Abstract
Background
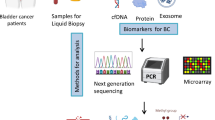
Bladder cancer (BCa) has a high incidence and recurrence rate worldwide. So far, there is no noninvasive detection of BCa therapy and prognosis based on urine multi-omics. Therefore, it is necessary to explore noninvasive predictive models and novel treatment modalities for BCa.
Methods
First, we performed protein analysis of urine from five BCa patients and five healthy individuals using liquid chromatography–tandem mass spectrometry (LC–MS/MS). Combining multi-omics data to mine particular and sensitive molecules to predict BCa prognosis. Second, urine proteomics data were combined with TCGA transcriptome data to select differential genes that were specifically highly expressed in urine and tissues. Further, the Lasso equation was used to screen specific molecules to construct a noninvasive prediction model of BCa. Finally, natural compounds of specific molecules were selected by combined network pharmacology and molecular docking to complete molecular structure docking.
Results
A noninvasive predictive model was constructed using PSMB5, P4HB, S100A16, GET3, CNP, TFRC, DCXR, and MPZL1, specific molecules screened by multi-omics, and clinical features, which had good predictive value at 1, 3, and 5 years of prediction. High expression of these target genes suggests a poor prognosis in patients with BCa, and they were mainly involved in cell adhesion molecules and the IGF pathway. In addition, the corresponding drugs and natural compounds were selected by network pharmacology, and the molecular structure 7NHT of PSMB5 was found to be well docked to Ellagic acid, a natural compound in Hetaoren that we found. The 3D structure 6I7S of P4HB was able to bind to Stigmasterol in Shanzha stably, and the structure 6WRV of TFRC as an iron transport carrier was also able to bind to Stigmasterol in Shanzha stably. The structures 1WOJ, 3D3W, and 6IGW of CNP, DCXR, and MPZL1 can also play an important role in combination with the natural compounds (S)-Stylopine, Kryptoxanthin, and Sitosterol in Maqianzi, Yumixu, and Laoguancao.
Conclusion
The noninvasive prediction model based on urinomics had excellent potential in predicting the prognosis of patients with BCa. The multi-omics screening of specific molecules combined with pharmacology and compound molecular docking can promote the research and development of novel drugs.










Similar content being viewed by others
Data availability
All data and material analyzed during this study are included in this article.
References
Barrio S et al (2019) Spectrum and functional validation of PSMB5 mutations in multiple myeloma. Leukemia 33(2):447–456
Bhat A, Kwon D, Soodana-Prakash N, Mouzannar A, Punnen S, Gonzalgo ML, Parekh DJ, Ritch CR (2021) Surveillance intensity in intermediate risk, nonmuscle invasive bladder cancer: Revisiting the optimal timing and frequency of cystoscopy. J Urol 206(1):22–28
Catellani C, Ravegnini G, Sartori C, Angelini S, Street ME (2021) GH and IGF System: The regulatory role of miRNAs and lncRNAs in Cancer. Front Endocrinol (Lausanne) 12:701246
Dong X, Premaratne ID, Sariibrahimoglu K, Limem S, Scott J, Gadjiko M, Berri N, Ginter P, Spector JA (2022) 3D-printed poly-4-hydroxybutyrate bioabsorbable scaffolds for nipple reconstruction. Acta Biomater 143:333–343
Dos Santos Maia M, Soares Rodrigues GC, Silva Cavalcanti AB, Scotti L, Scotti MT (2020) Consensus analyses in molecular docking studies applied to medicinal chemistry. Mini Rev Med Chem 20(14):1322–1340
Ebert B, Kisiela M, Maser E (2015) Human DCXR - another 'moonlighting protein' involved in sugar metabolism, carbonyl detoxification, cell adhesion and male fertility?. Biol Rev Camb Philos Soc 90(1):254–278
