Abstract
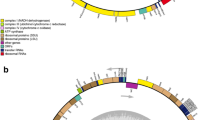
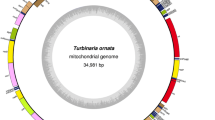
Recent development of DNA sequencing technologies has enabled successful construction of thousands of mitochondrial genomes. Nevertheless, only 33 mitochondrial genomes have been reported for species in Bacillariophyta, which includes harmful algal bloom (HAB) species. In this study, we determined the complete mitogenome of the Bacillariophyta HAB species Odontella regia. For comparison, we also constructed the mitogenome of Lithodesmium undulatum, another Bacillariophyta species. Both strains were isolated in the Jiaozhou Bay, an epitome of China’s coastal ecosystem and an ideal site for HAB research. These two mitogenomes were characterized as 37,617 bp and 37,057 bp circular-mapping molecules with AT content of 73.4% and 75.3%, respectively. The phylogenetic tree based on 32 protein-coding genes of the mitogenome encoded in 35 Bacillariophyta species revealed that the class Mediophyceae consisted of multi-phyletic clades. While O. regia formed an independent clade, L. undulatum was closely related to Thalassiosira pseudonana and Skeletonema marinoi. Synteny comparison of O. regia and L. undulatum mitogenomes and mitogenomes of three closely related species displayed substantial differences among lineages in Mediophyceae by a series of gene translocations and/or inversion with many conserved gene blocks such as rpl2-rps19-rps3-rpl16-atp9. These analyses suggested a complex evolutionary relationship in Mediophyceae in which the mitogenome of O. regia was the least conservative compared with the mitogenomes of four other Bacillariophyta species. Further studies are needed to clarify detailed phylogenetic relationships in Bacillariophyta.





Similar content being viewed by others
References
Adams K, Palme D (2003) Evolution of mitochondrial gene content: gene loss and transfer to the nucleus. Mol Phylogenet Evol 29:380–395
An SM, Noh JH, Choi DH, Lee JH, Yang EC (2016) Repeat region absent in mitochondrial genome of tube-dwelling diatom Berkeleya fennica (Naviculales, Bacillariophyceae). Mitochondrial DNA A 27:2137–2138
An SM, Kim SY, Noh JH, Yang EC (2017) Complete mitochondrial genome of Skeletonema marinoi (Mediophyceae, Bacillariophyta), a clonal chain forming diatom in the west coast of Korea. Mitochondrial DNA A 28:19–20
Anisimova M, Gil M, Dufayard J-F, Dessimoz C, Gascuel O (2011) Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Syst Biol 60:685–699
Armbrust EV, Berges JA, Bowler C, Green BR, Martinez D, Putnam NH, Zhou SG, Allen AE, Apt KE, Bechner M, Brzezinski MA, Chaal BK, Chiovitti A, Davis AK, Demarest MS, Detter JC, Glavina T, Goodstein D, Hadi MZ, Hellsten U, Hildebrand M, Jenkins BD, Jurka J, Kapitonov VV, Kroger N, Lau WWY, Lane TW, Larimer FW, Lippmeier JC, Lucas S, Medina M, Montsant A, Obornik M, Parker MS, Palenik B, Pazour GJ, Richardson PM, Rynearson TA, Saito MA, Schwartz DC, Thamatrakoln K, Valentin K, Vardi A, Wilkerson FP, Rokhsar DS (2004) The genome of the diatom Thalassiosira pseudonana: ecology, evolution, and metabolism. Science 306:79–86
Ashworth MP, Nakov T, Theriot EC (2013) Revisiting Ross and Sims (1971): toward a molecular phylogeny of the Biddulphiaceae and Eupodiscaceae (Bacillariophyceae). J Phycol 49:1207–1222
Aunins AW, Hamilton D, King TL (2018) The complete mitochondrial genome of the stalk-forming diatom Didymosphenia geminata. Mitochondrial DNA B 3:676–677
Badylak S (2004) Spatial and temporal patterns of phytoplankton composition in subtropical coastal lagoon, the Indian River Lagoon, Florida, USA. J Plankton Res 26:1229–1247
