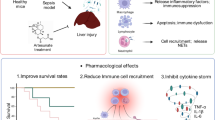
Abstract
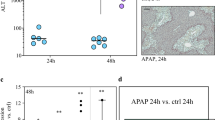
The role of non-parenchymal cells (NPCs) in the early phase of acetaminophen (APAP)-induced liver injury (AILI) remains unclear. Therefore, single-cell sequencing (scRNA-seq) was performed to explore the heterogeneity and immune network of NPCs in the livers of mice with AILI. Mice were challenged with saline, 300 mg/kg APAP, or 750 mg/kg APAP (n = 3 for each group). After 3 h, the liver samples were collected, digested, and subjected to scRNA-seq. Immunohistochemistry and immunofluorescence were performed to confirm the expression of Makorin ring finger protein 1 (Mkrn1). We identified 14 distinct cell subtypes among the 120,599 cells. A variety of NPCs were involved, even in the early stages of AILI, indicating highly heterogeneous transcriptome dynamics. Cholangiocyte cluster 3, which had high deleted in malignant brain tumors 1 (Dmbt1) expression, was found to perform drug metabolism and detoxification functions. Liver sinusoidal endothelial cells exhibited fenestrae loss and angiogenesis. Macrophage cluster 1 displayed a M1 polarization phenotype, whereas cluster 3 tended to exhibit M2 polarization. Kupffer cells (KCs) exhibited pro-inflammatory effects due to the high expression of Cxcl2. qRT-PCR and western blotting verified that the LIFR-OSM axis might promote the activation of MAPK signaling pathway in RAW264.7 macrophages. Mkrn1 was highly expressed in the liver macrophages of AILI mice and AILI patients. Interaction patterns between macrophages/KCs and other NPCs were complex and diverse. NPCs were highly heterogeneous and were involved in the immune network during the early phase of AILI. In addition, we propose that Mkrn1 may serve as a potential biomarker of AILI.








Similar content being viewed by others
Data availability
Data and materials used and/or analyzed during the current study are available from the corresponding authors on reasonable request.
Abbreviations
- NPCs:
-
Non-parenchymal cells
- APAP:
-
N-acetyl-p-aminophenol, acetaminophen
- AILI:
-
Acetaminophen-induced liver injury
- scRNA-seq:
-
Single-cell RNA sequencing
- Mkrn1:
-
Makorin ring finger protein 1
- Dmbt1:
-
Deleted in malignant brain tumor 1
- KCs:
-
Kupffer cells
- DILI:
-
Drug-induced liver injury
- NAPQI:
-
N-acetyl-p-benzoquinone imine
- LSECs:
-
Liver sinusoidal endothelial cells
- HSCs:
-
Hepatic stellate cells
- ALT:
-
Alanine aminotransferase
- AST:
-
Aspartate aminotransferase
- DAMPs:
-
Damage-associated molecular patterns
- CCR2:
-
Chemokine (C–C motif) receptor 2
- APCs:
-
Antigen-presenting cells
- NASH:
-
Non-alcoholic steatohepatitis
- UMI:
-
Unique molecular identifiers
- DEGs:
-
Differentially expressed genes
- GO:
-
Gene ontology
- KEGG:
-
Kyoto encyclopedia of genes and genomes
- HILI:
-
Herbal-induced liver injury
- NAFLD:
-
Non-alcoholic fatty liver disease
- CHB:
-
Chronic hepatitis B
- PBC:
-
Primary biliary cholangitis
- OSM:
-
Oncostatin M
- TUNEL:
-
Terminal dUTP nick‑end labeling
- ECs:
-
Endothelial cells
- pDCs:
-
Plasmacytoid dendritic cells
- Mps:
-
Mononuclear phagocytes
- SMCs:
-
Smooth muscle cells
- HepSCs:
-
Hepatic stellate cells
- UMAP:
-
Uniform manifold approximation and projection
- LIFR:
-
Leukemia inhibitory factor receptor
- MAPK:
-
Mitogen-activated protein kinase
- Spp1:
-
Secreted phosphoprotein 1
- OPN:
-
Osteopontin; VEGF, vascular endothelial growth factor
- CDKN1A:
-
Cyclin-dependent kinase inhibitor 1A
- LDH:
-
Lactate dehydrogenase
- Epcs:
-
Epithelial progenitors
- GSVA:
-
Gene set variation analysis
- ALP:
-
Alkaline phosphatase
- GGT:
-
γ-Glutamyl transferase
- TBIL:
-
Total bilirubin
References
Aithal GP, Grove JI (2015) Genome-wide association studies in drug-induced liver injury: step change in understanding the pathogenesis. Semin Liver Dis 35(4):421–431. https://doi.org/10.1055/s-0035-1567829
