020-00826-5
Maruyama T, Park H, Ozawa K, Tanaka Y, Sumino T, Hamana K, Hiraishi A, Kato K (2006) Sphingosinicella microcystinivorans gen. nov., sp. nov., a microcystin-degrading bacterium. Int J Syst Evol Microbiol 56:85–89. https://doi.org/10.1099/ijs.0.63789-0
Nikravesh Z, Arab F, Rezaeian V, Rahimian H (2006a a) Isolation of Sphingomonas sp. From peach and willow trees in Kashan. Proc. 17th Iranian Plant Protec Congr, (Abst.). 479
Nikravesh Z, Arab F, Rezaeian V, Rahimian H (2006b b) Sphingomonas spp. as epiphytic bacteria of peach trees and broccoli plants in Golestan and Mazandaran. Proc. 17th Iranian Plant Protec Congr, (Abst.). 475
Pan F, Meng Q, Wang Q, Luo S, Chen B, Khan KY, Yang X, Feng Y (2016) Endophytic bacterium Sphingomonas SaMR12 promotes cadmium accumulation by increasing glutathione biosynthesis in Sedum alfredii Hance. Chemosphere 154:358–366. https://doi.org/10.1016/j.chemosphere.2016.03.120
Rahimian H (1994) Bacterial leaf spot of zinnia in Mazandaran. Plant protec 17:1-11. (In Persian with English abstract). https://doi.org/10.22055/PPR.1995.12802
Rigano LA, Siciliano F, Enrique R, Sendín L, Filippone P, Torres PS, Qüesta J, Dow JM, Castagnaro AP, Vojnov AA, Marano MR (2007) Biofilm formation, epiphytic fitness, and canker development in Xanthomonas axonopodis pv. citri. Mol Plant-Microbe Interact 20:1222–1230. https://doi.org/10.1094/MPMI-20-10-1222
Schaad NW, Jones JB, Chun W (2001) Laboratory Guide for Identification of Plant Pathogenic Bacteria, 3rd edn. APS, St. Paul, MN, USA, 373 p. https://doi.org/10.1046/j.1365-3059.2001.00635.x
Silva C, Cabral JM, Van Keulen F (2004) Isolation of a beta-carotene over-producing soil bacterium, Sphingomonas sp. Biotechnol Lett 26:257–262. https://doi.org/10.1023/b:bile.0000013716.20116.dc
Stepanović S, Vuković D, Hola V, Bonaventura GD, Djukić S, Ćirković I, Ruzicka F (2007) Quantification of biofilm in microtiter plates: overview of testing conditions and practical recommendations for assessment of biofilm production by staphylococci. APMIS 115:891–899. https://doi.org/10.1111/j.1600-0463.2007.apm_630.x
Stolz A (2009) Molecular characteristics of xenobiotic-degrading sphingomonads. Appl Microbiol Biotechnol 81:793–811. https://doi.org/10.1007/s00253-008-1752-3
Stolz A (2014) Degradative plasmids from sphingomonads. FEMS Microbiol Lett 350:9–19. https://doi.org/10.1111/1574-6968.12283
Takeuchi M, Hamana K, Hiraishi A (2001) Proposal of the genus Sphingomonas sensu stricto and three new genera, Sphingobium, Novosphingobium and Sphingopyxis, on the basis of phylogenetic and chemotaxonomic analyses. Int J Syst Evol Microbiol 51:1405–1417. https://doi.org/10.1099/00207713-51-4-1405
Tamura K, Stecher G, Peterson D, Filipski A, Kumar S (2013) MEGA6: Molecular Evolutionary Genetics Analysis Version 6.0. Mol Biol Evol 30:2725–2729. https://doi.org/10.1093/molbev/mst197
Tanner K, Mancuso CP, Peretó J, Khalil AS, Vilanova C, Pascual J (2020)Sphingomonas solaris sp. nov., isolated from a solar panel in Boston, Massachusetts. Int J Syst Evol Microbiol 70:1814–1821. https://doi.org/10.1099/ijsem.0.003977
Verma H, Rani P, Kumar Singh A, Kumar R, Dwivedi V, Negi V, Lal R (2015) Sphingopyxis flava sp. nov., isolated from a hexachlorocyclohexane (HCH)-contaminated soil. Int J Syst Evol Microbiol 65:3720–3726. https://doi.org/10.1099/ijsem.0.000482
Weisburg WG, Barns SM, Pelletier DA, Lane DJ (1991) 16S ribosomal DNA amplification for phylogenetic study. J Bacteriol 173:697–703. https://doi.org/10.1128/jb.173.2.697-703.1991
Wilson K (2001) Preparation of Genomic DNA from Bacteria. Current Protocols in Molecular Biology 56:2.4.1–2.4.5. https://doi.org/10.1002/0471142727.mb0204s56
Wowk B, Fahy GM (2002) Inhibition of bacterial ice nucleation by polyglycerol polymers. Cryobiology 44:14–23. https://doi.org/10.1016/S0011-2240(02)00008-1
Yabuuchi E, Yano I, Oyaizu H, Hashimoto Y, Ezaki T, Yamamoto H (1990) Proposals of Sphingomonas paucimobilis gen. nov. and comb. nov., Sphingomonas parapaucimobilis sp. nov., Sphingomonas yanoikuyae sp. nov., Sphingomonas adhaesiva sp. nov., Sphingomonas capsulata comb. nov., and two genospecies of the genus Sphingomonas. Microbiol Immunol 34:99–119. https://doi.org/10.1111/j.1348-0421.1990.Tb00996.x
Yang S, Zhang X, Cao Z, Zhao K, Wang S, Chen M, Hu X (2014)Growth-promotingSphingomonas paucimobilis ZJSH1 associated with Dendrobium officinale through phytohormone production and nitrogen fixation. Microb Biotechnol 7:611–620. https://doi.org/10.1111/1751-7915.12148
Yu FB, Shan SD, Luo LP, Guan LB, Qin H (2013) Isolation and characterization of a Sphingomonas sp. strain F-7 degrading fenvalerate and its use in bioremediation of contaminated soil. J Environ Sci Health, Part B 48:198–207. https://doi.org/10.1080/03601234.2013.730299
Acknowledgments
This work was partly supported by Research and Technology Office, Sari Agricultural Sciences and Natural Resources University, Sari, Iran.
Availability of data and material
The datasets analyzed during the current study are available in the [NCBI, GenBank] repository, [http://www.ncbi.nlm.nih.gov].
Code availability
Software application or custom code is not applicable.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study’s conception and design. Conceiving and supervising the work and editing the final draft of the manuscript performed by [Hamideh Mortezania and Heshmatolah Rahimian]. Material preparation, data collection and analysis were performed by [Hamideh mortezania], [Valiollah Babaeizad], [Ali Barzegar] and [Ali Dehestani]. The first draft of the manuscript was written by [Hamideh mortezania] and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
We declare that the work is original, has not been published, or being submitted for publication elsewhere.
Conflict of interest
The authors have no conflicts of interest to declare relevant to the content of this article.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
ESM 1
(DOCX 3399 kb)
Rights and permissions
About this article
Cite this article
Mortezania, H., Rahimian, H., Babaeizad, V. et al. Preliminary characterization and phylogeny of sphingomonads occurring on Stone and Pome Fruit Trees in Northern Iran. Biologia 76, 3811–3825 (2021). https://doi.org/10.1007/s11756-021-00895-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11756-021-00895-7