Balmer-Hanchey EL, Jaykus L-A, Jaykus L-A, McClellan-Green P (2003) Marine biotoxins of algal origin and seafood safety. J Aquat Food Prod Technol 12:29–53
Bankevich A, Nurk S, Antipov D, Gurevich AA, Dvorkin M, Kulikov AS, Lesin VM, Nikolenko SI, Son P, Prjibelski AD, Pyshkin AV, Sirotkin AV, Vyahhi N, Tesler G, Alekseyev MA, Pevzner PA (2012) SPAdes: a new genome assembly algorithm and its applications to single-cell sequencing. J Comput Biol 19:455–477
Bolger AM, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for Illumina sequence data. Bioinformatics 30:2114–2120
Burger G, Nedelcu AM (2012) Mitochondrial genomes of algae. In: Bock R, Knoop V (eds) Genomics of chloroplasts and mitochondria. Springer, London, pp 127–157
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973
Carstensen J, Klais R, Cloern JE (2015) Phytoplankton blooms in estuarine and coastal waters: seasonal patterns and key species. Estuar Coast Shelf Sci 162:98–109
Chen S-C, Wei D-D, Shao R, Shi J-X, Dou W, Wang J-J (2014) Evolution of multipartite mitochondrial genomes in the booklice of the genus Liposcelis (Psocoptera). BMC Genomics 15:861
Chen B, Xie E, Gao Y, Ji W, Zhou Q (2015) Toxic effects of red tide caused by Karenia mikimotoi on marine organisms. J Fujian Fish 37:241–250 (In chinese, abstract in english)
Chen L, Huang JR, Dai J, Guo YF, Sun JT, Hong XY (2019) Intraspecific mitochondrial genome comparison identified CYTB as a high-resolution population marker in a new pest Athetis lepigone. Genomics 111:744–752
Crosman KM, Petrou EL, Rudd MB, Tillotson MD (2019) Clam hunger and the changing ocean: characterizing social and ecological risks to the Quinault razor clam fishery using participatory modeling. Ecol Soc 24:16
Crowell RM, Nienow JA, Cahoon AB (2019) The complete chloroplast and mitochondrial genomes of the diatom Nitzschia palea (Bacillariophyceae) demonstrate high sequence similarity to the endosymbiont organelles of the dinotom Durinskia baltica. J Phycol 55:352–364
Cunningham CW (1997) Can three incongruence tests predict when data should be combined? Mol Biol Evol 14:733–740
Darling AE, Mau B, Perna NT (2010) progressiveMauve: multiple genome alignment with gene gain, loss and rearrangement. PLoS One 5:e11147
Dyson K, Huppert DD (2010) Regional economic impacts of razor clam beach closures due to harmful algal blooms (HABs) on the Pacific coast of Washington. Harmful Algae 9:264–271
Felsenstein J (1985) Confidence-limits on phylogenies - an approach using the bootstrap. Evolution 39:783–791
Gastineau R, Kim S-Y, Lemieux C, Turmel M, Witkowski A, Park J-G, Kim B-S, Mann DG, Theriot EC (2019) Complete mitochondrial genome of a rare diatom (Bacillariophyta) Proschkinia and its phylogenetic and taxonomic implications. Mitochondrial DNA B 4:25–26
Giordano M, Beardall J, Raven JA (2005) CO2 concentrating mechanisms in algae: mechanisms, environmental modulation, and evolution. Annu Rev Plant Biol 56:99–131
Goer M-PO-LSSL-d, Olsen WTSJL (2006) Complete mitochondrial genome of the three brown algae (Heterokonta, Phaeophyceae) Dictyota dichotoma, Fucus vesiculosus and Desmarestia viridis. Curr Genet 49:47–58
Guillard RRL, Hargreaves PE (1994) Stichochrysis immobilis is a diatom, not a chrysophyte. Phycologia 32:234–236
Guillory WX, Onyshchenko A, Ruck EC, Parks M, Nakov T, Wickett NJ, Alverson AJ (2018) Recurrent loss, horizontal transfer, and the obscure origins of mitochondrial introns in diatoms (Bacillariophyta). Genome Biol Evol 10:1504–1515
Guo H (2004) Illustrations of planktons responsible for the blooms in Chinese coastal waters. Ocean Press, Beijing (In chinese)
