Abstract
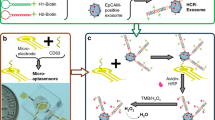
A simple nanoplatform based on molybdenum disulfide (MoS2) nanosheets, a fluorescence quencher (signal off), and a hybridization chain reaction (HCR) signal amplification (signal on) used for the enzyme-free, label-free, and low-background signal quantification of microRNA-21 in plasma exosome is reported. According to the sequence of microRNA-21, carboxy-fluorescein (FAM)-labeled hybridization probe 1 (FAM-H1) and hybridization probes 2 (FAM-H2) were designed with excitation maxima at 488 nm and emission maxima at 518 nm. MoS2 nanosheets could adsorb FAM-H1 and FAM-H2 and quenched their fluorescence signals to reduce the background signal. However, HCR was triggered when microRNA-21 was present. Consequently, HCR products containing a large number of FAM fluorophores can emit a strong fluorescence at 518 nm and could realize the detection of microRNA-21 as low as 6 pmol/L and had a wide linear relation of 0.01–25 nmol/L. This assay has the ability of single-base mismatch recognition and could identify microRNA-21 with high specificity. Most importantly, this approach was successfully applied to the detection of plasma exosomal microRNA-21 in patients with lung cancer, and it is proposed that other targets can also be detected by changing the FAM-H1 and FAM-H2 corresponding to the target sequence. Thus, a novel, hands-on strategy for liquid biopsy was proposed and has a potential application value in the early diagnosis of lung cancer.
Graphical abstract








Similar content being viewed by others
Data availability
All data and material generated or analyzed during this study are included in this published article and its supplementary information files.
Code availability
Not applicable.
References
Siegel RL, Miller KD, Fuchs HE, Jemal A (2021) Cancer statistics, 2021. CA Cancer J Clin 71:7–33. https://doi.org/10.3322/caac.21654
Terlizzi M, Colarusso C, Pinto A, Sorrentino R (2019) Drug resistance in non-small cell lung cancer (NSCLC): impact of genetic and non-genetic alterations on therapeutic regimen and responsiveness. Pharmacol Ther 202:140–148. https://doi.org/10.1016/j.pharmthera.2019.06.005
Goldstraw P, Chansky K, Crowley J et al (2016) The IASLC lung cancer staging project: proposals for revision of the TNM stage groupings in the forthcoming (Eighth) edition of the TNM classification for lung cancer. J Thorac Oncol 11:39–51. https://doi.org/10.1016/j.jtho.2015.09.009
Carvalho Â, Ferreira G, Seixas D et al (2021) Emerging lab-on-a-chip approaches for liquid biopsy in lung cancer: status in CTCs and ctDNA research and clinical validation. Cancers (Basel) 13:2101. https://doi.org/10.3390/cancers13092101
Pegtel DM, Gould SJ (2019) Exosomes. Annu Rev Biochem 88:487–514. https://doi.org/10.1146/annurev-biochem-013118-111902
Mashouri L, Yousefi H, Aref AR et al (2019) Exosomes: composition, biogenesis, and mechanisms in cancer metastasis and drug resistance. Mol Cancer 18:75. https://doi.org/10.1186/s12943-019-0991-5
Oliveto S, Mancino M, Manfrini N, Biffo S (2017) Role of microRNAs in translation regulation and cancer. World J Biol Chem 8:45–56. https://doi.org/10.4331/wjbc.v8.i1.45
Sanz-Rubio D, Martin-Burriel I, Gil A et al (2018) Stability of circulating exosomal miRNAs in healthy subjects. Sci Rep 8:10306. https://doi.org/10.1038/s41598-018-28748-5
Dejima H, Iinuma H, Kanaoka R et al (2017) Exosomal microRNA in plasma as a non-invasive biomarker for the recurrence of non-small cell lung cancer. Oncol Lett 13:1256–1263. https://doi.org/10.3892/ol.2017.5569
