Abstract
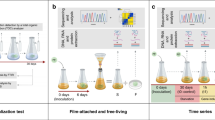
Biodegradability standards measure ultimate biodegradation of polymers by exposing the material under test to a natural microbial inoculum. Available tests developed by the International Organization for Standardization (ISO) use inoculums sampled from different environments e.g. soil, marine sediments, seawater. Understanding whether each inoculum is to be considered as microbially unique or not can be relevant for the interpretation of tests results. In this review, we address this question by consideration of the following: (i) the chemical nature of biodegradable plastics (virtually all biodegradable plastics are polyesters) (ii) the diffusion of ester bonds in nature both in simple molecules and in polymers (ubiquitous); (iii) the diffusion of decomposers capable of producing enzymes, called esterases, which accelerate the hydrolysis of esters, including polyesters (ubiquitous); (iv) the evidence showing that synthetic polyesters can be depolymerized by esterases (large and growing); (v) the evidence showing that these esterases are ubiquitous (growing and confirmed by bioinformatics studies). By combining the relevant available facts it can be concluded that if a certain polyester shows ultimate biodegradation when exposed to a natural inoculum, it can be considered biodegradable and need not be retested using other inoculums. Obviously, if the polymer does not show ultimate biodegradation it must be considered recalcitrant, until proven otherwise.




Similar content being viewed by others
References
Aguiar-Pulido V, Huang W, Suarez-Ulloa V, Cickovski T, Mathee K, Narasimhan G (2016) Metagenomics, metatranscriptomics, and metabolomics approaches for microbiome analysis. Evol Bioinform 12(Suppl 1):5–16. https://doi.org/10.4137/EBO.S36436
Alisch M, Feuerhack A, Müller H, Mensak B, Andreaus J, Zimmermann W (2004) Biocatalytic modification of polyethylene terephthalate fibres by esterases from actinomycete isolates. Biocatal Biotransform 22:347–351. https://doi.org/10.1080/10242420400025877
Alisch-Mark M, Herrmann A, Zimmermann W (2006) Increase of the hydrophilicity of polyethylene terephthalate fibres by hydrolases from Thermomonospora fusca and Fusarium solani f. sp. pisi. Biotechnol Lett 28:681–685. https://doi.org/10.1007/s10529-006-9041-7
Arnosti C, Bell C, Moorhead DL, Sinsabaugh RL, Steen AD, Stromberger M, Wallenstein M, Weintraub MN (2014) Extracellular enzymes in terrestrial, freshwater, and marine environments: perspectives on system variability and common research needs. Biogeochemistry 117:5–21. https://doi.org/10.1007/s10533-013-9906-5
Arya GC, Cohen H (2022) The multifaceted roles of fungal cutinases during infection. J Fungi 8(2):199. https://doi.org/10.3390/jof8020199
Atlas RM (1988) Biodegradation of hydrocarbons in the environment. In: Omenn GS (ed) Environmental biotechnology. Basic life sciences. Springer, Boston, pp 211–222
Bååth JA, Borch K, Jensen K, Brask J, Westh P (2021) Comparative biochemistry of four polyester (PET) hydrolases. ChemBioChem 22:1627–1637. https://doi.org/10.1002/cbic.202000793
Bairoch A (2000) The ENZYME database in 2000. Nucleic Acids Res 28(1):304–305. https://doi.org/10.1093/nar/28.1.304
Bakli M, Bouras N, Paşcalău R, Șmuleac L (2021) In silico characterization of a cutinase from Pseudomonas fluorescens. Res J Agric Sci 53(3):11–20
Balakirev ES, Ayala FJ (2003) Pseudogenes: are they “junk” or functional DNA? Annu Rev Genet 37:123–151. https://doi.org/10.1146/annurev.genet.37.040103.103949
Barrett T, Clark K, Gevorgyan R, Gorelenkov V, Gribov E, Karsch-Mizrachi I, Kimelman M, Pruitt KD, Resenchuk S, Tatusova T, Yaschenko E, Ostell J (2012) BioProject and BioSample databases at NCBI: facilitating capture and organization of metadata. Nucleic Acids Res 40(D1):D57–D63. https://doi.org/10.1093/nar/gkr1163
Barzkar N, Sohail M, Jahromi ST, Gozari M, Poormozaffar S, Nahavandi R, Hafezieh M (2021) Marine bacterial esterases: emerging biocatalysts for industrial applications. Appl Biochem Biotechnol 193(4):1187–1214. https://doi.org/10.1007/s12010-020-03483-8
Bastioli C, Bettarini F (2020) 6. General characteristics, processability, industrial applications and market evolution of biodegradable polymers. In: Bastioli C (ed) Handbook of biodegradable polymers. De Gruyter, Berlin, pp 147–182
