Abstract
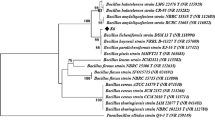
Laccases (EC 1.10.3.2) are considered one of the most prominent multicopper enzymes that exhibit the inherent properties of oxidizing a range of phenolic substrates. Mostly, reported laccases have been isolated from the plants and fungi species, whereas bacterial laccases are yet to be explored. Bacterial laccases have numerous distinctive properties over fungal laccases, including stability at high temperatures and high pH. This study includes the isolation of bacteria through the soil sample collected from the paper and pulp industry; the highest laccase-producing bacteria was identified as Bhargavaea bejingensis, using 16S rRNA gene sequencing. The extracellular and intracellular activities after 24 h incubation were 1.41 U/mL and 4.95 U/mL, respectively. The laccase-encoding gene of the bacteria was sequenced; moreover, the in vitro translated protein was bioinformatically characterized and asserted that the laccase produced by the bacteria Bhargavaea bejingensis was structurally and sequentially homologous to the CotA protein of Bacillus subtilis. The enzyme laccase produced from B. bejingensis was classified as three-domain laccase with several copper-binding residues, where a few crucial copper-binding residues of the laccase enzyme were also predicted.





Similar content being viewed by others
Data Availability
All the experimental data and material used are given within the manuscript.
Code Availability
Not applicable.
References
Kumar M, Kumar P, Das P, Solanki R, Kapur MK (2020) Potential applications of extracellular enzymes from Streptomyces spp. in various industries. Arch Microbiol 202(7):1597–1615. https://doi.org/10.1007/s00203-020-01898-9
Chio C, Sain M, Qin W (2019) Lignin utilization: a review of lignin depolymerization from various aspects. Renew Sustain Energy Rev 107:232–249. https://doi.org/10.1016/j.rser.2019.03.008
Mayolo-Deloisa K, González-González M, Rito-Palomares M (2020) Laccases in the food industry: bioprocessing, potential industrial and biotechnological applications. Front Bioeng Biotechnol 8:222. https://doi.org/10.3389/fbioe.2020.00222
Sharma N, Leung IK (2021) Novel thermophilic bacterial laccase for the degradation of aromatic organic pollutants. Front Chem. https://doi.org/10.3389/fchem.2021.711345
Chmelova D, Legerska B, Kunstova J, Ondrejovic M, Miertus S (2022) The production of laccases by white-rot fungi under solid-state fermentation conditions. World J Microbiol Biotechnol 38(2):1–20. https://doi.org/10.1007/s11274-021-03207-y
Chauhan PS, Goradia B, Saxena A (2017) Bacterial laccase: recent update on production, properties and industrial applications. 3 Biotech 7(5):1–20. https://doi.org/10.1007/s13205-017-0955-7
Prins A, Kleinsmidt L, Khan N, Kirby B, Kudanga T, Vollmer J, Le Roes-Hill M (2015) The effect of mutations near the T1 copper site on the biochemical characteristics of the small laccase from Streptomyces coelicolor A3 (2). Enzym Microb Technol 68:23–32. https://doi.org/10.1016/j.enzmictec.2014.10.003
Dai S, Yao Q, Yu G, Liu S, Yun J, Xiao X, Deng Z, Li H (2021) Biochemical characterization of a novel bacterial laccase and improvement of its efficiency by directed evolution on dye degradation. Front Microbiol 12:633004. https://doi.org/10.3389/fmicb.2021.633004
Givaudan A, Effosse A, Faure D, Potier P, Bouillant ML, Bally R (1993) Polyphenol oxidase in Azospirillum lipoferum isolated from rice rhizosphere: evidence for laccase activity in non-motile strains of Azospirillum lipoferum. FEMS Microbiol Lett 108(2):205–210. https://doi.org/10.1111/j.1574-6968.1993.tb06100.x