Aizarani N, Saviano A, Sagar et al (2019) A human liver cell atlas reveals heterogeneity and epithelial progenitors. Nature 572(7768):199–204. https://doi.org/10.1038/s41586-019-1373-2
Alvaro D, Mancino MG, Glaser S et al (2007) Proliferating cholangiocytes: a neuroendocrine compartment in the diseased liver. Gastroenterology 132(1):415–431. https://doi.org/10.1053/j.gastro.2006.07.023
Andrade RJ, Chalasani N, Björnsson ES et al (2019) Drug-induced liver injury. Nat Rev Dis Primers 5(1):58. https://doi.org/10.1038/s41572-019-0105-0
Antoniades CG, Quaglia A, Taams LS et al (2012) Source and characterization of hepatic macrophages in acetaminophen-induced acute liver failure in humans. Hepatology 56(2):735–746. https://doi.org/10.1002/hep.25657
Ben-Moshe S, Veg T, Manco R et al (2022) The spatiotemporal program of zonal liver regeneration following acute injury. Cell Stem Cell 29(6):973–989. https://doi.org/10.1016/j.stem.2022.04.008
Black AR, Black JD, Azizkhan-Clifford J (2001) Sp1 and krüppel-like factor family of transcription factors in cell growth regulation and cancer. J Cell Physiol 188(2):143–160. https://doi.org/10.1002/jcp.1111
Butler A, Hoffman P, Smibert P, Papalexi E, Satija R (2018) Integrating single-cell transcriptomic data across different conditions, technologies, and species. Nat Biotechnol 36(5):411–420. https://doi.org/10.1038/nbt.4096
Church RJ, Kullak-Ublick GA, Aubrecht J et al (2019) Candidate biomarkers for the diagnosis and prognosis of drug-induced liver injury: an international collaborative effort. Hepatology 69(2):760–773. https://doi.org/10.1002/hep.29802
Dura B, Choi J-Y, Zhang K et al (2019) scFTD-seq: freeze-thaw lysis based, portable approach toward highly distributed single-cell 3’ mRNA profiling. Nucleic Acids Res 47(3):e16. https://doi.org/10.1093/nar/gky1173
Friedman SL (2008) Hepatic stellate cells: protean, multifunctional, and enigmatic cells of the liver. Physiol Rev 88(1):125–172. https://doi.org/10.1152/physrev.00013.2007
Ghanem CI, Pérez MJ, Manautou JE, Mottino AD (2016) Acetaminophen from liver to brain: new insights into drug pharmacological action and toxicity. Pharmacol Res 109:119–131. https://doi.org/10.1016/j.phrs.2016.02.020
Holt MP, Yin H, Ju C (2010) Exacerbation of acetaminophen-induced disturbances of liver sinusoidal endothelial cells in the absence of Kupffer cells in mice. Toxicol Lett 194(1–2):34–41. https://doi.org/10.1016/j.toxlet.2010.01.020
Hoofnagle JH, Björnsson ES (2019) Drug-induced liver injury - types and phenotypes. N Engl J Med 381(3):264–273. https://doi.org/10.1056/NEJMra1816149
Hwang B, Lee JH, Bang D (2018) Single-cell RNA sequencing technologies and bioinformatics pipelines. Exp Mol Med 50(8):1–4. https://doi.org/10.1038/s12276-018-0071-8
Kim JH, Park S-M, Kang MR et al (2005) Ubiquitin ligase MKRN1 modulates telomere length homeostasis through a proteolysis of hTERT. Genes Dev 19(7):776–781. https://doi.org/10.1101/gad.1289405
Krenkel O, Tacke F (2017) Liver macrophages in tissue homeostasis and disease. Nat Rev Immunol 17(5):306–321. https://doi.org/10.1038/nri.2017.11
Lantieri F, Bachetti T (2022) OSM/OSMR and interleukin 6 family cytokines in physiological and pathological condition. Int J Mol Sci 23(19):11096. https://doi.org/10.3390/ijms231911096
Lee E-W, Lee M-S, Camus S et al (2009) Differential regulation of p53 and p21 by MKRN1 E3 ligase controls cell cycle arrest and apoptosis. EMBO J 28(14):2100–2113. https://doi.org/10.1038/emboj.2009.164
Lee M-S, Han H-J, Han SY et al (2018) Loss of the E3 ubiquitin ligase MKRN1 represses diet-induced metabolic syndrome through AMPK activation. Nat Commun 9(1):3404. https://doi.org/10.1038/s41467-018-05721-4
Lei X, Xu Q, Li C et al (2022) Egr1 confers protection against acetaminophen-induced hepatotoxicity via transcriptional upregulating of Acaa2. Int J Biol Sci 18(9):3800–3817. https://doi.org/10.7150/ijbs.71781