Haimeur A, Ulmann L, Mimouni V, Gueno F, Pineau-Vincent F, Meskini N, Tremblin G (2012) The role of Odontella aurita, a marine diatom rich in EPA, as a dietary supplement in dyslipidemia, platelet function and oxidative stress in high-fat fed rats. Lipids Health Dis 11:147
Hallegraeff GM (1993) A review of harmful algae blooms and their apparent global increase. Phycologia 32:79–99
Han X, Zou J, Zhang Y (2004) Harmful algae bloom species in Jiaozhou Bay and the features of distribution. Mar Sci 28:49–54
Hegde S, Narale DD, Anil AC (2011) Sexual reproduction in Odontella regia (Schultze) Simonsen 1974 (Bacillariophyta). Curr Sci 101:222–225
Hoagland P, Anderson DM, Kaoru Y, White AW (2002) The economic effects of harmful algal blooms in the United States: estimates, assessment issues, and information needs. Estuaries 25:819–837
Ibrahim, Imad A-S (2017) Study of phytoplankton blooms incident in Shatt Al-Arab river and marine coast Iraqi line. Mesop Environ J 3:18–25
Imanian B, Pombert J-F, Dorrell RG, Burki F, Keeling PJ (2012) Tertiary endosymbiosis in two dinotoms has generated little change in the mitochondrial genomes of their dinoflagellate hosts and diatom endosymbionts. PLoS One 7:e43763
Jin D, Thunberg E, Hoagland P (2008) Economic impact of the 2005 red tide event on commercial shellfish fisheries in New England. Ocean Coast Manag 51:420–429
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL (2008) NCBIBLAST: a better Web interface. Nucleic Acids Res 36:W5–W9
Kalyaanamoorthy S, Bui Quang M, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589
Kamikawa R, Azuma T, Ishii K, Matsuno Y, Miyashita H (2018) Diversity of organellar genomes in non-photosynthetic diatoms. Protist 169:351–361
Karp-Boss L, Gueta R, Rousso I (2014) Judging diatoms by their cover: variability in local elasticity of Lithodesmium undulatum undergoing cell division. PLoS One 9:e109089
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular Evolutionary Genetics Analysis Version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Li H, Durbin R (2009) Fast and accurate short read alignment with Burrows-Wheeler transform. Bioinformatics 25:1754–1760
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth G, Abecasis G, Durbin R, Genome Project Data P (2009) The Sequence Alignment/Map format and SAMtools. Bioinformatics 25:2078–2079
Liu F, Pang S, Li X, Li J (2014) Complete mitochondrial genome of the brown alga Sargassum horneri (Sargassaceae, Phaeophyceae): genome organization and phylogenetic analyses. J Appl Phycol 27:469–478
Liu F, Melton JT III, Bi Y (2017) Mitochondrial genomes of the green macroalga Ulva pertusa (Ulvophyceae, Chlorophyta): novel insights into the evolution of mitogenomes in the Ulvophyceae. J Phycol 53:1010–1019
Liu F, Liu S, Huang T, Chen N (2019) Construction and comparative analysis of mitochondrial genome in the brown tide forming alga Aureococcus anophagefferens (Pelagophyceae, Ochrophyta). J Appl Phycol 32:441–450
Mann DG (1999) The species concept in diatoms. Phycologia 38:437–495
Mann DG, Droop SJM (1996) Biodiversity, biogeography and conservation of diatoms. Hydrobiologia 336:19–32
McPartlin DA, Loftus JH, Crawley AS, Silke J, Murphy CS, O'Kennedy RJ (2017) Biosensors for the monitoring of harmful algal blooms. Curr Opin Biotechnol 45:164–169
Mimouni V, Ulmann L, Pasquet V, Mathieu M, Picot L, Bougaran G, Cadoret JP, Morant-Manceau A, Schoefs B (2012) The potential of microalgae for the production of bioactive molecules of pharmaceutical interest. Curr Pharm Biotechnol 13:2733–2750