Bertrand JC, Bonin P, Caumette P, Gattuso J-P, Gregori G, Guyoneaud R, Le Roux X, Matheron R, Poly F (2015) Biogeochemical cycles. In: Bertrand JC, Caumette P, Lebaron P, Matheron R, Normand P, Sime-Ngando T (eds) Environmental microbiology: fundamentals and applications. Springer, Dordrecht
Bher A, Mayekar PC, Auras RA, Schvezov CE (2022) Biodegradation of biodegradable polymers in mesophilic aerobic environments. Int J Mol Sci 23:12165. https://doi.org/10.3390/ijms232012165
Biundo A, Hromic A, Pavkov-Keller T, Gruber K, Quartinello F, Haernvall K, Perz V, Arrell MS, Zinn M, Ribitsch D, Guebitz GM (2016) Characterization of a poly(butylene adipate-co-terephthalate)-hydrolyzing lipase from Pelosinus fermentans. Appl Microbiol Biotechnol 100:1753–1764. https://doi.org/10.1007/s00253-015-7031-1
Boivin J, Costerton JW (1991) Biodeterioration of materials. In: Rossmoore HW (ed) Biodeterioration and biodegradation 8. Elsevier, Amsterdam, pp 53–62
Bollinger A, Thies S, Knieps-Grünhagen E, Gertzen C, Kobus S, Höppner A, Ferrer M, Gohlke H, Smits S, Jaeger KE (2020) A novel polyester hydrolase from the marine bacterium Pseudomonas aestusnigri -structural and functional insights. Front Microbiol 11:114. https://doi.org/10.3389/fmicb.2020.00114
Bosco F, Mollea C (2021) Biodegradation of natural rubber: microcosm study. Water Air Soil Pollut 232:227. https://doi.org/10.1007/s11270-021-05171-7
Boyadzhieva I, Atanasova N, Paunova-Krasteva T, Kambourova M (2022) Ɛ-Polycaprolactone degradation by a thermostable lipase isolated from Brevibacillus thermoruber strain 7, 29 November 2022, PREPRINT (Version 1) available at Research Square. https://doi.org/10.21203/rs.3.rs-2304161/v1
Brueckner T, Eberl A, Heumann S, Rabe M, Guebitz GM (2008) Enzymatic and chemical hydrolysis of poly(ethylene terephthalate) fabrics. J Polym Sci A Polym Chem 46:6435–6443. https://doi.org/10.1002/pola.22952
Buchholz PCF, Feuerriegel G, Zhang H, Perez-Garcia P, Nover LL, Chow J, Streit WR, Pleiss J (2022) Plastics degradation by hydrolytic enzymes: the plastics-active enzymes database-PAZy. Proteins 90(7):1443–1456. https://doi.org/10.1002/prot.26325
Budkum J, Thammasittirong SN, Thammasittirong A (2022) High poly ε-caprolactone biodegradation activity by a new Acinetobacter seifertii isolate. Folia Microbiol (praha) 67:659–669. https://doi.org/10.1007/s12223-022-00964-7
Carniel A, Valoni E, Nicomedes J, da Conceição GA, Machado de Castro A (2017) Lipase from Candida antarctica (CALB) and cutinase from Humicola insolens act synergistically for PET hydrolysis to terephthalic acid. Process Biochem 59:84–90. https://doi.org/10.1016/j.procbio.2016.07.023
Carr CM, de Oliveira BFR, Jackson SA, Laport MS, Clarke DJ, Dobson ADW (2022) Identification of BgP, a cutinase-like polyesterase from a deep-sea sponge-derived Actinobacterium. Front Microbiol 13:888343. https://doi.org/10.3389/fmicb.2022.888343
CEN/TR 17910:2022 Biodegradable plastics—Status of standardization and new prospects. https://standards.cencenelec.eu/dyn/www/f?p=205:110:0::::FSP_PROJECT:75718&cs=18794D961CDFBF10B7457BB33D3CA58F9. accessed 05 Mar 2023
Charnock C (2021) Norwegian Soils and Waters Contain Mesophilic. Plastic-Degrading Bacteria Microorganisms 9(1):94. https://doi.org/10.3390/microorganisms9010094
Chen S, Tong X, Woodard RW, Du G, Wu J, Chen J (2008) Identification and characterization of bacterial cutinase. J Biol Chem 283(38):25854–25862. https://doi.org/10.1074/jbc.M800848200
Chen S, Su L, Chen J, Wu J (2013) Cutinase: characteristics, preparation, and application. Biotechnol Adv 31(8):1754–1767. https://doi.org/10.1016/j.biotechadv.2013.09.005
Chinaglia S, Tosin M, Degli-Innocenti F (2018) Biodegradation rate of biodegradable plastics at molecular level. Pol Deg Stab 147:237–244. https://doi.org/10.1016/j.polymdegradstab.2017.12.011
Chothia C, Lesk AM (1986) The relation between the divergence of sequence and structure in proteins. EMBO J 5(4):823–826. https://doi.org/10.1002/j.1460-2075.1986.tb04288.x
Colwell RR, Walker JD, Cooney JJ (1977) Ecological aspects of microbial degradation of petroleum in the marine environment. CRC Crit Rev Microbiol 5(4):423–445. https://doi.org/10.3109/10408417709102813
Danso D, Schmeisser C, Chow J, Zimmermann W, Wei R, Leggewie D, Li X, Hazen T, Streit WR (2018) New insights into the function and global distribution of polyethylene terephthalate (PET)-degrading bacteria and enzymes in marine and terrestrial metagenomes. Appl Environ Microbiol 84(8):2773–2817. https://doi.org/10.1128/AEM.02773-17
