Abstract
Background
As a complex vascular malformation, fibro-adipose vascular anomaly was first proposed in 2014. Its overlap with other vascular malformations regarding imaging and clinical features often leads to misdiagnosis and improper management.
Objective
To construct a radiomics-based machine learning model to help radiologists differentiate fibro-adipose vascular anomaly from common venous malformations.
Materials and methods
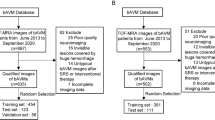
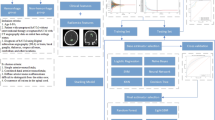
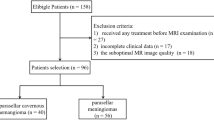
We retrospectively analyzed 178 children, adolescents and young adults with vascular malformations (41 fibro-adipose vascular anomaly and 137 common vascular malformation cases) who underwent MRI before surgery between May 2012 to January 2021. We extracted radiomics features from T1-weighted images and fat-saturated (FS) T2-weighted images and further selected features through least absolute shrinkage and selection operator (LASSO) and Boruta methods. We established eight weighted logistic regression classification models based on various combinations of feature-selection strategies (LASSO or Boruta) and sequence types (single- or multi-sequence). Finally, we evaluated the performance of each model by the mean area under the receiver operating characteristics curve (ROC-AUC), sensitivity and specificity in 10 runs of repeated k-fold (k = 10) cross-validation.
Results
Two multi-sequence models based on axial FS T2-W, coronal FS T2-W and axial T1-W images showed promising performance. The LASSO-based multi-sequence model achieved an AUC of 97%±3.8, a sensitivity of 94%±12.4 and a specificity of 89%±9.0. The Boruta-based multi-sequence model achieved an AUC of 97%±3.7, a sensitivity of 95%±10.5 and a specificity of 87%±9.0.
Conclusion
The radiomics-based machine learning model can provide a promising tool to help distinguish fibro-adipose vascular anomaly from common venous malformations.





Similar content being viewed by others
References
Alomari AI, Spencer SA, Arnold RW et al (2014) Fibro-adipose vascular anomaly: clinical–radiologic–pathologic features of a newly delineated disorder of the extremity. J Pediatr Orthop 34:109–117
International Society for the Study of Vascular Anomalies (2018) ISSVA classification of vascular anomalies. issva.org/classification. Accessed 1 Jan 2022
Amarneh M, Shaikh R (2020) Clinical and imaging features in fibro-adipose vascular anomaly (FAVA). Pediatr Radiol 50:380–387
Wassef M, Blei F, Adams D et al (2015) Vascular anomalies classification: recommendations from the International Society for the Study of Vascular Anomalies. Pediatrics 136:e203–e214
Vogel SA, Hess CP, Dowd CF et al (2013) Early versus later presentations of venous malformations: where and why? Pediatr Dermatol 30:534–540
Luks VL, Kamitaki N, Vivero MP et al (2015) Lymphatic and other vascular malformative/overgrowth disorders are caused by somatic mutations in PIK3CA. J Pediatr 166:1048–1054
Legiehn GM (2019) Sclerotherapy with adjunctive stasis of efflux (STASE) in venous malformations: techniques and strategies. Tech Vasc Interv Radiol 22:100630
Lipede C, Nikkhah D, Ashton R et al (2021) Management of fibro-adipose vascular anomalies (FAVA) in paediatric practice. JPRAS Open 29:71–81
Hori Y, Hirose K, Aramaki-Hattori N et al (2020) Fibro-adipose vascular anomaly (FAVA): three case reports with an emphasis on the mammalian target of rapamycin (mTOR) pathway. Diagn Pathol 15:98
Erickson J, McAuliffe W, Blennerhassett L, Halbert A (2017) Fibroadipose vascular anomaly treated with sirolimus: successful outcome in two patients. Pediatr Dermatol 34:e317–e320
Shaikh R, Alomari AI, Kerr CL et al (2016) Cryoablation in fibro-adipose vascular anomaly (FAVA): a minimally invasive treatment option. Pediatr Radiol 46:1179–1186
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577
Liu Z, Wang S, Dong D et al (2019) The applications of radiomics in precision diagnosis and treatment of oncology: opportunities and challenges. Theranostics 9:1303–1322