Esquela-Kerscher A, Slack FJ (2006) Oncomirs - microRNAs with a role in cancer. Nat Rev Cancer 6:259–269. https://doi.org/10.1038/nrc1840
Markou A, Tsaroucha EG, Kaklamanis L et al (2008) Prognostic value of mature microRNA-21 and microRNA-205 overexpression in non-small cell lung cancer by quantitative real-time RT-PCR. Clin Chem 54:1696–1704. https://doi.org/10.1373/clinchem.2007.101741
O’Bryan S, Dong S, Mathis JM, Alahari SK (2017) The roles of oncogenic miRNAs and their therapeutic importance in breast cancer. Eur J Cancer 72:1–11. https://doi.org/10.1016/j.ejca.2016.11.004
Dillhoff M, Liu J, Frankel W et al (2008) MicroRNA-21 is overexpressed in pancreatic cancer and a potential predictor of survival. J Gastrointest Surg 12:2171–2176. https://doi.org/10.1007/s11605-008-0584-x
Huang Q, Liu L, Liu C-H et al (2013) MicroRNA-21 regulates the invasion and metastasis in cholangiocarcinoma and may be a potential biomarker for cancer prognosis. Asian Pac J Cancer Prev 14:829–834. https://doi.org/10.7314/apjcp.2013.14.2.829
Zheng W, Zhao J, Tao Y et al (2018) MicroRNA-21: A promising biomarker for the prognosis and diagnosis of non-small cell lung cancer. Oncol Lett 16:2777–2782. https://doi.org/10.3892/ol.2018.8972
Chen C, Ridzon DA, Broomer AJ et al (2005) Real-time quantification of microRNAs by stem-loop RT-PCR. Nucleic Acids Res 33:e179. https://doi.org/10.1093/nar/gni178
Egloff S, Melnychuk N, Reisch A et al (2021) Enzyme-free amplified detection of cellular microRNA by light-harvesting fluorescent nanoparticle probes. Biosens Bioelectron 179:113084. https://doi.org/10.1016/j.bios.2021.113084
Li M, Xu X, Zhou Z et al (2020) Label-free detection of microRNA: two-stage signal enhancement with hairpin assisted cascade isothermal amplification and light-up DNA-silver nanoclusters. Microchim Acta 187:141. https://doi.org/10.1007/s00604-019-4094-1
Li F, Zhou Y, Yin H, Ai S (2020) Recent advances on signal amplification strategies in photoelectrochemical sensing of microRNAs. Biosens Bioelectron 166:112476. https://doi.org/10.1016/j.bios.2020.112476
Yang B, Zhang S, Fang X, Kong J (2019) Double signal amplification strategy for ultrasensitive electrochemical biosensor based on nuclease and quantum dot-DNA nanocomposites in the detection of breast cancer 1 gene mutation. Biosens Bioelectron 142:111544. https://doi.org/10.1016/j.bios.2019.111544
Borum RM, Jokerst JV (2021) Hybridizing clinical translatability with enzyme-free DNA signal amplifiers: recent advances in nucleic acid detection and imaging. Biomater Sci 9:347–366. https://doi.org/10.1039/D0BM00931H
Dirks RM, Pierce NA (2004) Triggered amplification by hybridization chain reaction. Proc Natl Acad Sci U S A 101:15275–15278. https://doi.org/10.1073/pnas.0407024101
Miao P, Tang Y (2020) Dumbbell hybridization chain reaction based electrochemical biosensor for ultrasensitive detection of exosomal miRNA. Anal Chem 92:12026–12032. https://doi.org/10.1021/acs.analchem.0c02654
Li N, Chen J, Luo M et al (2017) Highly sensitive chemiluminescence biosensor for protein detection based on the functionalized magnetic microparticles and the hybridization chain reaction. Biosens Bioelectron 87:325–331. https://doi.org/10.1016/j.bios.2016.08.067
Li X, Shan J, Zhang W et al (2017) Recent advances in synthesis and biomedical applications of two-dimensional transition metal dichalcogenide nanosheets. Small 13:1602660. https://doi.org/10.1002/smll.201602660