Danso D, Chow J, Streit WR (2019) Plastics: environmental and biotechnological perspectives on microbial degradation. Appl Environ Microbiol 85(19):e01095-e1119. https://doi.org/10.1128/AEM.01095-19
Degli-Innocenti F, Breton T (2020) Intrinsic biodegradability of plastics and ecological risk in the case of leakage. ACS Sustainable Chem Eng 8(25):9239–9249. https://doi.org/10.1021/acssuschemeng.0c01230
Dimarogona M, Nikolaivits E, Kanelli M, Christakopoulos P, Sandgren M (2015) Structural and functional studies of a Fusarium oxysporum cutinase with polyethylene terephthalate modification potential. Biochim Biophys Acta 11:2308–2317. https://doi.org/10.1016/j.bbagen.2015.08.009
Domergue F, Miklaszewska M (2022) The production of wax esters in transgenic plants: towards a sustainable source of bio-lubricants. J Exp Bot 73(9):2817–2834. https://doi.org/10.1093/jxb/erac046
Doolittle RF (1981) Similar amino acid sequences: chance or common ancestry? Science 214(4517):149–159. https://doi.org/10.1126/science.7280687
EN 17615:2022 Plastics. Environmental aspects. Vocabulary. https://standards.cencenelec.eu/dyn/www/f?p=205:110:0::::FSP_PROJECT:70390&cs=1EBDE354922407F3E9FCE8F7EED378607 accessed 12 Nov 2022
European Bioplastics, Bioplastics market development update report (2021), https://docs.european-bioplastics.org/publications/market_data/Report_Bioplastics_Market_Data_2021_short_version.pdf accessed 20 Oct 2022
Farzi A, Dehnad A, Fotouhi AF (2019) Biodegradation of polyethylene terephthalate waste using Streptomyces species and kinetic modeling of the process. Biocatal Agric Biotechnol 17:25–31. https://doi.org/10.1016/j.bcab.2018.11.002
Fischer-Colbrie G, Heumann S, Liebminger S, Almansa EM, Cavaco-Paulo A, Guebitz GM (2004) New enzymes with potential for PET surface modification. Biocatalysis 22(5–6):341–346. https://doi.org/10.1080/10242420400024565
Fried JR (2014) Polymer science and technology. Prentice Hall, Upper Saddle River
Gambarini V, Pantos O, Kingsbury JM, Weaver L, Handley KM, Lear G (2022) PlasticDB: a database of microorganisms and proteins linked to plastic biodegradation. Database 2022:baac008. https://doi.org/10.1093/database/baac008
Gan Z, Zhang H (2019) PMBD: a comprehensive plastics microbial biodegradation database. Database 2019:baz119. https://doi.org/10.1093/database/baz119
Gao R, Sun C (2021) A marine bacterial community capable of degrading poly(ethylene terephthalate) and polyethylene. J Hazard Mater 416:125928. https://doi.org/10.1016/j.jhazmat.2021.125928
Haernvall K, Zitzenbacher S, Wallig K, Yamamoto M, Schick MB, Ribitsch D, Guebitz GM (2017a) Hydrolysis of ionic phthalic acid based polyesters by wastewater microorganisms and their enzymes. Environ Sci Technol 51(8):4596–4605. https://doi.org/10.1021/acs.est.7b00062
Haernvall K, Zitzenbacher S, Yamamoto M, Schick MB, Ribitsch D, Guebitz GM (2017b) A new arylesterase from Pseudomonas pseudoalcaligenes can hydrolyze ionic phthalic polyesters. J Biotechnol 257:70–77. https://doi.org/10.1016/j.jbiotec.2017.01.012
Hahn S, Hennecke D (2022) Fraunhofer ITEM, Fraunhofer IME, CEFIC—Finale Report WP4—Comparison between natural and synthetic polymers—May 2022 http://cefic-lri.org/wp-content/uploads/2022/07/ECO52-WP4-Report-polymer-Final.pdf accessed 20 Oct 2022
Hajighasemi M, Tchigvintsev A, Nocek B, Flick R, Popovic A, Hai T, Khusnutdinova AN, Brown G, Xu X, Cui H, Anstett J, Chernikova TN, Brüls T, Le Paslier D, Yakimov MN, Joachimiak A, Golyshina OV, Savchenko A, Golyshin PN, Edwards EA, Yakunin AF (2018) Screening and characterization of novel polyesterases from environmental metagenomes with high hydrolytic activity against synthetic polyesters. Environ Sci Technol 52(21):12388–12401. https://doi.org/10.1021/acs.est.8b04252
Hefetz A, Fales HM, Batra SWT (1979) Natural polyesters: DuFour’s gland macrocyclic lactones form brood cell laminesters in colletes bees. Science 204(4391):415–417. https://doi.org/10.1126/science.204.4391.415
Hu X, Gao Z, Wang Z, Su T, Yang L, Li P (2016) Enzymatic degradation of poly(butylene succinate) by cutinase cloned from Fusarium solani. Pol Deg Stab 134:211–219. https://doi.org/10.1016/j.polymdegradstab.2016.10.012