Li L, Zeng Z (2020) Live imaging of innate and adaptive immune responses in the liver. Front Immunol 11:564768. https://doi.org/10.3389/fimmu.2020.564768
Li C, Ming Y, Hong W et al (2018) Comparison of hepatic transcriptome profiling between acute liver injury and acute liver failure induced by acetaminophen in mice. Toxicol Lett 283:69–76. https://doi.org/10.1016/j.toxlet.2017.11.020
Li X, Tang J, Mao Y (2022) Incidence and risk factors of drug-induced liver injury. Liver Int 42(9):1999–2014. https://doi.org/10.1111/liv.15262
Liao Y, Smyth GK, Shi W (2014) FeatureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics 30(7):923–930. https://doi.org/10.1093/bioinformatics/btt656
Liu Z-X, Govindarajan S, Kaplowitz N (2004) Innate immune system plays a critical role in determining the progression and severity of acetaminophen hepatotoxicity. Gastroenterology 127(6):1760–1774. https://doi.org/10.1053/j.gastro.2004.08.053
MacParland SA, Liu JC, Ma X-Z et al (2018) Single cell RNA sequencing of human liver reveals distinct intrahepatic macrophage populations. Nat Commun 9(1):4383. https://doi.org/10.1038/s41467-018-06318-7
Marrone G, Maeso-Díaz R, García-Cardena G et al (2015) KLF2 exerts antifibrotic and vasoprotective effects in cirrhotic rat livers: behind the molecular mechanisms of statins. Gut 64(9):1434–1443. https://doi.org/10.1136/gutjnl-2014-308338
Massalha H, Bahar Halpern K, Abu-Gazala S et al (2020) A single cell atlas of the human liver tumor microenvironment. Mol Syst Biol 16(12):9682. https://doi.org/10.15252/msb.20209682
Mossanen JC, Krenkel O, Ergen C et al (2016) Chemokine (C-C motif) receptor 2-positive monocytes aggravate the early phase of acetaminophen-induced acute liver injury. Hepatology 64(5):1667–1682. https://doi.org/10.1002/hep.28682
Nelson DR, Zeldin DC, Hoffman SMG et al (2004) Comparison of cytochrome P450 (CYP) genes from the mouse and human genomes, including nomenclature recommendations for genes, pseudogenes and alternative-splice variants. Pharmacogenetics 14(1):1–18. https://doi.org/10.1097/00008571-200401000-00001
Niu N, Xu S, Xu Y, Little PJ, Jin Z-G (2019) Targeting mechanosensitive transcription factors in atherosclerosis. Trends Pharmacol Sci 40(4):253–266. https://doi.org/10.1016/j.tips.2019.02.004
Poisson J, Lemoinne S, Boulanger C et al (2017) Liver sinusoidal endothelial cells: physiology and role in liver diseases. J Hepatol 66(1):212–227. https://doi.org/10.1016/j.jhep.2016.07.009
Qiu X, Hill A, Packer J et al (2017) Single-cell mRNA quantification and differential analysis with census. Nat Methods 14(3):309–315. https://doi.org/10.1038/nmeth.4150
Ramachandran P, Dobie R, Wilson-Kanamori JR et al (2019) Resolving the fibrotic niche of human liver cirrhosis at single-cell level. Nature 575(7783):512–518. https://doi.org/10.1038/s41586-019-1631-3
Satija R, Farrell JA, Gennert D, Schier AF, Regev A (2015) Spatial reconstruction of single-cell gene expression data. Nat Biotechnol 33(5):495–502. https://doi.org/10.1038/nbt.3192
Shen T, Liu Y, Shang J et al (2019) Incidence and etiology of drug-induced liver injury in Mainland China. Gastroenterology. https://doi.org/10.1053/j.gastro.2019.02.002
Starkey Lewis P, Campana L, Aleksieva N et al (2020) Alternatively activated macrophages promote resolution of necrosis following acute liver injury. J Hepatol 73(2):349–360. https://doi.org/10.1016/j.jhep.2020.02.031
Stephens C, Andrade RJ, Lucena MI (2014) Mechanisms of drug-induced liver injury. Curr Opin Allergy Clin Immunol 14(4):286–292. https://doi.org/10.1097/ACI.0000000000000070
Terkelsen MK, Bendixen SM, Hansen D et al (2020) Transcriptional dynamics of hepatic sinusoid-associated cells after liver injury. Hepatology 72(6):2119–2133. https://doi.org/10.1002/hep.31215
Warkentin TE (2022) Platelet-activating anti-PF4 disorders: an overview. Semin Hematol 59(2):59–71. https://doi.org/10.1053/j.seminhematol.2022.02.005