Mohammadian M, Fathi-Roudsari M, Mollania N, Badoei-Dalfard A, Khajeh K (2010) Enhanced expression of a recombinant bacterial laccase at low temperature and microaerobic conditions: purification and biochemical characterization. J Ind Microbiol Biotechnol 37(8):863–869. https://doi.org/10.1007/s10295-010-0734-5
Callejon S, Sendra R, Ferrer S, Pardo I (2016) Cloning and characterization of a new laccase from Lactobacillus plantarum J16 CECT 8944 catalyzing biogenic amines degradation. Appl Microbiol Biotechnol 100(7):3113–3124. https://doi.org/10.1007/s00253-015-7158-0
Jiang YP, Cai JL, Pei JJ, Li Q, Zhao LG (2021) Cloning, overexpression, and characterization of a thermostable, organic solvent-tolerant laccase from Bacillus pumilus ARA and its application to dye decolorization. ACS Omega 6(14):9741–9749. https://doi.org/10.1021/acsomega.1c00370
Chopra NK, Sondhi S (2022) Cloning, expression and characterization of laccase from Bacillus licheniformis NS2324 in E. coli application in dye decolorization. Int J Biol Macromol 206:1003–1011. https://doi.org/10.1016/j.ijbiomac.2022.03.104
Haq I, Mazumder P, Kalamdhad AS (2020) Recent advances in removal of lignin from paper industry wastewater and its industrial applications—a review. Bioresour Technol 312:123636. https://doi.org/10.1016/j.biortech.2020.123636
Singh AK, Chandra R (2019) Pollutants released from the pulp paper industry: aquatic toxicity and their health hazards. Aquat Toxicol 211:202–216. https://doi.org/10.1016/j.aquatox.2019.04.007
Degryse E, Glansdorff N, Piérard A (1978) A comparative analysis of extreme thermophilic bacteria belonging to the genus Thermus. Arch Microbiol 117(2):189–196. https://doi.org/10.1007/BF00402307
Dhiman K, Shirkot P (2015) Bioprospecting and molecular characterization of laccase producing bacteria from paper mills of Himachal Pradesh. Proc Natl Acad Sci India Sect B Biol Sci 85(4):1095–1103. https://doi.org/10.1007/s40011-015-0541-x
Kalra K, Chauhan R, Shavez M, Sachdeva S (2013) Isolation of laccase producing Trichoderma Spp. and effect. Int J ChemTech Res 5:2229–2235. https://doi.org/10.20902/IJCR.2013.6547861
Wang YS, Liu L, Fu Q, Sun J, An ZY, Ding R, Li Y, Zhao XD (2020) Effect of Bacillus subtilis on corrosion behavior of 10MnNiCrCu steel in marine environment. Sci Rep 10(1):1–9. https://doi.org/10.1038/s41598-020-62809-y
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215(3):403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Kumar S, Tamura K, Nei M (1994) MEGA: molecular evolutionary genetics analysis software for microcomputers. Bioinformatics 10(2):189–191. https://doi.org/10.1093/bioinformatics/10.2.189
Deleage G, Roux B (1987) An algorithm for protein secondary structure prediction based on class prediction. Protein Eng Des Sel 1(4):289–294. https://doi.org/10.1093/protein/1.4.289
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105–132. https://doi.org/10.1016/0022-2836(82)90515-0
Vos P, Garrity G, Jones D, Krieg NR, Ludwig W, Rainey FA, Whitman WB (eds) (2011) Bergey’s manual of systematic bacteriology: volume 3: the Firmicutes, vol 3. Springer Science & Business Media, Berlin. https://doi.org/10.1007/b92997
Enguita FJ, Matias PM, Martins LIO, Plácido D, Henriques AO, Carrondo MA (2002) Spore-coat laccase CotA from Bacillus subtilis: crystallization and preliminary X-ray characterization by the MAD method. Acta Crystallogr D 58(9):1490–1493. https://doi.org/10.1107/S0907444902011575