ISO 22404 (2019) Plastics—determination of the aerobic biodegradation of non-floating materials exposed to marine sediment—method by analysis of evolved carbon dioxide https://www.iso.org/standard/73123.html accessed 22 Nov 2022
ISO 23977–1:2020 Plastics—Determination of the aerobic biodegradation of plastic materials exposed to seawater—Part 1: Method by analysis of evolved carbon dioxide. https://www.iso.org/standard/77499.html. accessed 18 Jan 2022
ISO 17556:2019 Plastics — Determination of the ultimate aerobic biodegradability of plastic materials in soil by measuring the oxygen demand in a respirometer or the amount of carbon dioxide evolved. https://www.iso.org/standard/74993.html accessed 12 Nov 2022
Jha SK, Jain P, Sharma HP (2015) Xenobiotic degradation by bacterial enzymes. Int J Curr Microbiol App Sci 4(6):48–62
Kanwal A, Zhang M, Sharaf F, Chengtao L (2022) Screening and characterization of novel lipase producing Bacillus species from agricultural soil with high hydrolytic activity against PBAT poly (butylene adipate co terephthalate) co-polyesters. Polym Bull 79:10053–10076. https://doi.org/10.1007/s00289-021-03992-4
Kanwal A, Zhang M, Sharaf F (2023) Characterization of poly (butylene adipate-co-terephthalate) PBAT co-polyesters degrading bacteria from farmland soil of Xinjiang, 10 January 2023, PREPRINT (Version 1) available at Research Square. https://doi.org/10.21203/rs.3.rs-2448998/v1
Kim SH, Lee JW, Kim JS, Lee W, Park MS, Lim YW (2022) Plastic-inhabiting fungi in marine environments and PCL degradation activity. Antonie Van Leeuwenhoek 115(12):1379–1392. https://doi.org/10.1007/s10482-022-01782-0
Kleeberg I, Welzel K, Vandenheuvel J, Müller RJ, Deckwer WD (2005) Characterization of a new extracellular hydrolase from Thermobifida fusca degrading aliphatic-aromatic copolyesters. Biomacromol 6(1):262–270. https://doi.org/10.1021/bm049582t
Kržan A, Hemjinda S, Miertus S, Corti A, Chiellini E (2006) Standardization and certification in the area of environmentally degradable plastics. Polym Degrad Stab 91:2819–2833. https://doi.org/10.1016/j.polymdegradstab.2006.04.034
Li-Beisson Y (2011) Cutin and suberin. Encyclopedia of life sciences (eLS). JWS Ltd, Chichester
Liebminger S, Eberl A, Sousa F, Heumann S, Fischer-Colbrie G, Cavaco-Paulo A, Guebitz GM (2007) Hydrolysis of PET and bis-(benzoyloxyethyl) terephthalate with a new polyesterase from Penicillium citrinum. Biocatal 25(2–4):171–177. https://doi.org/10.1080/10242420701379734
Liu Y, Liu C, Liu H, Zeng Q, Tian X, Long L, Yang J (2022) Catalytic features and thermal adaptation mechanisms of a deep sea bacterial cutinase-type poly (ethylene terephthalate) hydrolase. Front Bioeng Biotechnol 10:865787. https://doi.org/10.3389/fbioe.2022.865787
Martínez A, Maicas S (2021) Cutinases: characteristics and insights in industrial production. Catalysts 11:1194. https://doi.org/10.3390/catal11101194
Meyer-Cifuentes IE, Werner J, Jehmlich N, Will SE, Neumann-Schaal M, Öztürk B (2020) Synergistic biodegradation of aromatic-aliphatic copolyester plastic by a marine microbial consortium. Nat Commun 11:5790. https://doi.org/10.1038/s41467-020-19583-2
Mishra M, Singh SK, Kumar A (2021) Chapter 5-Environmental factors affecting the bioremediation potential of microbes. In: Kumar A, Singh VK, Singh P, Mishra VK (eds) Woodhead publishing series in food science, technology and nutrition, microbe mediated remediation of environmental contaminants. Woodhead Publishing, Sawston, pp 47–58
Müller R (2006) Biological degradation of synthetic polyesters—enzymes as potential catalysts for polyester recycling. Process Biochem 41:2124–2128. https://doi.org/10.1016/j.procbio.2006.05.018
Müller RJ, Schrader H, Profe J, Dresler K, Deckwer WD (2005) Enzymatic degradation of poly(ethylene terephthalate): rapid hydrolyse using a hydrolase from T. fusca. Macromol Rapid Commun 26:1400–1405. https://doi.org/10.1002/marc.200500410
Muroi F, Tachibana Y, Soulenthone P, Yamamoto K, Mizuno T, Sakurai T, Kobayashi Y, Kasuya K (2017) Characterization of a poly(butylene adipate-co-terephthalate) hydrolase from the aerobic mesophilic bacterium Bacillus pumilus. Polym Degrad Stab 137:11–22. https://doi.org/10.1016/j.polymdegradstab.2017.01.006
Nawaz A, Hasan F, Shah AA (2015) Degradation of poly(ε-caprolactone) (PCL) by a newly isolated Brevundimonas sp. strain MRL-AN1 from soil. FEMS Microbiol Lett 362(1):1–7. https://doi.org/10.1093/femsle/fnu004
