Abstract
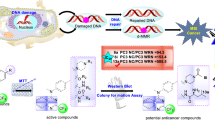
A series of novel 1,2,3-triazole-benzoxazepine hybrid molecules were synthesized and evaluated for their anticancer properties against four cancer cell lines (Caco-2, PC3, MCF-7, and HL60). Most of the synthesized compounds exhibited moderate to good cytotoxicity against tested cancer cell lines. Three of the prepared compounds, namely, 2, 3, and 4, showed excellent anticancer properties with micromolar IC50 values ranging from 6 to 18 μM. In silico pharmacophore profiling followed by in-vitro enzyme assays showed 2, 3, and 4 to have micromolar inhibitory IC50 values against the oncogenic kinase FLT3. These compounds represent new inhibitory leads of novel chemotypes against FLT3.
Graphical Abstract









Similar content being viewed by others
References
Islam MS, Wang C, Zheng J, Paudyal N, Zhu Y, Sun H. The potential role tubeimosides in cancer prevention and treatment. Eur J Med Chem. 2019;162:109–21. https://doi.org/10.1016/j.ejmech.2018.11.001
Zhang J, Wang S, Ba Y, Xu Z. Tetrazole hybrids with potential anticancer activity. Eur J Med Chem. 2019;178:341–51. https://doi.org/10.1016/j.ejmech.2019.05.071
Shaveta, Mishra S, Singh P. Hybrids molecules: the privileged scaffolds for various pharmaceuticals. Eur J Med Chem. 2016;124:500–36. https://doi.org/10.1016/j.ejmech.2016.08.039
Hu YQ, Zhang S, Xu Z, Lv ZS, Liu ML, Feng LS. 4-Quinolone hybrids and their antibacterial activities. Eur J Med Chem. 2017;141:335–45. https://doi.org/10.1016/j.ejmech.2017.09.050
Zhang B. Comprehensive review on the anti-bacterial activity of 1,2,3- triazole hybrids. Eur J Med Chem. 2019;168:357–72. https://doi.org/10.1016/j.ejmech.2019.02.055
Kalaria PN, Karad SC, Raval DK. A review on diverse heterocyclic compounds as the privileged scaffolds in antimalarial drug discovery. Eur J Med Chem. 2018;158:917–36. https://doi.org/10.1016/j.ejmech.2018.08.040
Emami S, Ghobadi E, Saednia S, Hashemi SM. Current advances of triazole alcohols derived from fluconazole: design, in vitro and in silico studies. Eur J Med Chem. 2019;170:173–94. https://doi.org/10.1016/j.ejmech.2019.03.020
Kaoukabi H, Kabri Y, Curti C, Taourirte M, Rodríguez-Ubis JC, Snoeck R, et al. Dihydropyrimidinone/1,2,3-triazole hybrid molecules: synthesis and anti-varicella-zoster virus (VZV) evaluation. Eur J Med Chem. 2018;155:772–81. https://doi.org/10.1016/j.ejmech.2018.06.028
Keri RS, Patil SA, Budagumpi S, Nagaraja BM. Triazole, A promising antitubercular agent. Chem Biol Drug Des. 2015;86:410–23. https://doi.org/10.1111/cbdd.12527
Lal K, Yadav P. Recent advancements in 1,4-disubstituted 1H-1,2,3-triazoles as potential anticancer agents. Anti Cancer Agents Med Chem. 2018;18:21–37. https://doi.org/10.2174/1871520616666160811113531
Akhtar J, Khan AA, Ali Z, Haider R, Yar MS. Structure-activity relationship (SAR) study and design strategies of nitrogen-containing heterocyclic moieties for their anticancer activities. Eur J Med Chem. 2017;125:143–89. https://doi.org/10.1016/j.ejmech.2016.09.023
Senwar KR, Sharma P, Reddy TS, Jeengar MK, Nayak VL, Naidu VGM, et al. Spirooxindole-derived morpholine-fused-1,2,3-triazoles: Design, synthesis, cytotoxicity and apoptosis inducing studies. Eur J Med Chem. 2015;102:413–24. https://doi.org/10.1016/j.ejmech.2015.08.017
Wei G, Luan W, Wang S, Cui S, Li F, Liu Y, et al. A library of 1,2,3-triazole-substituted oleanolic acid derivatives as anticancer agents: design, synthesis, and biological evaluation. Org Biomol Chem. 2015;13:1507–14. https://doi.org/10.1039/C4OB01605J
Yadav P, Lal K, Kumar A, Guru SK, Jaglan S, Bhushan S. Green synthesis and anticancer potential of chalcone linked-1,2,3-triazoles. Eur J Med Chem. 2017;126:944–53. https://doi.org/10.1016/j.ejmech.2016.11.030
O’Neil IA, Murray CL, Hunter RC, Kalindjian SB, Jenkins TC. The synthesis of functionalized pyrrolo-[2, 1-c][1, 4]-benzodiazepines. Synlett. 1997;1:75–78. https://doi.org/10.1055/s-1997-693.
