Abstract
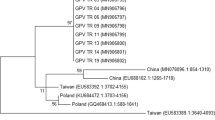
Since its first recognition in the early 1960s, Derzsy’s disease has caused significant economic losses in the goose meat industry through the world. Today, Derzsy’s disease still maintains its importance for small-scale waterfowl farming, despite not having a significant impact on public health. In the present study, we investigated the distribution of goose parvovirus (GPV) and its potential variants from a 2019 outbreak in Turkey. Tissue samples were obtained from infected eggs and goslings that were raised in distinct farming areas of the various provinces. For this purpose, a novel primer set for amplification of a 630-bp region of VP3 was designed to confirm GPV infection by conventional PCR method. A 4709-base nucleotide sequence including the structural, non-structural, and 5' inverted terminal repeat regions was obtained from three samples from the Central Anatolian region. Multiple sequence comparisons and phylogenetic analysis demonstrated that the field strains clustered with European group 2 and contained a series of unique amino acid substitutions that might affect the virulence of the virus. These results confirmed that European-related field strains caused the outbreak in Asia Minor, and this might assist in understanding the circulation of GPV in Asia and Europe.



Similar content being viewed by others
Abbreviations
- BLAST:
-
Basic Local Alignment Search Tool
- dN:
-
Non-synonymous mutation
- dS:
-
Synonymous mutation
- GPV:
-
Goose parvovirus
- ITR:
-
Inverted terminal repeat
- MDPV:
-
Muscovy duck parvovirus
- MUSCLE:
-
Multiple Sequence Comparison by Log-Expectation
- MSA:
-
Multiple sequence alignment
- NS:
-
Non-structural
- PCR:
-
Polymerase chain reaction
- Rep:
-
Replication
- RBE:
-
Ribosome binding element
- SBDS:
-
Short beak and dwarfism syndrome
- USA:
-
United States of America
- VP:
-
Viral protein
References
Woolcock PR (2013) Viral infections of waterfowl. Diseases of poultry. Wiley, New York, pp 417–463
Nagy Z, Derzsy D (1968) A viral disease of goslings. II. Microscopic lesions. Acta Vet Acad Sci Hung 18:3–17
Cotmore SF, Agbandje-McKenna M, Canuti M et al (2019) ICTV virus taxonomy profile: Parvoviridae. J Gen Virol 100:367–368. https://doi.org/10.1099/jgv.0.001212
Bian G, Ma H, Luo M et al (2019) Identification and genomic analysis of two novel duck-origin GPV-related parvovirus in China. BMC Vet Res 15:1–10. https://doi.org/10.1186/s12917-019-1833-9
Parrish CR (2011) Parvoviridae. Fenner’s veterinary virology. Elsevier, Amsterdam, pp 225–235
Tatár-kis T, Mató T, Markos B, Palya V (2004) Phylogenetic analysis of Hungarian goose parvovirus isolates and vaccine strains. Avian Pathol 33:438–444. https://doi.org/10.1080/03079450410001724067
Le Gall-Reculé G, Jestin V (1994) Biochemical and genomic characterization of muscovy duck parvovirus. Arch Virol 139:121–131. https://doi.org/10.1007/BF01309459
Zádori Z, Stefancsik R, Rauch T, Kisary J (1995) Analysis of the complete nucleotide sequences of goose and muscovy duck pervoviruses indicates common ancestral origin with adeno-associated virus 2. Virology 212:562–573. https://doi.org/10.1006/viro.1995.1514
Zádori Z, Erdei J, Nagy J, Kisary J (1994) Characteristics of the genome of goose parvovirus. Avian Pathol 23:359–364. https://doi.org/10.1080/03079459408419004
Schettler CH (1971) Virus hepatitis of geese II. Host range of goose hepatitis virus. Avian Dis 15:809. https://doi.org/10.2307/1588871
Schettler CH (1971) Goose virus hepatitis in the Canada goose and snow goose. J Wildl Dis 7:147–148. https://doi.org/10.7589/0090-3558-7.3.147
Glávits R, Zolnai A, Szabó É et al (2005) Comparative pathological studies on domestic geese (Anser Anser Domestica) and muscovy ducks (Cairina Moschata) experimentally infected with parvovirus strains of goose and muscovy duck origin. Acta Vet Hung 53:73–89. https://doi.org/10.1556/AVet.53.2005.1.8