Nikolaivits E, Kanelli M, Dimarogona M, Topakas E (2018) A middle-aged enzyme still in its prime: recent advances in the field of cutinases. Catalysts 8(12):612. https://doi.org/10.3390/catal8120612
Nimchua T, Punnapayak H, Zimmermann W (2007) Comparison of the hydrolysis of polyethylene terephthalate fibers by a hydrolase from Fusarium oxysporum LCH I and Fusarium solani f. sp. pisi. Biotechnol J 2:361–364. https://doi.org/10.1002/biot.200600095
Nimchua T, Eveleigh DE, Sangwatanaroj U, Punnapayak H (2008) Screening of tropical fungi producing polyethylene terephthalate-hydrolyzing enzyme for fabric modification. J Ind Microbiol Biotechnol 35(8):843–850. https://doi.org/10.1007/s10295-008-0356-3
Oda M, Numoto N, Bekker GJ, Kamiya N, Kawai F (2021) Cutinases from thermophilic bacteria (actinomycetes): from identification to functional and structural characterization. Methods Enzymol 648:159–185. https://doi.org/10.1016/bs.mie.2020.12.031
OECD (2006) Revised introduction to the OECD guidelines for testing of chemicals, section 3, OECD guidelines for the testing of chemicals, section 3. OECD Publishing, Paris. https://doi.org/10.1787/9789264030213-en
Ollis DL, Cheah E, Cygler M, Dijkstra B, Frolow F, Franken SM, Harel M, Remington SJ, Silman I, Schrag J (1992) The alpha/beta hydrolase fold. Protein Eng 5(3):197–211. https://doi.org/10.1093/protein/5.3.197
Penkhrue W, Khanongnuch C, Masaki K, Pathom-Aree W, Punyodom W, Lumyong S (2015) Isolation and screening of biopolymer-degrading microorganisms from northern Thailand. World J Microbiol Biotechnol 31(9):1431–1442. https://doi.org/10.1007/s11274-015-1895-1
Pérez-García P, Danso D, Zhang H, Chow J, Streit WR (2021) Chapter Seven-Exploring the global metagenome for plastic-degrading enzymes. Methods Enzymol 648:137–157. https://doi.org/10.1016/bs.mie.2020.12.022
Perz V, Baumschlager A, Bleymaier K, Zitzenbacher S, Hromic A, Steinkellner G, Pairitsch A, Łyskowski A, Gruber K, Sinkel C, Küper U, Ribitsch D, Guebitz GM (2015) Hydrolysis of synthetic polyesters by Clostridium botulinum esterases. Biotechnol Bioeng 113(5):1024–1034. https://doi.org/10.1002/bit.25874
Perz V, Bleymaier K, Sinkel C, Kueper U, Bonnekessel M, Ribitsch D, Guebitz GM (2016a) Substrate specificities of cutinases on aliphatic-aromatic polyesters and on their model substrates. N Biotechnol 33(2):295–304. https://doi.org/10.1016/j.nbt.2015.11.004
Perz V, Hromic A, Baumschlager A, Steinkellner G, Pavkov-Keller T, Gruber K, Bleymaier K, Zitzenbacher S, Zankel A, Mayrhofer C, Sinkel C, Kueper U, Schlegel K, Ribitsch D, Guebitz GM (2016b) An esterase from anaerobic Clostridium hathewayi can hydrolyze aliphatic–aromatic polyesters. Environ Sci Technol 50:2899–2907. https://doi.org/10.1021/acs.est.5b04346
Peters EN (2005) Plastics: information and properties of polymeric materials. In: Kutz M (ed) Mechanical engineers’ handbook: materials and mechanical design, vol 1, 3rd edn. Wiley, Hoboken, pp 336–375
Ping LF, Chen XY, Yuan XL, Zhang M, Chai YJ, Shan SD (2017) Application and comparison in biosynthesis and biodegradation by Fusarium solani and Aspergillus fumigatus cutinases. Int J Biol Macromol 104(Pt A):1238–1245. https://doi.org/10.1016/j.ijbiomac.2017.06.118
Pischedda A, Tosin M, Degli-Innocenti F (2019) Biodegradation of plastics in soil: the effect of temperature. Pol Deg Stab 170:109017. https://doi.org/10.1016/j.polymdegradstab.2019.109017
Plastics Europe, 2021, Facts (2021) https://plasticseurope.org/knowledge-hub/plastics-the-facts-2021/ accessed on October 20, 2022
Popovic A, Hai T, Tchigvintsev A, Hajighasemi M, Nocek B, Khusnutdinova AN, Brown G, Glinos J, Flick R, Skarina T, Chernikova TN, Yim V, Brüls T, Paslier DL, Yakimov MM, Joachimiak A, Ferrer M, Golyshina OV, Savchenko A, Golyshin PN, Yakunin AF (2017) Activity screening of environmental metagenomic libraries reveals novel carboxylesterase families. Sci Rep 7:44103. https://doi.org/10.1038/srep44103
Pundir S, Martin MJ, O’Donovan C et al (2016) Uniprot tools. Curr Protoc Bioinform 53:115. https://doi.org/10.1002/0471250953.bi0129s53
Rauwerdink A, Kazlauskas RJ (2015) How the same core catalytic machinery catalyzes 17 different reactions: the serine-histidine-aspartate catalytic triad of α/β-hydrolase fold enzymes. ACS Catal 5(10):6153–6176. https://doi.org/10.1021/acscatal.5b01539