Liao Y, Venhuis BJ, Rodenhuis N, Timmerman W, Wikstro¨m H, Meier E, et al. New (sulfonyloxy)piperazinyldibenzazepines as potential atypical antipsychotics: chemistry and pharmacological evaluation. J Med Chem. 1999;42:2235–44. https://doi.org/10.1021/jm991005d
Goutham K, Kumar DA, Suresh S, Sridhar B, Narender R, Karunakar GV. Gold-catalyzed intramolecular cyclization of N-propargylic β-enaminones for the synthesis of 1,4-oxazepine derivatives. J Org Chem. 2015;80:11162–8. https://doi.org/10.1021/acs.joc.5b01733
McElligott AM, Maginn EN, Greene LM, McGuckin S, Hayat A, Browne PV, et al. The novel tubulin-targeting agent pyrrolo-1,5-benzoxazepine-15 induces apoptosis in poor prognostic subgroups of chronic lymphocytic leukemia. Cancer Res. 2009;69:8366–75. https://doi.org/10.1158/0008-5472.CAN-09-0131
Yin Y, Zhang YQ, Jin B, Shao S, Wu X, Sangani CB, et al. 6,7-Dihydrobenzo[f]benzo[4,5]imidazo[1,2-d][1,4]oxazepine derivatives as selective inhibitors of PI3Kα. Bioorg Med Chem. 2015;23:1231–40. https://doi.org/10.1016/j.bmc.2015.01.052
Abutayeh RF, Taha MO. Discovery of novel Flt3 inhibitory chemotypes through extensive ligand-based and new structure-based pharmacophore modelling methods. J Mol Graph Model. 2019;88:128–51. https://doi.org/10.1016/j.jmgm.2019.01.011
Kuntala N, Telu JR, Banothu V, Nallapati SB, Anireddy JS, Pal S. Novel benzoxepine-1,2,3-triazole hybrids: synthesis and pharmacological evaluation as potential antibacterial and anticancer agents. Med Chem Commun. 2015;6:1612–9. https://doi.org/10.1039/C5MD00224A
Ouahrouch A, Ighachane H, Taourirte M, Engels JW, Sedra MH, Lazrek HB. Benzimidazole-1,2,3-triazole hybrid molecules: synthesis and evaluation for antibacterial/antifungal activity. Arch Pharm Chem Life Sci. 2014;347:1–8. https://doi.org/10.1002/ardp.201400142
Sumangala V, Poojary B, Chidananda N, Fernandes J, Kumari NS. Synthesis and antimicrobial activity of 1,2,3-triazoles containing quinoline moiety. Arch Pharm Res. 2010;33:1911–8. https://doi.org/10.1007/s12272-010-1204-3
Rathwell K, Sperry J, Brimble MA. Synthesis of triazole analogues of the nanaomycin antibiotics using ‘click chemistry’. Tetrahedron. 2010;66:4002–9. https://doi.org/10.1016/j.tet.2010.04.048
Kumar A, Ahmad I, Chhikara BS, Tiwari R, Mandal D, Parang K. Synthesis of 3-phenylpyrazolopyrimidine-1,2,3-triazole conjugates and evaluation of their Src kinase inhibitory and anticancer activities. Bioorg Med Chem Lett. 2011;21:1342–6. https://doi.org/10.1016/j.bmcl.2011.01.047
Gill C, Jadhav G, Shaikh M, Kale R, Ghawalkar A, Nagargoje D, et al. Clubbed [1,2,3] triazoles by fluorine benzimidazole: A novel approach to H37Rv inhibitors as a potential treatment for tuberculosis. Bioorg Med Chem Lett. 2008;18:6244–7. https://doi.org/10.1016/j.bmcl.2008.09.096
Mizyed SA, Ashram M, Awwadi FF. A new and convenient synthetic method for 1,2,3,5,6,11bhexahydroimidazo[1,2-d][1,4]benzoxazepine and its derivatives. Arkivoc. 2011;x:277–86. https://doi.org/10.3998/ark.5550190.0012.a22
Ashram M, Awwadi FF. A new, simple and efficient method for the synthesis of tricyclic [1,3]oxazolo[3,2- d][1,4]benzoxazepine, [1,3]oxazino[3,2-d][1,4]benzoxazepine, pyrimido[1,2- d][1,4]benzoxazepine and their derivatives. Arch Org Chem. 2019;v:142–51. https://doi.org/10.24820/ark.5550190.p010.780.