Chen VB, Arendall WB, Headd JJ, Keedy DA, Immormino RM, Kapral GJ, Richardson DC (2010) MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr D 66(1):12–21. https://doi.org/10.1107/S0907444909042073
Kamali M, Khodaparast Z (2015) Review on recent developments on pulp and paper mill wastewater treatment. Ecotoxicol Environ Saf 114:326–342. https://doi.org/10.1016/j.ecoenv.2014.05.005
Sharma D, Chaudhary R, Kaur J, Arya SK (2020) Greener approach for pulp and paper industry by xylanase and laccase. Biocatal Agric Biotechnol 25:101604. https://doi.org/10.1016/j.bcab.2020.101604
Singh G, Arya SK (2019) Utility of laccase in pulp and paper industry: a progressive step towards the green technology. Int J Biol Macromol 134:1070–1084. https://doi.org/10.1016/j.ijbiomac.2019.05.168
Janusz G, Pawlik A, Świderska-Burek U, Polak J, Sulej J, Jarosz-Wilkołazka A, Paszczyński A (2020) Laccase properties, physiological functions, and evolution. Int J Mol Sci 21(3):966. https://doi.org/10.3390/ijms21030966
Arregui L, Ayala M, Gómez-Gil X, Gutiérrez-Soto G, Hernández-Luna CE, de Los H, Santos M, Levin L, Rojo-Domínguez A, Romero-Martínez D, Saparrat MC, Trujillo-Roldán MA, Valdez-Cruz NA (2019) Laccases: structure, function, and potential application in water bioremediation. Microb Cell Fact 18(1):1–33. https://doi.org/10.1186/s12934-019-1248-0
Martins LO, Soares CM, Pereira MM, Teixeira M, Costa T, Jones GH, Henriques AO (2002) Molecular and biochemical characterization of a highly stable bacterial laccase that occurs as a structural component of the Bacillus subtilis endospore coat. J Biol Chem 277(21):18849–18859. https://doi.org/10.1074/jbc.M200827200
Trubitsina LI, Tishchenko SV, Gabdulkhakov AG, Lisov AV, Zakharova MV, Leontievsky AA (2015) Structural and functional characterization of two-domain laccase from Streptomyces viridochromogenes. Biochimie 112:151–159. https://doi.org/10.1016/j.biochi.2015.03.005
Singh G, Singh S, Kaur K, Arya SK, Sharma P (2019) Thermo and halo tolerant laccase from Bacillus sp. SS4: evaluation for its industrial usefulness. J Gen Appl Microbiol 65(1):26–33. https://doi.org/10.2323/jgam.2018.04.002
Dubé E, Shareck F, Hurtubise Y, Daneault C, Beauregard M (2008) Homologous cloning, expression, and characterisation of a laccase from Streptomyces coelicolor and enzymatic decolourisation of an indigo dye. Appl Microbiol Biotechnol 79:597–603. https://doi.org/10.1007/s00253-008-1475-5
Brander S, Mikkelsen JD, Kepp KP (2014) Characterization of an alkali- and halide-resistant laccase expressed in E. coli: CotA from Bacillus clausii. PLoS ONE 9(6):e99402. https://doi.org/10.1371/journal.pone.0099402
Garg G, Dhiman SS, Mahajan R, Kaur A, Sharma J (2011) Bleach-boosting effect of crude xylanase from Bacillus stearothermophilus SDX on wheat straw pulp. New Biotechnol 28(1):58–64. https://doi.org/10.1016/j.nbt.2010.07.020
Mehandia S, Sharma SC, Arya SK (2020) Isolation and characterization of an alkali and thermostable laccase from a novel Alcaligenes faecalis and its application in decolorization of synthetic dyes. Biotechnol Rep 25:e00413. https://doi.org/10.1016/j.btre.2019.e00413
Dhiman SS, Garg G, Mahajan R, Garg N, Sharma J (2009) ‘Single lay out’ and ‘mixed lay out’ enzymatic processes for bio-bleaching of kraft pulp. Bioresour Technol 100(20):4736–4741. https://doi.org/10.1016/j.biortech.2009.04.059
