Abstract
Background
The role of mitophagy in various cancer-associated biological processes is well recognized. Nonetheless, the comprehensive implications of mitophagy in clear cell renal cell carcinoma (ccRCC) necessitate further exploration.
Methods
Based on the transcriptomic data encompassing 25 mitophagy-related genes (MRGs), we identified the distinct mitophage patterns in 763 ccRCC samples. Subsequently, a mitophage-related predictive signature with machine learning algorithms was constructed, designated as RiskScore, to quantify the individual mitophagy status in ccRCC patients. Employing multispectral immunofluorescence (mIF) and immunohistochemistry (IHC) staining, we detected the effect of PTEN-induced putative kinase 1 (PINK1) in the prognosis and immune microenvironment of ccRCC.
Results
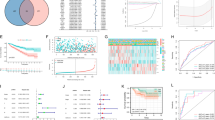
Our analysis initially encompassed a comprehensive assessment of the expression profiling, genomic variations, and interactions among the 25 MRGs in ccRCC. Subsequently, the consensus clustering algorithm was applied to stratify ccRCC patients into three clusters with distinct prognostic outcomes, tumor microenvironment (TME) characteristics, and underlying biological pathways. We screened eight pivotal genes (CLIC4, PTPRB, SLC16A12, ENPP5, FLRT3, HRH2, PDK4, and SCD5) to construct a mitophagy-related predictive signature, which showed excellent prognostic value for ccRCC patients. Moreover, patient subgroups divided by the RiskScore showed contrasting expression levels of immune checkpoints (ICPs), abundance of immune cells, and immunotherapy response. Additionally, a nomogram was established with robust predictive power integrating the RiskScore and clinical features. Notably, we observed that PINK1 expression markedly correlated with favorable treatment response and advanced maturation stages of tertiary lymphoid structures, which potentially shed light on enhancing anti-tumor immunity of ccRCC.
Conclusion
Collectively, this study initially developed a signature associated with mitophagy, which demonstrated an excellent ability to predict the clinical prognosis, TME characterization, and responsiveness to targeted therapy and immunotherapy for ccRCC patients. Of particular note is the pivotal role of PINK1 in mediating the treatment response and immune microenvironment for ccRCC patients.








Similar content being viewed by others
Data availability
The datasets used and/or analyzed in the study are available from the corresponding author upon reasonable request.
Abbreviations
- ccRCC:
-
Clear cell renal cell carcinoma
- CDF:
-
Cumulative distribution function
- CNVs:
-
Copy number variations
- CPTAC:
-
Clinical Proteomic Tumor Analysis Consortium
- DAMPs:
-
Danger-associated molecular patterns
- DCA:
-
Decision curve analysis
- DEGs:
-
Differential expression genes
- EMBL:
-
European Molecular Biology Laboratory
- FPKM:
-
Fragments per kilobase of transcript per million
- FUSCC:
-
Fudan University Shanghai Cancer Center
- GO:
-
Gene Ontology
- GSVA:
-
Gene set variation analysis
- ICGC:
-
International Cancer Genome Consortium
- ICIs:
-
Immune checkpoint inhibitors
- ICPs:
-
Immune checkpoints
- IHC:
-
Immunohistochemistry
- IMDC:
-
International Metastatic Renal Cell Carcinoma Database Consortium
- KEGG:
-
Kyoto Encyclopedia of Genes and Genomes
- LASSO:
-
Least absolute shrinkage and selector operation
- LOH:
-
Loss of heterozygosity
- mIF:
-
Multispectral immunofluorescence
- MRGs:
-
Mitophagy-related genes
- MSKCC:
-
Memorial Sloan Kettering Cancer Center
- OS:
-
Overall survival
- PCA:
-
Principal component analysis
- PD-1:
-
Programmed cell death protein 1
- PINK1:
-
PTEN-induced putative kinase 1
- PPI:
-
Protein–protein interaction
- RCC:
-
Renal cell carcinoma
- ROC:
-