Ashram M, Awwadi FF. A new and efficient synthesis of unsaturated benzoxazepines using sodium metabisulfite and potassium permanganate as oxidative reagent. Arkivoc. 2019;vi:239–51. https://doi.org/10.24820/ark.5550190.p011.061
Wang L, Sun Y. Efflux mechanism and pathway of verapamil pumping by human P-glycoprotein. Arch Biochem Biophys. 2020;696:108675 https://doi.org/10.1016/j.abb.2020.108675
Ali AAS, Khan D, Naqvi A, Al-Blewi FF, Rezki N, Aouad MR, Hagar M. Design, synthesis, molecular modeling, anticancer studies, and density functional theory calculations of 4-(1, 2, 4-Triazol-3-ylsulfanylmethyl)-1, 2, 3-triazole derivatives. ACS Omega. 2021;6:301–16. https://doi.org/10.1021/acsomega.0c04595.
Vanaparthi S, Bantu R, Jain N, Janardhan S, Nagarapu L. Synthesis and anti-proliferative activity of a novel 1,2,3-triazole tethered chalcone acetamide derivatives. Bioorg Med Chem Lett 2020;30:127304 https://doi.org/10.1016/j.bmcl.2020.127304
Macan M, Perin N, Jakopec S, Miloc M, Stojkovic MR, Kralj M, et al. Synthesis, antiproliferative activity and DNA/RNA-binding properties of mono- and bis-(1,2,3-triazolyl)-appended benzimidazo[1,2-a]quinoline derivatives. Eur J Med Chem. 2020;185:111845. 0.1016/j.ejmech.2019.111845
Hijjawi MS, Abutayeh RF, Taha MO. Structure-based discovery and bioactivity evaluation of novel aurora-A kinase inhibitors as anticancer agents via docking-based comparative intermolecular contacts analysis (dbCICA). Molecules. 2020;25:6003. https://doi.org/10.3390/molecules25246003.
Aboalhaija NH, Zihlif MA, Taha MO. Discovery of new selective cytotoxic agents against Bcl-2 expressing cancer cells using ligand-based modeling. Chem Biol Interact. 2016;250:12–26. https://doi.org/10.1016/j.cbi.2016.03.006
Alabed SJ, Khanfar M, Taha MO. Computer-aided discovery of new FGFR-1 inhibitors followed by in vitro validation. Future Med Chem. 2016;8:1841–69. https://doi.org/10.4155/fmc-2016-0056
Taha MO, Bustanji Y, Al-Ghussein MAS, Mohammad M, Zalloum H, Al-Masri IM, et al. Pharmacophore modeling, quantitative structure–activity relationship analysis, and in silico screening reveal potent glycogen synthase Kinase-3β inhibitory activities for cimetidine, hydroxychloroquine, and gemifloxacin. J Med Chem 2008;51:2062–77. https://doi.org/10.1021/jm7009765
Al-Sha’er MA, Taha MO. Elaborate ligand-based modeling reveal new nanomolar heat shock protein 90a inhibitors. J Chem Inf Model. 2010;50:1706–23. https://doi.org/10.1021/ci100222k
Tuffaha GO, Hatmal MM, Taha MO. Discovery of new JNK3 inhibitory chemotypes via QSAR-Guided selection of docking-based pharmacophores and comparison with other structure-based pharmacophore modeling methods. J Mol Graph Model. 2019;91:30–51. https://doi.org/10.1016/j.jmgm.2019.05.015
Khanfar MA, Taha MO. Elaborate ligand-based modeling coupled with multiple linear regression and k nearest neighbor QSAR analyses unveiled new nanomolar mTOR inhibitors. J Chem Inf Model. 2013;53:2587–612. https://doi.org/10.1021/ci4003798
Alassaf SA, Hijjawi MS, Abuhammad A, Taha MO. Structure-based discovery of new polo-like kinase 1 (PLK1) inhibitors as potential anticancer agents via docking-based comparative intermolecular contacts analysis (dbCICA). Med Chem Res. 2021;1:1747–66. https://doi.org/10.1007/s00044-021-02774-x.