Regulation (EC) No 1907/2006 of the European Parliament and of the Council of 18 December 2006 concerning the Registration, Evaluation, Authorisation and Restriction of Chemicals (REACH), establishing a European Chemicals Agency, amending Directive 1999/45/EC and repealing Council Regulation (EEC) No 793/93 and Commission Regulation (EC) No 1488/94 as well as Council Directive 76/769/EEC and Commission Directives 91/155/EEC, 93/67/EEC, 93/105/EC and 2000/21/EC, Official Journal of the European Union, 2006. https://eur-lex.europa.eu/legal-content/EN/TXT/PDF/?uri=CELEX:32006R1907&from=en accessed 05 Mar 2023
Ribitsch D, Heumann S, Trotscha E, Herrero Acero E, Greimel K, Leber R, Birner-Gruenberger R, Deller S, Eiteljoerg I, Remler P, Weber T, Siegert P, Maurer KH, Donelli I, Freddi G, Schwab H, Guebitz GM (2011) Hydrolysis of polyethyleneterephthalate by p-nitrobenzylesterase from Bacillus subtilis. Biotechnol Prog 27:951–960. https://doi.org/10.1002/btpr.610
Rizzarelli P, DegliInnocenti F, Valenti G, Rapisarda M (2020) Biodegradation of green polymer composites: laboratory procedures and standard test methods, materials research foundations. In: Al-Ahmed A, Inamuddin (eds) Advanced applications of bio-degradable green composites. Materials Research Forum LLC, Millersville. pp 1–44
Ronkvist ÅM, Xie W, Lu W, Gross RA (2009) Cutinase-catalyzed hydrolysis of poly(ethylene terephthalate). Macromolecules 42(14):5128–5138. https://doi.org/10.1021/ma9005318
Rosato A, Romano A, Totaro G, Celli A, Fava F, Zanaroli G, Sisti L (2022) Enzymatic degradation of the most common aliphatic bio-polyesters and evaluation of the mechanisms involved: an extended study. Polymers 14:1850. https://doi.org/10.3390/polym14091850
Rosenboom JG, Langer R, Traverso G (2022) Bioplastics for a circular economy. Nat Rev Mater 7:117–137. https://doi.org/10.1038/s41578-021-00407-8
Rost B (1999) Twilight zone of protein sequence alignments. Protein Eng 12(2):85–94. https://doi.org/10.1093/protein/12.2.85
Rueda Rueda H, Prieto-Correaa E (2020) Cutinases obtained from filamentous fungi: comparison of screening methods. DYNA 87(214):183–190
Sander C, Schneider R (1991) Database of homology-derived protein structures and the structural meaning of sequence alignment. Proteins 9(1):56–68. https://doi.org/10.1002/prot.340090107
Santos-Beneit F, Chen LM, Bordel S, Frutos de la Flor R, García-Depraect O, Lebrero R, Rodriguez-Vega S, Muñoz R, Börner RA, Börner T (2023) Screening enzymes that can depolymerize commercial biodegradable polymers: heterologous expression of Fusarium solani cutinase in Escherichia coli. Microorganisms 11(2):328. https://doi.org/10.3390/microorganisms11020328
Satti SM, Shah AA (2020) Polyester-based biodegradable plastics: an approach towards sustainable development. Lett Appl Microbiol 70(6):413–430. https://doi.org/10.1111/lam.13287
Sebastian J, Chandra AK, Kolattukudy PE (1987) Discovery of a cutinase-producing Pseudomonas sp. cohabiting with an apparently nitrogen-fixing Corynebacterium sp. in the phyllosphere. J Bacteriol 169(1):131–136
Shah AA, Kato S, Shintani N, Kamini NR, Nakajima-Kambe T (2014) Microbial degradation of aliphatic and aliphatic-aromatic co-polyesters. Appl Microbiol Biotechnol 98:3437–3447. https://doi.org/10.1007/s00253-014-5558-1
Sharma R, Chisti Y, Banerjee UC (2001) Production, purification, characterization, and applications of lipases. Biotechnol Adv 19(8):627–662. https://doi.org/10.1016/s0734-9750(01)00086-6
Sharma T, Sharma A, Kanwar SS (2017) An overview on esterases: structure, classification, sources and their application. Recent Adv Biotechnol 2:216–229
Sharma A, Sharma T, Sharma T, Sharma S, Kanwar SS (2019) Role of microbial hydrolases in bioremediation. Microbes Environ 16:149–164. https://doi.org/10.1007/978-981-13-9117-0_7
Sharon C, Sharon M (2012) Studies on biodegradation of polyethylene terephthalate: a synthetic polymer. J Microbiol Biotechnol Res 2(2):248–257
Shi K, Jing J, Song L, Su T, Wang Z (2020) Enzymatic hydrolysis of polyester: degradation of poly(ε-caprolactone) by Candida antarctica lipase and Fusarium solani cutinase. Int J Biol Macromol 144:183–189. https://doi.org/10.1016/j.ijbiomac.2019.12.105
Shukla E, Bendre AD, Gaikwad SM (2022) Hydrolases: The Most Diverse Class of Enzymes. In: Haider S, Haider A, Català A (eds) Hydrolases. IntechOpen, London