Receiver operating characteristic
- ROS:
-
Reactive oxygen species
- ssGSEA:
-
Single-sample gene set enrichment analysis
- TCGA:
-
The cancer genome atlas
- TIDE:
-
Tumor immune dysfunction and exclusion
- TKI:
-
Tyrosine kinase inhibitors
- TMB:
-
Tumor mutation burden
- TME:
-
Tumor microenvironment
- TPM:
-
Transcripts per million
- VEGF:
-
Vascular endothelial growth factor
- VHL:
-
Von Hippel–Lindau
References
Anwaier A et al (2023) Tumor microenvironment-based signatures distinguish intratumoral heterogeneity, prognosis, and immunogenomic features of clear cell renal cell carcinoma. J Natl Cancer Center. https://doi.org/10.1016/j.jncc.2023.08.003
Braun DA et al (2021) Progressive immune dysfunction with advancing disease stage in renal cell carcinoma. Cancer Cell 39(5):632-648 e8
Cai J et al (2019) The role of PD-1/PD-L1 axis and macrophage in the progression and treatment of cancer. J Cancer Res Clin Oncol 145(6):1377–1385
Capitanio U et al (2021) Parenchymal biopsy in the management of patients with renal cancer. World J Urol 39(8):2961–2968
Chen W et al (2016) Cancer statistics in China, 2015. CA Cancer J Clin 66(2):115–132
Chevrier S et al (2017) An immune atlas of clear cell renal cell carcinoma. Cell 169(4):736-749 e18
Clark DJ et al (2019) Integrated proteogenomic characterization of clear cell renal cell carcinoma. Cell 179(4):964-983 e31
Doblado L et al (2021) Mitophagy in human diseases. Int J Mol Sci 22(8):3903
Geissler K et al (2015) Immune signature of tumor infiltrating immune cells in renal cancer. Oncoimmunology 4(1):e985082
Gilkes DM, Semenza GL, Wirtz D (2014) Hypoxia and the extracellular matrix: drivers of tumour metastasis. Nat Rev Cancer 14(6):430–439
Hanzelmann S, Castelo R, Guinney J (2013) GSVA: gene set variation analysis for microarray and RNA-seq data. BMC Bioinformatics 14:7
Helmink BA et al (2020) B cells and tertiary lymphoid structures promote immunotherapy response. Nature 577(7791):549–555
Horeweg N et al (2022) Tertiary lymphoid structures critical for prognosis in endometrial cancer patients. Nat Commun 13(1):1373
Hu X et al (2022) Bioinformatics-led discovery of osteoarthritis biomarkers and inflammatory infiltrates. Front Immunol 13:871008
Hunter MV et al (2021) Spatially resolved transcriptomics reveals the architecture of the tumor-microenvironment interface. Nat Commun 12(1):6278
Jiang P et al (2018) Signatures of T cell dysfunction and exclusion predict cancer immunotherapy response. Nat Med 24(10):1550–1558
Kim MC et al (2021) Updates on immunotherapy and immune landscape in renal clear cell carcinoma. Cancers (Basel) 13(22):5856
Korolchuk VI et al (2017) Mitochondria in cell senescence: is mitophagy the weakest link? EBioMedicine 21:7–13
Leek JT et al (2012) The sva package for removing batch effects and other unwanted variation in high-throughput experiments. Bioinformatics 28(6):882–883
Li Y et al (2022a) Histopathologic and proteogenomic heterogeneity reveals features of clear cell renal cell carcinoma aggressiveness. Cancer Cell 41(1):139–163
Li X et al (2022b) Immune checkpoint blockade in pancreatic cancer: trudging through the immune desert. Semin Cancer Biol. https://doi.org/10.1016/j.semcancer.2022.08.009
Li Y et al (2023) PINK1-mediated mitophagy promotes oxidative phosphorylation and redox homeostasis to induce drug-tolerant persister cancer cells. Cancer Res 83(3):398–413
Luo J et al (2023) Enhanced mitophagy driven by ADAR1-GLI1 editing supports the self-renewal of cancer stem cells in hepatocellular carcinoma. Hepatology. https://doi.org/10.1097/HEP.0000000000000299
Marquardt A et al (2021) Subgroup-independent mapping of renal cell carcinoma-machine learning reveals prognostic mitochondrial gene signature beyond histopathologic boundaries. Front Oncol 11:621278