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195
Moore SK, Dreyer SJ, Ekstrom JA, Moore K, Norman K, Klinger T, Allison EH, Jardine SL (2020) Harmful algal blooms and coastal communities: socioeconomic impacts and actions taken to cope with the 2015 U.S. West Coast domoic acid event. Harmful Algae 96:101799
Oudot-Le Secq MP, Green BR (2011) Complex repeat structures and novel features in the mitochondrial genomes of the diatoms Phaeodactylum tricornutum and Thalassiosira pseudonana. Gene 476:20–26
Oudot-Le Secq MP, Fontaine JM, Rousvoal S, Kloareg B, Loiseaux-De Goer S (2001) The complete sequence of a brown algal mitochondrial genome, the ectocarpale Pylaiella littoralis (L.) Kjellm. J Mol Evol 53:80–88
Palmer JD, Adams KL, Cho YR, Parkinson CL, Qiu YL, Song KM (2000) Dynamic evolution of plant mitochondrial genomes: mobile genes and introns and highly variable mutation rates. Proc Natl Acad Sci U S A 97:6960–6966
Pogoda CS, Keepers KG, Hamsher SE, Stepanek JG, Kane NC, Kociolek JP (2019) Comparative analysis of the mitochondrial genomes of six newly sequenced diatoms reveals group II introns in the barcoding region of cox1. Mitochondrial DNA A 30:43–51
Pollock DD, Zwickl DJ, McGuire JA, Hillis DM (2002) Increased taxon sampling is advantageous for phylogenetic inference. Syst Biol 51:664–671
Prasetiya FS, Gastineau R, Poulin M, Lemieux C, Turmel M, Syakti AD, Hardivillier Y, Widowati I, Risjani Y, Iskandar I, Subroto T, Falaise C, Arsad S, Safitri I, Mouget J-L, Leignel V (2019) Haslea nusantara (Bacillariophyceae), a new blue diatom from the Java Sea, Indonesia: morphology, biometry and molecular characterization. Plant Ecol Evol 152:188–202
Qi Y, Zou J, Liang S (2004) Red tide along the coast of China. Science Press, Beijing (In chinese)
Ravin NV, Galachyants YP, Mardanov AV, Beletsky AV, Petrova DP, Sherbakova TA, Zakharova YR, Likhoshway YV, Skryabin KG, Grachev MA (2010) Complete sequence of the mitochondrial genome of a diatom alga Synedra acus and comparative analysis of diatom mitochondrial genomes. Curr Genet 56:215–223
Saitou N, Nei M (1987) The neighbor-joining method-a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Sevcikova T, Klimes V, Zbrankova V, Strnad H, Hroudova M, Vlcek C, Elias M (2016) A comparative analysis of mitochondrial genomes in eustigmatophyte algae. Genome Biol Evol 8:705–722
Shibl AA, Isaac A, Ochsenkuhn MA, Cardenas A, Fei C, Behringer G, Arnoux M, Drou N, Santos MP, Gunsalus KC, Voolstra CR, Amin SA (2020) Diatom modulation of select bacteria through use of two unique secondary metabolites. Proc Natl Acad Sci U S A 117:27445–27455
Smith DR (2016) The past, present and future of mitochondrial genomics: have we sequenced enough mtDNAs? Brief Funct Genomics 15:47–54
Smith SA, Dunn CW (2008) Phyutility: a phyloinformatics tool for trees, alignments and molecular data. Bioinformatics 24:715–716
Song H, Liu F, Li Z, Xu Q, Chen Y, Yu Z, Chen N (2020) Development of a high-resolution molecular marker for tracking Phaeocystis globosa genetic diversity through comparative analysis of chloroplast genomes. Harmful Algae 99:101911
Starkenburg SR, Kwon KJ, Jha RK, McKay C, Jacobs M, Chertkov O, Twary S, Rocap G, Cattolico RA (2014) A pangenomic analysis of the Nannochloropsis organellar genomes reveals novel genetic variations in key metabolic genes. BMC Genomics 15:212
Swofford DL (2002) PAUP*: Phylogenetic Analysis Using Parsimony. Version 40b10 Sinauer Associates, Sunderland
Tamura K, Nei M, Kumar S (2004) Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc Natl Acad Sci U S A 101:11030–11035