Silva CM, Carneiro F, O’Neill A, Fonseca LP, Cabral JSM, Guebitz G, Cavaco-Paulo A (2005) Cutinase—a new tool for biomodification of synthetic fibers. J Polym Sci A 43:2448–2450. https://doi.org/10.1002/pola.20684
Sood S, Sharma A, Sharma N, Kanwar SS (2016) Carboxylesterases: sources, characterization and broader applications. Insights Enzyme Res 1:1. https://doi.org/10.21767/2573-4466.100002
Soulenthone P, Tachibana Y, Muroi F, Suzuki M, Ishii N, Ohta Y, Kasuya K (2020) Characterization of a mesophilic actinobacteria that degrades poly(butylene adipate-co-terephthalate). Polym Degrad Stab 181:109335. https://doi.org/10.1016/j.polymdegradstab.2020.109335
Sparkman OD, Penton ZE, Kitson FG (2011) Chapter 15—Esters. In: Sparkman OD, Penton ZE, Kitson FG (eds) Gas chromatography and mass spectrometry, 2nd edn. Academic Press, Boca Raton, pp 285–292
Streit WR, Schmitz RA (2004) Metagenomics-the key to the uncultured microbes. Curr Opin Microbiol 7(5):492–498. https://doi.org/10.1016/j.mib.2004.08.002
Sulaiman S, Yamato S, Kanaya E, Kim JJ, Koga Y, Takano K, Kanaya S (2012) Isolation of a novel cutinase homolog with polyethylene terephthalate-degrading activity from leaf-branch compost by using a metagenomic approach. Appl Environ Microbiol 78(5):1556–1562. https://doi.org/10.1128/AEM.06725-11
Sun X, Kostka JE (2019) Hydrocarbon-degrading microbial communities are site specific, and their activity is limited by synergies in temperature and nutrient availability in surface ocean waters. Appl Environ Microbiol 85(15):e00443-e519. https://doi.org/10.1128/AEM.00443-19
Suyama T, Tokiwa Y, Ouichanpagdee P, Kanagawa T, Kamagata Y (1998) Phylogenetic affiliation of soil bacteria that degrade aliphatic polyesters available commercially as biodegradable plastics. Appl Environ Microbiol 64(12):5008–5011. https://doi.org/10.1128/AEM.64.12.5008-5011.1998
Suzuki M, Tachibana Y, Ji K, Takizawa R, Muroi F, Ki K (2017) Difference in environmental degradability between poly(ethylene succinate) and poly(3-hydroxybutyrate). J Polym Res 24:217. https://doi.org/10.1007/s10965-017-1383-4
Suzuki M, Tachibana Y, Kasuya KI (2021) Biodegradability of poly(3-hydroxyalkanoate) and poly(ε-caprolactone) via biological carbon cycles in marine environments. Polym J 53:47–66. https://doi.org/10.1038/s41428-020-00396-5
Thouand G (2014) Biodegradability assessments of organic substances and polymers. Environ Sci Pollut Res 21:9443–9444. https://doi.org/10.1007/s11356-014-2663-8
TNO- Netherlands Organization for Applied Scientific Research (2005) From PEC_PNEC ratio to quantitative risk level using species sensitivity distributions; methodology applied in the environmental impact factor. https://www.sintef.no/globalassets/project/erms/reports/erms-report-no-10_pec_pnec-to-risk-ssd_tno.pdf accessed 11 Nov 2022
Todea A, Cespugli M, Ebert C, Asaro F, Tomada S, Hollan F, Fortuna S, Gardossi L (2022) Rational guidelines for the two-step scalability of enzymatic polycondensation: experimental and computational optimization of the enzymatic synthesis of poly(glycerolazelate). Chem Sus Chem. https://doi.org/10.1002/cssc.202102657
Tokiwa Y, Calabia BP (2007) Biodegradability and biodegradation of polyesters. J Polym Environ 15:259–267. https://doi.org/10.1007/s10924-007-0066-3
Tokiwa Y, Jarerat A (2003) Microbial degradation of aliphatic polyesters. Macromol Symp 201(1):283–289. https://doi.org/10.1002/masy.200351131
Tokiwa Y, Suzuki T, Takeda K (1986) Hydrolysis of polyesters by Rhizopus arrhizus lipase. Agric Biol Chem 50(5):1323–1325. https://doi.org/10.1080/00021369.1986.10867565
Torena P, Alvarez-Cuenca M, Reza M (2021) Biodegradation of polyethylene terephthalate microplastics by bacterial communities from activated sludge. Can J Chem Eng 99:S69–S82. https://doi.org/10.1002/cjce.24015
Urbanek AK, Rymowicz W, Strzelecki MC, Kociuba W, Franczak L, Mirończuk AM (2017) Isolation and characterization of Arctic microorganisms decomposing bioplastics. AMB Express 7:148. https://doi.org/10.1186/s13568-017-0448-4
Urbanek AK, Mirończuk AM, García-Martín A, Saborido A, de la Mata I (2020) Biochemical properties and biotechnological applications of microbial enzymes involved in the degradation of polyester-type plastics. Biochim Biophys Acta Proteins Proteom 2:140315. https://doi.org/10.1016/j.bbapap.2019.140315
Vivek K, Sandhia GS, Subramaniyan S (2022) Extremophilic lipases for industrial applications: a general review. Biotechnol Adv 60:108002. https://doi.org/10.1016/j.biotechadv.2022.108002