Wei J, Marisetty A, Schrand B et al (2019) Osteopontin mediates glioblastoma-associated macrophage infiltration and is a potential therapeutic target. J Clin Invest 129(1):137–149. https://doi.org/10.1172/JCI121266
Xie Y, McGill MR, Dorko K et al (2014) Mechanisms of acetaminophen-induced cell death in primary human hepatocytes. Toxicol Appl Pharmacol 279(3):266–274. https://doi.org/10.1016/j.taap.2014.05.010
Xiong X, Kuang H, Ansari S et al (2019) Landscape of intercellular crosstalk in healthy and NASH liver revealed by single-cell secretome gene analysis. Mol Cell. https://doi.org/10.1016/j.molcel.2019.07.028
Yan M, Huo Y, Yin S, Hu H (2018) Mechanisms of acetaminophen-induced liver injury and its implications for therapeutic interventions. Redox Biol 17:274–283. https://doi.org/10.1016/j.redox.2018.04.019
Yang W, He H, Wang T et al (2021) Single-cell transcriptomic analysis reveals a hepatic stellate cell-activation roadmap and myofibroblast origin during liver fibrosis in mice. Hepatology 74(5):2774–2790. https://doi.org/10.1002/hep.31987
Yang T, Wang H, Wang X, Li J, Jiang L (2022) The dual role of innate immune response in acetaminophen-induced liver injury. Biology (Basel) 11(7):1057. https://doi.org/10.3390/biology11071057
Yokomori H, Oda M, Ogi M et al (2003) Endothelin-1 suppresses plasma membrane Ca++-ATPase, concomitant with contraction of hepatic sinusoidal endothelial fenestrae. Am J Pathol 162(2):557–566. https://doi.org/10.1016/S0002-9440(10)63849-7
Yu G, Wang L-G, Han Y, He Q-Y (2012) clusterProfiler: an R package for comparing biological themes among gene clusters. OMICS 16(5):284–287. https://doi.org/10.1089/omi.2011.0118
Yuan L, Kaplowitz N (2013) Mechanisms of drug-induced liver injury. Clin Liver Dis 17(4):507–518. https://doi.org/10.1016/j.cld.2013.07.002
Zhang Y, Liu T, Hu X et al (2021) Cell Call: integrating paired ligand-receptor and transcription factor activities for cell-cell communication. Nucleic Acids Res 49(15):8520–8534. https://doi.org/10.1093/nar/gkab638
Acknowledgements
This work was supported by the National Natural Science Foundation of China (NSFC 81970513, NSFC 82270619, and NSFC 32171175), National Key R&D Program of China (2022YFC3502101), Clinical Research Innovation and Development Fund of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (PY120-05).
Funding
This work was supported by the National Natural Science Foundation of China (NSFC 81970513, NSFC 82270619, and NSFC 32171175), the National Key R&D Program of China (2022YFC3502101), and the Clinical Research Innovation and Development Fund of Renji Hospital, School of Medicine, Shanghai Jiao Tong University (PY120-05).
Author information
Authors and Affiliations
Contributions
YXL, YZ, JL, and BXL performed the experiment and wrote and drafted the manuscript. HXL, YJ, TYZ, and FYZ retrieved the data and performed the statistical analysis. MXK, FX, WZ, and YXC performed the bioinformatics analysis. TJT, BXL, and MYM acquired funding. BXL and MYM revised the manuscript. All authors read and approved the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Ethical approval
All animal experiments were approved by the Animal Ethics Committee of Shanghai Jiao Tong University Renji Hospital. Animal research was conducted in accordance with international guidelines. The clinical research was approved by the Ethics Committee of Shanghai Jiaotong University Renji Hospital in accordance with the ethical guidelines of the 1975 Declaration of Helsinki.
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Li, X., Zhi, Y., Li, J. et al. Single-cell RNA sequencing to reveal non-parenchymal cell heterogeneity and immune network of acetaminophen-induced liver injury in mice. Arch Toxicol 97, 1979–1995 (2023). https://doi.org/10.1007/s00204-023-03513-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00204-023-03513-4