Fan J, Du W, Zhang H, Wang Y, Li K, Meng Y, Wang J (2020) Transcriptional downregulation of miR-127-3p by CTCF promotes prostate cancer bone metastasis by targeting PSMB5. FEBS Lett 594(3):466–476
Fang J, Ji WH, Wang FZ, Xie TM, Wang L, Fu ZF, Wang Z, Yan FQ, Shen QL, Ye ZM (2021) Circular RNA hsa_circ_0000700 promotes cell proliferation and migration in Esophageal Squamous Cell Carcinoma by sponging miR-1229. J Cancer 12(9):2610–2623
Ferragut F, Vachetta VS, Troncoso MF, Rabinovich GA, Elola MT (2021) ALCAM/CD166: A pleiotropic mediator of cell adhesion, stemness and cancer progression. Cytokine Growth Factor Rev 61:27–37
Gaczynska M, Osmulski PA (2018) Targeting protein-protein interactions in the ubiquitin-proteasome pathway. Adv Protein Chem Struct Biol 110:123–165
Goel A, Ward DG, Noyvert B, Yu M, Gordon NS, Abbotts B, Colbourne JK, Kissane S, James ND, Zeegers MP, Cheng KK, Cazier JB, Whalley CM, Beggs AD, Palles C, Arnold R, Bryan RT (2022) Combined exome and transcriptome sequencing of non-muscle-invasive bladder cancer: associations between genomic changes, expression subtypes, and clinical outcomes. Genome Med 14(1):59
Huang Y, Huang J, Huang Y, Gan L, Long L, Pu A, Xie R (2020) TFRC promotes epithelial ovarian cancer cell proliferation and metastasis via up-regulation of AXIN2 expression. Am J Cancer Res 10(1):131–147
Ikeda A et al (2020) Risk for intravesical recurrence of bladder cancer stratified by the results on two consecutive UroVysion fluorescence in situ hybridization tests: a prospective follow-up study in Japan. Int J Clin Oncol 25(6):1163–1169
Karreman MA, Winkler F (2020) Targeting an adhesion molecule to prevent brain colonization of lung cancer. Neuro Oncol 22(7):899–900
Kohada Y et al (2021) Recurrence- and progression-free survival in intermediate-risk non-muscle-invasive bladder cancer: the impact of conditional evaluation and subclassification. BJU Int 127(4):473–485
Liang Y, Ye F, Xu C, Zou L, Hu Y, Hu J, Jiang H (2021) A novel survival model based on a Ferroptosis-related gene signature for predicting overall survival in bladder cancer. BMC Cancer 21(1):943
Lindskrog SV et al (2021) An integrated multi-omics analysis identifies prognostic molecular subtypes of non-muscle-invasive bladder cancer. Nat Commun 12(1):2301
Lopez-Beltran A, Cheng L, Gevaert T, Blanca A, Cimadamore A, Santoni M, Massari F, Scarpelli M, Raspollini MR, Montironi R (2020) Current and emerging bladder cancer biomarkers with an emphasis on urine biomarkers. Expert Rev Mol Diagn 20(2):231–243
Lu X, Meng J, Su L, Jiang L, Wang H, Zhu J, Huang M, Cheng W, Xu L, Ruan X, Yeh S, Liang C, Yan F (2021) Multi-omics consensus ensemble refines the classification of muscle-invasive bladder cancer with stratified prognosis, tumour microenvironment and distinct sensitivity to frontline therapies. Clin Transl Med 11(12):e601
Lyu L, Xiang W, Zheng F, Huang T, Feng Y, Yuan J, Zhang C (2020) Significant Prognostic Value of the Autophagy-Related Gene P4HB in Bladder Urothelial Carcinoma. Front Oncol 10:1613
Mancarella C, Morrione A, Scotlandi K (2021) Novel regulators of the IGF system in cancer. Biomolecules 11(2):273
Nogales C, Mamdouh ZM, List M, Kiel C, Casas AI, Schmidt HHHW (2022) Network pharmacology: curing causal mechanisms instead of treating symptoms. Trends Pharmacol Sci 43(2):136–150
Pinzi L, Rastelli G (2019) Molecular docking: Shifting paradigms in drug discovery. Int J Mol Sci 20(18):4331
Rahrmann EP, Wolf NK, Otto GM, Heltemes-Harris L, Ramsey LB, Shu J, LaRue RS, Linden MA, Rathe SK, Starr TK, Farrar MA, Moriarity BS, Largaespada DA (2019) Sleeping Beauty Screen Identifies RREB1 and Other Genetic Drivers in Human B-cell Lymphoma. Mol Cancer Res 17(2):567–582