Kim J-H (2009) Estimating classification error rate: repeated cross-validation, repeated hold-out and bootstrap. Comput Stat Data Anal 53:3735–3745
Varma S, Simon R (2006) Bias in error estimation when using cross-validation for model selection. BMC Bioinformatics 7:91
Tustison NJ, Avants BB, Cook PA et al (2010) N4ITK: improved N3 bias correction. IEEE Trans Med Imaging 29:1310–1320
Zwanenburg A, Leger S, Vallières M, Löck S (2016) Image biomarker standardisation initiative. arXiv:1612.07003
Tibshirani R (2011) Regression shrinkage and selection via the lasso: a retrospective. J R Stat Soc Series B Stat Methodol 73:267–288
Kursa MB, Rudnicki WR (2010) Feature selection with Boruta package. J Stat Softw 36:1–13
Batista GEAPA, Prati RC, Monard MC (2004) A study of the behavior of several methods for balancing machine learning training data. ACM SIGKDD Explorations Newsl 6:20–29
Corani G, Benavoli A (2015) A Bayesian approach for comparing cross-validated algorithms on multiple data sets. Mach Learn 100:285–304
Benavoli A, Corani G, Demsar J, Zaffalon M (2017) Time for a change: a tutorial for comparing multiple classifiers through Bayesian analysis. J Mach Learn Res 18:2653–2688
Kruschke J, Liddell T (2015) The Bayesian new statistics: two historical trends converge. SSRN. https://doi.org/10.2139/ssrn.2606016. Accessed 15 Aug 2022
Nadeau C, Bengio Y (2003) Inference for the generalization error. Mach Learn 52:239–281
Youden WJ (1950) Index for rating diagnostic tests. Cancer 3:32–35
Lee BB, Baumgartner I, Berlien P et al (2015) Diagnosis and treatment of venous malformations. Consensus document of the International Union of Phlebology (IUP): updated 2013. Int Angiol 34:97–149
Hage AN, Chick JFB, Srinivasa RN et al (2018) Treatment of venous malformations: the data, where we are, and how it is done. Tech Vasc Interv Radiol 21:45–54
Rabe E, Breu FX, Cavezzi A et al (2014) European guidelines for sclerotherapy in chronic venous disorders. Phlebology 29:338–354
Armato SG 3rd, McLennan G, Bidaut L et al (2011) The Lung Image Database Consortium (LIDC) and Image Database Resource Initiative (IDRI): a completed reference database of lung nodules on CT scans. Med Phys 38:915–931
Kumar V, Gu Y, Basu S et al (2012) Radiomics: the process and the challenges. Magn Reson Imaging 30:1234–1248
Chicklore S, Goh V, Siddique M et al (2013) Quantifying tumour heterogeneity in 18F-FDG PET/CT imaging by texture analysis. Eur J Nucl Med Mol Imaging 40:133–140
Parekh V, Jacobs MA (2016) Radiomics: a new application from established techniques. Expert Rev Precis Med Drug Dev 1:207–226
Haralick RM, Shanmugam K, Dinstein I (1973) Textural features for image classification. IEEE Trans Syst Man Cybernet Syst SMC–3:610–621
Hein KD, Mulliken JB, Kozakewich H et al (2002) Venous malformations of skeletal muscle. Plast Reconstr Surg 110:1625
Bewick V, Cheek L, Ball JJ (2005) Statistics review 14: logistic regression. Crit Care 9:112–118
Tomek I (1976) Two modifications of CNN. IEEE Trans Syst Man Cybernet Syst SMC–6:769–772
Chawla NV, Bowyer KW, Hall LO, Kegelmeyer WP (2002) SMOTE: synthetic minority over-sampling technique. J Artif Int Res 16:321–357
Funding
This study has received funding from the National Key R&D Program of China (2017YFE0103600), the National Natural Science Foundation of China (81720108021) and the Henan provincial science and technology research projects (SB201901070 and 212102310689).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflicts of interest
None
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 3.34 MB)
Rights and permissions
Springer Nature or its licensor holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Dong, J., Gong, Y., Liu, Q. et al. Radiomics-based machine learning approach in differentiating fibro-adipose vascular anomaly from venous malformation. Pediatr Radiol 53, 404–414 (2023). https://doi.org/10.1007/s00247-022-05520-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00247-022-05520-6