Kapoor M, Kapoor RK, Kuhad RC (2007) Differential and synergistic effects of xylanase and laccase mediator system (LMS) in bleaching of soda and waste pulps. J Appl Microbiol 103(2):305–317. https://doi.org/10.1111/j.1365-2672.2006.03251.x
Sharma A, Thakur VV, Shrivastava A, Jain RK, Mathur RM, Gupta R, Kuhad RC (2014) Xylanase and laccase based enzymatic kraft pulp bleaching reduces adsorbable organic halogen (AOX) in bleach effluents: a pilot scale study. Bioresour Technol 169:96–102. https://doi.org/10.1016/j.biortech.2014.06.066
Qiu F, Zhang X, Liu L, Sun L, Schumann P, Song W (2009) Bacillus beijingensis sp. nov. and Bacillus ginsengi sp. nov., isolated from ginseng root. Int J Syst Evol Microbiol 59(4):729–734. https://doi.org/10.1099/ijs.0.65861-0
Verma P, Pandey PK, Gupta AK, Seong CN, Park SC, Choe HN, Baik KS, Patole MS, Shouche YS (2012) Reclassification of Bacillus beijingensis Qiu et al. 2009 and Bacillus ginsengi Qiu et al. 2009 as Bhargavaea beijingensis comb. nov. and Bhargavaea ginsengi comb. nov. and emended description of the genus Bhargavaea. Int J Syst Evol Microbiol 62(Pt_10):2495–2504. https://doi.org/10.1099/ijs.0.034850-0
Shivaji S, Ara S, Begum Z, Ruth M, Singh A, Kumar Pinnaka A (2013) Draft genome sequence of Bhargavaea cecembensis strain DSE10T, isolated from a deep-sea sediment sample collected at a depth of 5,904 m from the Chagos-Laccadive ridge system in the Indian Ocean. Genome Announc 1(3):e00346-e413. https://doi.org/10.1128/genomeA.00346-13
Tian FH, Fan DY, Zhang C, Jia CW, Gao W, Li Y, Li CT (2018) Bhargavaea changchunensis sp. nov. isolated from soil in China. Arch Microbiol 200(10):1465–1470. https://doi.org/10.1007/s00203-018-1563-6
Janusz G, Pawlik A, Swiderska-Burek U, Polak J, Sulej J, Jarosz-Wilkołazka A, Paszczynski A (2020) Laccase properties, physiological functions, and evolution. Int J Mol Sci 21(3):966. https://doi.org/10.3390/ijms21030966
Machczynski MC, Vijgenboom E, Samyn B, Canters GW (2004) Characterization of SLAC: a small laccase from Streptomyces coelicolor with unprecedented activity. Protein Sci 13(9):2388–2397. https://doi.org/10.1110/ps.04759104
Gunne M, Höppner A, Hagedoorn PL, Urlacher VB (2014) Structural and redox properties of the small laccase S sl1 from Streptomyces sviceus. FEBS J 281(18):4307–4318. https://doi.org/10.1111/febs.12755
Acknowledgements
Not applicable.
Funding
Not applicable.
Author information
Authors and Affiliations
Contributions
SC and SP designed the methodology and conceptualized the experiments, analyzed the data, and interpreted the results. SC and RP performed the investigation, analysis, and validation. SC, RP, MM, and SP were involved in editing. AV provided the laboratory to perform the experiments. All the authors reviewed and approved the manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that there is no conflict of interest.
Ethical Approval
Not applicable.
Consent to Participate
We Dr. Shalini Porwal and Dr. Ram Prasad give the consent to participate in Current Microbiology.
Consent for Publication
We Dr. Shalini Porwal and Dr. Ram Prasad give the consent to publish in Current Microbiology.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chaudhary, S., Varma, A., Mandal, M. et al. Isolation and Characterization of a Novel Laccase-Producing Bacteria Bhargavaea beijingensis from Paper and Pulp Effluent-Treated Soil Using In Silico Approaches. Curr Microbiol 80, 241 (2023). https://doi.org/10.1007/s00284-023-03346-5
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00284-023-03346-5