Tang X, Bi G (2016) Complete mitochondrial genome of Fistulifera solaris (Bacillariophycidae). Mitochondrial DNA A 27:4405–4406
Theriot EC, Ashworth M, Ruck E, Nakov T, Jansen RK (2010) A preliminary multigene phylogeny of the diatoms (Bacillariophyta): challenges for future research. Plant Ecol Evol 143:278–296
Thorvaldsdottir H, Robinson JT, Mesirov JP (2013) Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief Bioinform 14:178–192
Trifinopoulos J, Lam-Tung N, von Haeseler A, Minh BQ (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44:W232–W235
Villain A, Kojadinovic M, Puppo C, Prioretti L, Hubert P, Zhang Y, Gregori G, Roulet A, Roques C, Claverie J-M, Gontero B, Blanc G (2017) Complete mitochondrial genome sequence of the freshwater diatom Asterionella formosa. Mitochondrial DNA B 2:97–98
Ward BL, Anderson RS, Bendich AJ (1981) The mitochondrial genome is large and variable in a family of plants (Cucurbitaceae). Cell 25:793–803
Wei L, Xin Y, Wang D, Jing X, Zhou Q, Su X, Jia J, Ning K, Chen F, Hu Q, Xu J (2013) Nannochloropsis plastid and mitochondrial phylogenomes reveal organelle diversification mechanism and intragenus phylotyping strategy in microalgae. BMC Genomics 14:534
Williams DM (2007) Diatom phylogeny: fossils, molecules and the extinction of evidence. C R Palevol 6:505–514
Wu Y, Sun S, Zhang Y (2005) Long-term change of environment and it’s influence on phytoplankton community structure in Jiaozhou Bay. Oceanol Limnol Sin 36:487–498 (In chinese, abstract in english)
Xia S, Wang K, Wan LL, Li AF, Hu Q, Zhang CW (2013) Production, characterization, and antioxidant activity of fucoxanthin from the marine diatom Odontella aurita. Mar Drugs 11:2667–2681
Yang S, Dong S (2006) A map of common planktonic diatoms in the waters of China. Ocean University of China Press, Qingdao (In chinese)
Yu Z, Chen N (2019) Emerging trends in red tide and major research progresses. Oceanol Limnol Sin 50:474–486 (In chinese, abstract in english)
Yuan X-L, Cao M, Bi G-Q (2016) The complete mitochondrial genome of Pseudo-nitzschia multiseries (Bacillariophyta). Mitochondrial DNA A 27:2777–2778
Acknowledgments
We are thankful to all staffs of the marine ecological environment genomics research group in the Institute of Oceanology, Chinese Academy of Sciences. We also want to express our gratitude to the two anonymous reviewers for their critical comments and suggestions.
Funding
This research was supported by the Marine S & T Fund of Shandong Province for Pilot National Laboratory for Marine Science and Technology (Qingdao) (No. 2018SDKJ0504); the Strategic Priority Research Program of Chinese Academy of Sciences (Grant No. XDB42000000) (to Nansheng Chen); the Chinese Academy of Sciences Pioneer Hundred Talents Program (to Nansheng Chen); the Taishan Scholar Project Special Fund (to Nansheng Chen); the Qingdao Innovation and Creation Plan (Talent Development Program-5th Annual Pioneer and Innovator Leadership Award to Nansheng Chen, 19-3-2-16-zhc); the Key Research Program of Frontier Sciences, Chinese Academy of Sciences (No. QYZDB-SSW-DQC023) (to Feng Liu); and the Major Scientific and Technological Innovation Project of Shandong Province (No. 2019JZZY020706) (to Feng Liu).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
About this article
Cite this article
Wang, Y., Chen, Y., Wang, J. et al. Mitochondrial genome of the harmful algal bloom species Odontella regia (Mediophyceae, Bacillariophyta). J Appl Phycol 33, 855–868 (2021). https://doi.org/10.1007/s10811-020-02364-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10811-020-02364-1