Meylan M et al (2022) Tertiary lymphoid structures generate and propagate anti-tumor antibody-producing plasma cells in renal cell cancer. Immunity 55(3):527-541 e5
Munoz-Erazo L et al (2020) Tertiary lymphoid structures in cancer—considerations for patient prognosis. Cell Mol Immunol 17(6):570–575
Panigrahi DP et al (2020) The emerging, multifaceted role of mitophagy in cancer and cancer therapeutics. Semin Cancer Biol 66:45–58
Peter MR et al (2021) Investigating urinary circular RNA biomarkers for improved detection of renal cell carcinoma. Front Oncol 11:814228
Poole LP, Macleod KF (2021) Mitophagy in tumorigenesis and metastasis. Cell Mol Life Sci 78(8):3817–3851
Qu Y et al (2022) A proteogenomic analysis of clear cell renal cell carcinoma in a Chinese population. Nat Commun 13(1):2052
Senbabaoglu Y et al (2016) Tumor immune microenvironment characterization in clear cell renal cell carcinoma identifies prognostic and immunotherapeutically relevant messenger RNA signatures. Genome Biol 17(1):231
Siegel RL et al (2023) Cancer statistics, 2023. CA Cancer J Clin 73(1):17–48
Silina K et al (2018) Germinal centers determine the prognostic relevance of tertiary lymphoid structures and are impaired by corticosteroids in lung squamous cell carcinoma. Cancer Res 78(5):1308–1320
Siska PJ et al (2017) Mitochondrial dysregulation and glycolytic insufficiency functionally impair CD8 T cells infiltrating human renal cell carcinoma. JCI Insight. https://doi.org/10.1172/jci.insight.93411
Tang H et al (2023) Heterogeneity and function of cancer-associated fibroblasts in renal cell carcinoma. J Natl Cancer Center 3(2):100–105
Tian X et al (2023) Special issue “The advance of solid tumor research in China”: Multi-omics analysis based on 1311 clear cell renal cell carcinoma samples identifies a glycolysis signature associated with prognosis and treatment response. Int J Cancer 152(1):66–78
Vesely MD, Zhang T, Chen L (2022) Resistance Mechanisms to Anti-PD Cancer Immunotherapy. Annu Rev Immunol 40:45–74
Wang L et al (2018) PTEN-L is a novel protein phosphatase for ubiquitin dephosphorylation to inhibit PINK1-Parkin-mediated mitophagy. Cell Res 28(8):787–802
Xu WH et al (2019a) Prognostic implications of aquaporin 9 expression in clear cell renal cell carcinoma. J Transl Med 17(1):363
Xu WH et al (2019b) Prognostic value and immune infiltration of novel signatures in clear cell renal cell carcinoma microenvironment. Aging (Albany NY) 11(17):6999–7020
Xu W et al (2021a) Prognostic immunophenotyping clusters of clear cell renal cell carcinoma defined by the unique tumor immune microenvironment. Front Cell Dev Biol 9:785410
Xu W et al (2021b) Systematic genome-wide profiles reveal alternative splicing landscape and implications of splicing regulator DExD-box helicase 21 in aggressive progression of adrenocortical carcinoma. Phenomics 1(6):243–256
Xu W et al (2021c) Hexokinase 3 dysfunction promotes tumorigenesis and immune escape by upregulating monocyte/macrophage infiltration into the clear cell renal cell carcinoma microenvironment. Int J Biol Sci 17(9):2205–2222
Xu W et al (2022) The unique genomic landscape and prognostic mutational signature of Chinese clear cell renal cell carcinoma. J Natl Cancer Center. https://doi.org/10.1016/j.jncc.2022.07.001
Xu W et al (2022a) Genomic alteration of MTAP/CDKN2A predicts sarcomatoid differentiation and poor prognosis and modulates response to immune checkpoint blockade in renal cell carcinoma. Front Immunol 13:953721
Xu W et al (2022b) Tumor-associated macrophage-derived chemokine CCL5 facilitates the progression and immunosuppressive tumor microenvironment of clear cell renal cell carcinoma. Int J Biol Sci 18(13):4884–4900
Xu Y et al (2022c) PINK1 deficiency in gastric cancer compromises mitophagy, promotes the Warburg effect, and facilitates M2 polarization of macrophages. Cancer Lett 529:19–36