Ravvaz K, Walz ME, Weissert JA, Downs TM (2017) Predicting nonmuscle invasive bladder cancer recurrence and progression in a united states population. J Urol 198(4):824–831
Sharouf F, Zaben M, Lammie A, Leach P, Bhatti MI (2018) Neurocutaneous melanosis presenting with hydrocephalus and malignant transformation: case-based update. Childs Nerv Syst 34(8):1471–1477
Siegel RL, Miller KD, Fuchs HE, Jemal A (2022) Cancer statistics. CA Cancer J Clin 72(1):7–33
Springer S.U et al (2018) Non-invasive detection of urothelial cancer through the analysis of driver gene mutations and aneuploidy. Elife 7:e32143
Su H, Tao T, Yang Z, Kang X, Zhang X, Kang D, Wu S, Li C (2019) Circular RNA cTFRC acts as the sponge of MicroRNA-107 to promote bladder carcinoma progression. Mol Cancer 18(1):27
Suh J, Han D, Ku JH, Kim HH, Kwak C, Jeong CW (2022) Next-generation Proteomics-Based Discovery, Verification, and Validation of Urine Biomarkers for Bladder Cancer Diagnosis. Cancer Res Treat 54(3):882–893
Sun D, Gou H, Wang D, Li C, Li Y, Su H, Wang X, Zhang X, Yu J (2022) LncRNA TNFRSF10A-AS1 promotes gastric cancer by directly binding to oncogenic MPZL1 and is associated with patient outcome. Int J Biol Sci 18(8):3156–3166
Valenberg F et al (2019) Prospective validation of an mRNA-based urine test for surveillance of patients with bladder cancer. Eur Urol 75(5):853–860
Wang CY, Li CY, Hsu HP, Cho CY, Yen MC, Weng TY, Chen WC, Hung YH, Lee KT, Hung JH, Chen YL, Lai MD (2017) PSMB5 plays a dual role in cancer development and immunosuppression. Am J Cancer Res 7(11):2103–2120
Wang Y, Chen L, Yu M, Fang Y, Qian K, Wang G, Ju L, Xiao Y, Wang X (2020) Immune-related signature predicts the prognosis and immunotherapy benefit in bladder cancer. Cancer Med 9(20):7729–7741
Wu J, Minikes AM, Gao M, Bian H, Li Y, Stockwell BR, Chen ZN, Jiang X (2019) Intercellular interaction dictates cancer cell ferroptosis via NF2-YAP signalling. Nature 572(7769):402–406
Wu Y, Peng Y, Guan B, He A, Yang K, He S, Gong Y, Li X, Zhou L (2021) P4HB: A novel diagnostic and prognostic biomarker for bladder carcinoma. Oncol Lett 21(2):95
Xu N et al (2022) Integrated proteogenomic characterization of urothelial carcinoma of the bladder. J Hematol Oncol 15(1):76
Zhang J, Guo S, Wu Y, Zheng ZC, Wang Y, Zhao Y (2019) P4HB, a Novel Hypoxia Target Gene Related to Gastric Cancer Invasion and Metastasis. Biomed Res Int 2019:9749751
Funding
This work was supported by the regulatory mechanism of AMPK in ischemic–reperfusion injury and fibrosis in renal transplantation (CY2015-YJRC08); The Open Foundation of Gansu Key Laboratory of Functional Genomics and Molecular Diagnostics; Gansu Province Intellectual Property Planning project (21ZSCQ012); The Second Hospital of Lanzhou University "Cuiying Science and Technology Innovation" project (CY2021-QN-A20).
Author information
Authors and Affiliations
Contributions
SW wrote and edited the manuscript. JC and SC edited the manuscript. JY performed the statistical analyses and generated the figures. HW, CW, and KL collected the public data. LY revised and reviewed the manuscript. All authors subsequently critically edited and revised the report. All authors read and approved the final report. The corresponding author had full access to all the data and final responsibility to submit for publication.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest regarding the publication of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Wan, S., Cao, J., Chen, S. et al. Construction of noninvasive prognostic model of bladder cancer patients based on urine proteomics and screening of natural compounds. J Cancer Res Clin Oncol 149, 281–296 (2023). https://doi.org/10.1007/s00432-022-04524-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00432-022-04524-x