Stoute ST, Tsai H, Metwally SA et al (2020) Viral infections of waterfowl. Diseases of poultry. Wiley, New York, pp 446–497
Palya V, Zolnai A, Benyeda Z et al (2009) Short beak and dwarfism syndrome of mule duck is caused by a distinct lineage of goose parvovirus. Avian Pathol 38:175–180. https://doi.org/10.1080/03079450902737839
Ning K, Liang T, Wang M et al (2017) Genetic detection and characterization of goose parvovirus: Implications for epidemiology and pathogenicity in Cherry Valley Pekin ducks. Infect Genet Evol 51:101–103. https://doi.org/10.1016/j.meegid.2017.03.024
Ning K, Liang T, Wang M et al (2018) Pathogenicity of a variant goose parvovirus, from short beak and dwarfism syndrome of Pekin ducks, in goose embryos and goslings. Avian Pathol 47:391–399. https://doi.org/10.1080/03079457.2018.1459040
Hlinak A, Müller T, Kramer M et al (1998) Serological survey of viral pathogens in bean and white-fronted geese from Germany. J Wildl Dis 34:479–486. https://doi.org/10.7589/0090-3558-34.3.479
Irvine R, Ceeraz V, Cox B et al (2008) Goose parvovirus in Great Britain. Vet Rec 163:461. https://doi.org/10.1136/vr.163.15.461
Wozniakowski G (2009) Genetic variance of Derzsy’s disease strains isolated in Poland. J Mol Genet Med 03:210–216. https://doi.org/10.4172/1747-0862.1000037
Jansson DS, Feinstein R, Kardi V et al (2007) Epidemiologic investigation of an outbreak of goose parvovirus infection in Sweden. Avian Dis Dig 2:e18–e18. https://doi.org/10.1637/0005-2086(2007)51[609:EIOAOO]2.0.CO;2
Edgar RC (2004) MUSCLE: A multiple sequence alignment method with reduced time and space complexity. BMC Bioinformatics 5:1–19. https://doi.org/10.1186/1471-2105-5-113
Kearse M, Moir R, Wilson A et al (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649. https://doi.org/10.1093/bioinformatics/bts199
Sterky F, Lundeberg J (2000) Sequence analysis of genes and genomes. J Biotechnol 76:1–31. https://doi.org/10.1016/S0168-1656(99)00176-5
Sievers F, Higgins DG (2018) Clustal Omega for making accurate alignments of many protein sequences. Protein Sci 27:135–145. https://doi.org/10.1002/pro.3290
Kumar S, Stecher G, Li M et al (2018) MEGA X: Molecular evolutionary genetics analysis across computing platforms. Mol Biol Evol 35:1547–1549. https://doi.org/10.1093/molbev/msy096
Martin DP, Murrell B, Golden M et al (2015) RDP4: detection and analysis of recombination patterns in virus genomes. Virus Evol 1:1–5. https://doi.org/10.1093/ve/vev003
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Guindon S, Dufayard JF, Lefort V et al (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Kardoğan Ö, Müştak HK, Müştak İB (2021) The first detection and characterization of goose parvovirus (GPV) in Turkey. Trop Anim Health Prod. https://doi.org/10.1007/s11250-020-02463-8
Poonia B, Dunn P, Lu H et al (2006) Isolation and molecular characterization of a new Muscovy duck parvovirus from Muscovy ducks in the USA. Avian Pathol 35:435–441. https://doi.org/10.1080/03079450601009563
Shien J-H, Wang Y-S, Chen C-H et al (2008) Identification of sequence changes in live attenuated goose parvovirus vaccine strains developed in Asia and Europe. Avian Pathol 37:499–505. https://doi.org/10.1080/03079450802356979
Wan C, Chen C, Cheng L et al (2019) Specific detection and differentiation of classic goose parvovirus and novel goose parvovirus by TaqMan real-time PCR assay, coupled with host specificity. BMC Vet Res 15:1–8. https://doi.org/10.1186/s12917-019-2090-7
Jinlong Y, Rui Y, Anchun C et al (2010) A simple and rapid method for detection of Goose Parvovirus in the field by loop-mediated isothermal amplification. Virol J 7:2–8. https://doi.org/10.1186/1743-422X-7-14
Yu X, Wei L, Chen H et al (2018) Development of colloidal gold-based immunochromatographic assay for rapid detection of goose parvovirus. Front Microbiol 9:1–7. https://doi.org/10.3389/fmicb.2018.00953
Chang PC, Shien JH, Wang MS, Shieh HK (2000) Phylogenetic analysis of parvoviruses isolated in Taiwan from ducks and geese. Avian Pathol 29:45–49. https://doi.org/10.1080/03079450094270