Xu W et al (2023) Stimuli-responsive nanodelivery systems for amplifying immunogenic cell death in cancer immunotherapy. Immunol Rev. https://doi.org/10.1111/imr.13237
Yang JF et al (2019) Screening, identification and validation of CCND1 and PECAM1/CD31 for predicting prognosis in renal cell carcinoma patients. Aging (Albany NY) 11(24):12057–12079
Zaffuto E, Karakiewicz PI, Capitanio U (2016) Complete response after treatment with first-line targeted anti-vascular endothelial growth factor therapy in metastatic renal cancer: what next? Ann Transl Med 4(15):291
Zheng J et al (2021) Traditional Chinese medicine Bu-Shen-Jian-Pi-Fang attenuates glycolysis and immune escape in clear cell renal cell carcinoma: results based on network pharmacology. Biosci Rep. https://doi.org/10.1042/BSR20204421
Zheng Y et al (2021) STOML2 potentiates metastasis of hepatocellular carcinoma by promoting PINK1-mediated mitophagy and regulates sensitivity to lenvatinib. J Hematol Oncol 14(1):16
Zheng R et al (2022) Cancer incidence and mortality in China, 2016. J Natl Cancer Center. https://doi.org/10.1016/j.jncc.2022.02.002
Acknowledgements
We are grateful to all patients for their dedicated participation in the study.
Funding
This study was partially supported by the grant of Longhua Hospital of Shanghai University of Traditional Chinese Medicine (YW002.017).
Author information
Authors and Affiliations
Contributions
JY, JX, and WL: helped in conceptualization. JX, WL, and SL: contributed to data curation and formal analysis. JX, WL, and HT: worked in investigation and methodology. HT and TW: worked in resources and software. JY, HT, and TW: worked in supervision. JX, JY, and WL: contributed to original draft. JY, HT, and TW: helped in editing.
Corresponding authors
Ethics declarations
Conflict of interest
The authors report no potential conflicts of interest.
Ethical approval and consent to participate
The study design and test procedures followed the Helsinki Declaration II. The ethics approval and consent to participate of the Urology and Pathology departments in this study were approved by the ethics committee of Renji Hospital, Shanghai Jiao Tong University School of Medicine.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
432_2023_5349_MOESM1_ESM.pdf
Supplementary file1 SUPPLEMENTARY FIGURE 1. Survival analysis of ccRCC patients stratified by the expression of MRGs. (A-H) Kaplan–Meier analysis of the OS of ccRCC patients stratified by the expression of MRGs. SUPPLEMENTARY FIGURE 2. Generation of mitophagy-related clusters by unsupervised clustering analysis. (A-H) Consensus matrix heatmaps of unsupervised clustering from k = 2 to k = 9. (I-K) The cumulative distribution function plot, delta plot, and tracking plot from k = 2 to k = 9. SUPPLEMENTARY FIGURE 3. Development of GeneClusters by unsupervised clustering analysis. (A-H) Consensus matrix heatmaps of unsupervised clustering from k = 2 to k = 9. (I-K) The cumulative distribution function plot, delta plot, and tracking plot from k = 2 to k = 9. SUPPLEMENTARY FIGURE 4. Correlation between RiskGroup and clinical features. (A-F) Proportion of clinical features (age and gender) in low and high RiskGroup. (G-L) Survival analysis of ccRCC patients between two RiskGroups in different subgroups stratified by clinical features (age and gender). (PDF 1834 KB)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Xiang, J., Liu, W., Liu, S. et al. Deciphering the implications of mitophagy-related signatures in clinical outcomes and microenvironment heterogeneity of clear cell renal cell carcinoma. J Cancer Res Clin Oncol 149, 16015–16030 (2023). https://doi.org/10.1007/s00432-023-05349-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-023-05349-y