Shahin R, AlQtaishat S, Taha MO. Elaborate ligand-based modeling reveal new submicromolar Rho kinase inhibitors. J Comput Aided Mol Des. 2012;26:249–66. https://doi.org/10.1007/s10822-011-9509-y
Al-Barghouthy E, Abuhammad A, Taha MO. QSAR-guided pharmacophore modeling and subsequent virtual screening identify novel TYK2 inhibitor. Med Chem Res. 2019;28:1368–87. https://doi.org/10.1007/s00044-019-02377-7
Al-Sha’er MA, Basheer HA, Taha MO. Discovery of new PKN2 inhibitory chemotypes via QSAR-guided selection of docking-based pharmacophores. Mol Divers. 2022;4:1–20. https://doi.org/10.1007/s11030-022-10434-4
Al-Tawil MF, Daoud S, Hatmal MM, Taha MO. Discovery of new Cdc2-like kinase 4 (CLK4) inhibitors via pharmacophore exploration combined with flexible docking-based ligand/receptor contact fingerprints and machine learning. RSC Adv. 2022;12:10686–10700. https://doi.org/10.1039/D2RA00136E
Mousa LA, Hatmal MM, Taha MO. Exploiting activity cliffs for building pharmacophore models and comparison with other pharmacophore generation methods: sphingosine kinase 1 as case study. J Comput Aided Mol Des. 2022;36:39–62. https://doi.org/10.1007/s10822-021-00435-0
Al-Sha’er MA, Al-aqtash RA, Taha MO. Discovery of new phosphoinositide 3-kinase delta (PI3Kδ) inhibitors via virtual screening using crystallography-derived pharmacophore modelling and QSAR analysis. Med Chem. 2019;15:588–601. https://doi.org/10.2174/1573406415666190222125333
Hatmal MM, Taha MO. Combining stochastic deformation/relaxation and intermolecular contacts analysis for extracting pharmacophores from ligand-peceptor complexes. J Chem Inf Model. 2018;58:879–93. https://doi.org/10.1021/acs.jcim.7b00708
Wu G, Robertson DH, Brooks CL 3rd, Vieth M. Detailed analysis of grid-based molecular docking: a case study of CDOCKER – A CHARMm based MD docking program. J Comput Chem. 2003;24:1549–62. https://doi.org/10.1002/jcc.10306
Nassar H, Abu-Dahab R, Taha MO. Inhibition of protein kinases by proton pump inhibitors: computational screening and in vitro evaluation. Med Chem Res. 2021;30:2266–76. https://doi.org/10.1007/s00044-021-02812-8
Tirado-Rives J, Jorgensen WL. Contribution of conformer focusing to the uncertainty in predicting free energies for protein− ligand binding. J Med Chem. 2006;49:5880–4. https://doi.org/10.1021/jm060763i
Ma H, Deacon S, Horiuchi K. The challenge of selecting protein kinase assays for lead discovery optimization. Expert Opin Drug Disco. 2008;3:607–62. https://doi.org/10.1517/17460441.3.6.607
Bardaweel S, Aljanabi R, Sabbah D, Sweidan K. Design, synthesis, and biological evaluation of novel MAO-A inhibitors targeting lung cancer. Molecules. 2022;27:2887 https://doi.org/10.3390/molecules27092887
Gucký T, Řezníčková E, Radošová Muchová T, Jorda R, Klejová Z, Malínková V, et al. Discovery of N2-(4-Amino-cyclohexyl)-9-cyclopentyl-N6-(4-morpholin-4-ylmethyl-phenyl)-9H-purine-2,6-diamine as a Potent FLT3 Kinase Inhibitor for Acute Myeloid Leukemia with FLT3 Mutations. J Med Chem. 2018;61:3855–69. https://doi.org/10.1021/acs.jmedchem.7b01529
Smith CC, Zhang DC, Lin KC, Lasater EA. Characterizing and overriding the structural mechanism of the quizartinib-resistant FLT3 “gatekeeper” F691L mutation with PLX3397. Cancer Disco. 2015;5:668–79. https://doi.org/10.1158/2159-8290.CD-15-0060
Acknowledgements
The Chemistry Department of the University of Jordan-Amman is gratefully acknowledged for measuring the NMR and HRMS spectra. This project was funded by the Deanship of Academic Research at the University of Jordan.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Ashram, M., Habashneh, A.Y., Bardaweel, S. et al. A Click Synthesis, Molecular Docking and Biological Evaluation of 1,2,3-triazoles-benzoxazepine hybrid as potential anticancer agents. Med Chem Res 32, 271–287 (2023). https://doi.org/10.1007/s00044-022-03001-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00044-022-03001-x