Tsai H-J, Tseng C-H, Chang P-C et al (2004) Genetic variation of viral protein 1 genes of field strains of waterfowl parvoviruses and their attenuated derivatives. Avian Dis 48:512–521. https://doi.org/10.1637/7172
Chen S, Liu P, He Y et al (2018) The 164 K, 165 K and 167 K residues in 160YPVVKKPKLTEE171 are required for the nuclear import of goose parvovirus VP1. Virology 519:17–22. https://doi.org/10.1016/j.virol.2018.03.020
Tu M, Liu F, Chen S et al (2015) Role of capsid proteins in parvoviruses infection. Virol J. https://doi.org/10.1186/s12985-015-0344-y
Yu TF, Ma B, Gao MC, Wang JW (2012) Localization of linear B-cell epitopes on goose parvovirus structural protein. Vet Immunol Immunopathol 145:522–526. https://doi.org/10.1016/j.vetimm.2011.11.022
Cotmore SF, Agbandje-McKenna M, Chiorini JA et al (2014) The family Parvoviridae. Arch Virol 159:1239–1247. https://doi.org/10.1007/s00705-013-1914-1
Yu T-F, Ma B, Wang J-W (2016) Identification of linear B-cell epitopes on goose parvovirus non-structural protein. Vet Immunol Immunopathol 179:85–88. https://doi.org/10.1016/j.vetimm.2016.08.009
Qiu Z, Tian W, Yu T et al (2012) Monoclonal antibodies against NS1 protein of goose parvovirus. Hybridoma 31:125–130. https://doi.org/10.1089/hyb.2011.0098
Wan C, Chen H, Fu Q et al (2015) Genomic characterization of goose parvovirus and muscovy duck parvovirus co-infection in Fujian, China. Kafkas Univ Vet Fak Derg 21:923–928. https://doi.org/10.9775/kvfd.2015.13848
Liu W-J, Yang Y-T, Zou H-Y et al (2020) Identification of recombination in novel goose parvovirus isolated from domesticated Jing-Xi partridge ducks in South China. Virus Genes 56:600–609. https://doi.org/10.1007/s11262-020-01781-1
Wang J, Ling J, Wang Z et al (2017) Molecular characterization of a novel Muscovy duck parvovirus isolate: evidence of recombination between classical MDPV and goose parvovirus strains. BMC Vet Res 13:1–10. https://doi.org/10.1186/s12917-017-1238-6
Yang Z, Bielawski JR (2000) Statistical methods for detecting molecular adaptation. Trends Ecol Evol 15:496–503. https://doi.org/10.1016/S0169-5347(00)01994-7
Stamenković GG, Ćirković VS, Šiljić MM et al (2016) Substitution rate and natural selection in parvovirus B19. Sci Rep 6:35759. https://doi.org/10.1038/srep35759
Battilani M, Scagliarini A, Ciulli S et al (2006) High genetic diversity of the VP2 gene of a canine parvovirus strain detected in a domestic cat. Virology 352:22–26. https://doi.org/10.1016/j.virol.2006.06.002
Kryazhimskiy S, Plotkin JB (2008) The population genetics of dN/dS. PLoS Genet. https://doi.org/10.1371/journal.pgen.1000304
Estevez C, Villegas P (2004) Sequence analysis, viral rescue from infectious clones and generation of recombinant virions of the avian adeno-associated virus. Virus Res 105(2):195–208
Shehata AA, Gerry DM, Heenemann K, Halami MY, Tokarzewski S, Wencel P, Vahlenkamp TW (2016) Goose parvovirus and circovirus coinfections in ornamental ducks. Avian Dis 60(2):516–522
Li P, Lin S, Zhang R, Chen J, Sun D, Lan J, Song S, Xie Z, Jiang S (2018) Isolation and characterization of novel goose parvovirus-related virus reveal the evolution of waterfowl parvovirus. Transbound Emerg Dis 65(2):e284–e295
Acknowledgements
This study did not receive any financial support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Human and animal participants
This article does not contain any studies with human or animal subjects performed by any of the authors.
Additional information
Handling Editor: Sheela Ramamoorthy.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Isidan, H., Turan, T., Atasoy, M.O. et al. Molecular analysis of goose parvovirus field strains from a Derzsy’s disease outbreak reveals local European-associated variants. Arch Virol 166, 1931–1942 (2021). https://doi.org/10.1007/s00705-021-05086-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00705-021-05086-y