Wallace PW, Haernvall K, Ribitsch D, Zitzenbacher S, Schittmayer M, Steinkellner G, Gruber K, Guebitz GM, Birner-Gruenberger R (2016) PpEst is a novel PBAT degrading polyesterase identified by proteomic screening of Pseudomonas pseudoalcaligenes. Appl Microbiol Biotechnol 101:2291–2303. https://doi.org/10.1007/s00253-016-7992-8
Walter T, Augusta J, Müller RJ, Widdecke H, Klein J (1995) Enzymatic degradation of a model polyester by lipase from Rhizopus delemar. Enzyme Microb Technol 17(3):218–224. https://doi.org/10.1016/0141-0229(94)00007-E
Wei R, Zimmermann W (2017) Microbial enzymes for the recycling of recalcitrant petroleum-based plastics: how far are we? Microb Biotechnol 10(6):1308–1322. https://doi.org/10.1111/1751-7915.12710
Wei S, Zhao Y, Zhou R, Lin J, Su T, Tong H, Wang Z (2022) Biodegradation of polybutylene adipate-co-terephthalate by Priestia megaterium, Pseudomonas mendocina, and Pseudomonas pseudoalcaligenes following incubation in the soil. Chemosphere 307(Pt1):135700. https://doi.org/10.1016/j.chemosphere.2022.135700
Weinberger S, Canadell J, Quartinello F, YeniadB AA, Pellis A, Guebitz GM (2017) Enzymatic degradation of poly(ethylene 2,5-furanoate) powders and amorphous film. Catalysts 7(11):318. https://doi.org/10.3390/catal7110318
Witt U, Müller RJ, Deckwer WD (1995) New biodegradable polyester–copolymers from commodity chemicals with favorable use properties. J Environ Polym Degrad 3:215–223. https://doi.org/10.1007/BF02068676
Witt U, Einig T, Yamamoto M, Kleeberg I, Deckwer WD, Müller RJ (2001) Biodegradation of aliphatic-aromatic copolyesters: evaluation of the final biodegradability and ecotoxicological impact of degradation intermediates. Chemosphere 44(2):289–299. https://doi.org/10.1016/s0045-6535(00)00162-4
Wu P, Li Z, Gao J, Zhao Y, Wang H, Qin H, Gu Q, Wei R, Liu W, Han X (2023) Characterization of a PBAT degradation carboxylesterase from Thermobacillus composti KWC4. Catalysts 13(2):340. https://doi.org/10.3390/catal13020340
Wufuer R, Li W, Wang S, Duo J (2022) Isolation and degradation characteristics of PBAT film degrading bacteria. Int J Environ Res Public Health 19(24):17087. https://doi.org/10.3390/ijerph192417087
Yang Y, Yang J, Jiang L (2016) Comment on “A bacterium that degrades and assimilates poly(ethylene terephthalate).” Science 353(6301):759. https://doi.org/10.1126/science.aaf8305
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016a) Response to Comment on “A bacterium that degrades and assimilates poly(ethylene terephthalate).” Science 353(6301):759. https://doi.org/10.1126/science.aaf8625
Yoshida S, Hiraga K, Takehana T, Taniguchi I, Yamaji H, Maeda Y, Toyohara K, Miyamoto K, Kimura Y, Oda K (2016b) A bacterium that degrades and assimilates poly(ethylene terephthalate). Science 351(6278):1196–1199. https://doi.org/10.1126/science.aad6359
Zee M (2020) Methods for evaluating the biodegradability of environmentally degradable polymers. In: Bastioli C (ed) Handbook of biodegradable polymers. De Gruyter, Boston, pp 1–22
Zuckerkandl E, Pauling L (1965) Evolutionary divergence and convergence in proteins. In: Bryson V, Vogel HJ (eds) Evolving genes and proteins. Academic Press, Boca Raton, pp 97–166
Acknowledgements
Many thanks to Prof. Lucia Gardossi for useful discussions and advice.
Funding
No funding was received for conducting this study.
Author information
Authors and Affiliations
Contributions
FDI: conceptualization, investigation, writing—original draft preparation, writing—review & editing); T.B.: writing—review & editing; S.C.: investigation, writing—original draft preparation, writing—review & editing; E.E.: Writing—review & editing; M.P.: supervision, writing—review & editing; A.Pe.: investigation, writing—original draft preparation; A.Pi.: investigation, conceptualization, visualization, writing—original draft preparation, writing—review & editing; M.T.: Writing—review & editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they work for a company that produces biodegradable plastics.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Degli-Innocenti, F., Breton, T., Chinaglia, S. et al. Microorganisms that produce enzymes active on biodegradable polyesters are ubiquitous. Biodegradation 34, 489–518 (2023). https://doi.org/10.1007/s10532-023-10031-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10532-023-10031-8