Kalantar-zadeh K, Ou JZ, Daeneke T et al (2015) Two-dimensional transition metal dichalcogenides in biosystems. Adv Funct Mater 25:5086–5099. https://doi.org/10.1002/adfm.201500891
Zhu C, Zeng Z, Li H et al (2013) Single-layer MoS2-based nanoprobes for homogeneous detection of biomolecules. J Am Chem Soc 135:5998–6001. https://doi.org/10.1021/ja4019572
Zhang Y, Zheng B, Zhu C et al (2015) Single-layer transition metal dichalcogenide nanosheet based nanosensors for rapid, sensitive, and multiplexed detection of DNA. Adv Mater 27:935–939. https://doi.org/10.1002/adma.201404568
Théry C, Amigorena S, Raposo G, Clayton A (2006) Isolation and characterization of exosomes from cell culture supernatants and biological fluids. Curr Protoc Cell Biol Chapter 3:Unit 3.22. https://doi.org/10.1002/0471143030.cb0322s30
Mahani M, Mousapour Z, Divsar F et al (2019) A carbon dot and molecular beacon based fluorometric sensor for the cancer marker microRNA-21. Microchim Acta 186:132. https://doi.org/10.1007/s00604-019-3233-z
Wang S, Wang L, Xu X et al (2019) MnO2 nanosheet-mediated ratiometric fluorescence biosensor for MicroRNA detection and imaging in living cells. Anal Chim Acta 1063:152–158. https://doi.org/10.1016/j.aca.2019.02.049
Xia Y, Chen T, Zhang L et al (2021) Colorimetric detection of exosomal microRNA through switching the visible-light-induced oxidase mimic activity of acridone derivate. Biosens Bioelectron 173:112834. https://doi.org/10.1016/j.bios.2020.112834
Fang H, Xie N, Ou M et al (2018) Detection of nucleic acids in complex samples via magnetic microbead-assisted catalyzed hairpin assembly and “DD-A” FRET. Anal Chem 90:7164–7170. https://doi.org/10.1021/acs.analchem.8b01330
Fakhri N, Abarghoei S, Dadmehr M et al (2020) Paper based colorimetric detection of miRNA-21 using Ag/Pt nanoclusters. Spectrochim Acta Part A Mol Biomol Spectrosc 227:117529. https://doi.org/10.1016/j.saa.2019.117529
Raposo G, Stoorvogel W (2013) Extracellular vesicles: exosomes, microvesicles, and friends. J Cell Biol 200:373–383. https://doi.org/10.1083/jcb.201211138
Watabe S, Kikuchi Y, Morita S et al (2020) Clinicopathological significance of microRNA-21 in extracellular vesicles of pleural lavage fluid of lung adenocarcinoma and its functions inducing the mesothelial to mesenchymal transition. Cancer Med 9:2879–2890. https://doi.org/10.1002/cam4.2928
Gao Z, Yuan H, Mao Y et al (2021) In situ detection of plasma exosomal microRNA for lung cancer diagnosis using duplex-specific nuclease and MoS2 nanosheets. Analyst 146:1924–1931. https://doi.org/10.1039/d0an02193h
Funding
This research was supported by the National Natural Science Foundation of China (No. 81973099).
Author information
Authors and Affiliations
Contributions
Conceptualization: Yongjun Wu. Methodology: Yanhua Mao and Jinlan Wei. Formal analysis and investigation: Jinlan Wei, Longjie Wu. Writing, original draft preparation: Sitian He and Jinlan Wei. Writing review and editing: Yongjun Wu, Hongchao Guo, Clement Yaw Effah, Jinlan Wei, Xinlian Liu. Resources: Yanhua Mao, Sitian He, Jinlan Wei, Yongjun Wu. Supervision: Yongjun Wu, Sitian He.
Corresponding author
Ethics declarations
Ethics approval
Not applicable.
Consent to participate
All sampled subjects were informed and consented.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Wei, J., He, S., Mao, Y. et al. A simple “signal off–on” fluorescence nanoplatform for the label-free quantification of exosome-derived microRNA-21 in lung cancer plasma. Microchim Acta 188, 397 (2021). https://doi.org/10.1007/s00604-021-05051-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00604-021-05051-1