Abstract
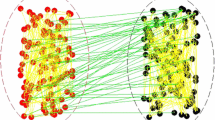
Human communities are organized in complex webs of contacts that may be represented by a graph or network. In this graph, vertices identify individuals and edges establish the existence of some type of relations between them. In real communities, the possible edges may be active or not for variable periods of time. These so-called temporal networks typically result from an endogenous social dynamics, usually coupled to the process under study taking place in the community. For instance, disease spreading may be affected by local information that makes individuals aware of the health status of their social contacts, allowing them to reconsider maintaining or not their social contacts. Here we investigate the impact of such a dynamical network structure on disease dynamics, where infection occurs along the edges of the network. To this end, we define an endogenous network dynamics coupled with disease spreading. We show that the effective infectiousness of a disease taking place along the edges of this temporal network depends on the population size, the number of infected individuals in the population and the capacity of healthy individuals to sever contacts with the infected, ultimately dictated by availability of information regarding each individual’s health status. Importantly, we also show how dynamical networks strongly decrease the average time required to eradicate a disease.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
13.1 Introduction
Understanding disease spreading and evolution involves overcoming a multitude of complex, multi-scale challenges of mathematical and biological nature [1, 2]. Traditionally, the contact process between an infected individual and the susceptible ones was assumed to affect equally any susceptible in a population (mean-field approximation, well-mixed population approximation) or, alternatively, all those susceptible living in the physical neighborhood of the infected individual (spatial transmission). During recent years, however, it has become clear that disease spreading [2,3,4,5] transcends geography: the contact process is no longer restricted to the immediate geographical neighbors, but exhibits the stereotypical small-world phenomenon [6,7,8,9], as testified by recent global pandemics (together with the impressive amount of research that has been carried out to investigate them) or, equally revealing, the dynamics associated with the spreading of computer viruses [5, 10,11,12,13,14,15,16,17,18,19,20,21,22,23]. Recent advances in the science of networks [3, 4, 19, 24, 25] also provided compelling evidence of the role that the networks of contacts between individuals or computers play in the dynamics of infectious diseases [4, 7]. In the majority of cases in which complex networks of disease spreading have been considered [9], they were taken to be a single, static entity. However, contact networks are intrinsically temporal entities and, in general, one expects the contact process to proceed along the lines of several networks simultaneously [11, 13,14,15,16, 18, 23, 24, 26,27,28,29,30,31,32,33,34,35,36]. In fact, modern societies have developed rapid means of information dissemination, both at local and at centralized levels, which one naturally expects to alter individuals’ response to vaccination policies, their behavior with respect to other individuals and their perception of likelihood and risk of infection [37]. In some cases one may even witness the adoption of centralized measures, such as travel restrictions [38, 39] or the imposition of quarantine spanning parts of the population [40], which may induce abrupt dynamical features onto the structure of the contact networks. In other cases, social media can play a determinant role in defining the contact network, providing crucial information on the dynamical patterns of disease spreading [41]. Furthermore, the knowledge an individual has (based on local and/or social media information) about the health status of acquaintances, partners, relatives, etc., combined with individual preventive strategies [42,43,44,45,46,47,48,49,50] (such as condoms, vaccination, the use of face masks or prophylactic drugs, avoidance of visiting specific web-pages, staying away from public places, etc.), also leads to changes in the structure and shape of the contact networks that naturally acquire a temporal dimension that one should not overlook.
Naturally, the temporal dimension and multitude of contact networks involved in the process of disease spreading render this problem intractable from an analytic standpoint. Recently, sophisticated computational platforms have been developed to deal with disease prevention and forecast [5, 10, 11, 18, 27, 29,30,31,32,33,34,35,36, 51,52,53,54,55]. The computational complexity of these models reflects the intrinsic complexity of the problem at stake, and their success relies on careful calibration and validation procedures requiring biological and socio-geographic knowledge of the process at stake.
Our goal here, instead, will be to answer the following question: What is the impact of a temporal contact network structure in the overall dynamics of disease progression? Does one expect that it will lead to a rigid shift of the critical parameters driving disease evolution, as one witnesses whenever one includes spatial transmission patterns? Or even to an evanescence of their values whenever one models the contact network as a (static and infinite) scale-free network, such that the variance of the network degree distribution becomes arbitrarily large? Or will the temporal nature of the contact network lead to new dynamical features? And, if so, which features will emerge from the inclusion of this temporal dimension?
To answer this question computationally constitutes, in general, a formidable challenge. We shall attempt to address the problem analytically, and to this end some simplifications will be required. However, the simplifications we shall introduce become plausible taking into consideration recent results (i) in the evolutionary dynamics of social dilemmas of cooperation, (ii) in the dynamics of peer-influence, and even (iii) in the investigation of how individual behavior determines and is determined by the global, population wide behavior. All these recent studies point out to the fact that the impact of temporal networks in the population dynamics stems mostly from the temporal part itself, and not so much from the detailed shape and structure of the network [56,57,58,59,60,61,62,63]. Indeed, we now know that (i) different models of adaptive network dynamics lead to similar qualitative features regarding their impact in what concerns the evolution of cooperation [56,57,58,59,60,61,62,63], (ii) the degree of peer-influence is robust to the structural patterns associated with the underlying social networks [62], and (iii) the impact of temporal networks in connecting individual to collective behavior in the evolution of cooperation is very robust and related to a problem of N-body coordination [61, 63]. Altogether, these features justify that we model the temporal nature of the contact network in terms of a simple, adaptive network, the dynamics of which can be approximately described in terms a coupled system of ODEs. This “adaptive-linking” dynamics, as it was coined [28, 57,58,59], leads to network snapshot structures that do not replicate what one observes in real-life, in the same sense that the small-world model of Watts and Strogatz does not lead to the heterogeneous and diverse patterns observed in data snapshots of social networks. Notwithstanding, the active-linking dynamics allows us to include, analytically, the temporal dimension into the problem of disease dynamics. The results [28], as we elaborate in Sects. 3 and 4, prove rewarding, showing that the temporal dimension of a contact network leads to a shift of the critical parameters (defined below) which is no longer rigid but, instead, becomes dependent on the frequency of infected individuals in the population. This, we believe, constitutes a very strong message with a profound impact whenever one tries to incorporate the temporal dimension into computational models of disease forecast.
This chapter is organized as follows. In the following Sect. 2, we introduce the standard disease models we shall employ, as well as the details of the temporal contact network model. Section 3 is devoted to present and discuss the results, and Sect. 4 contains a summary of the main conclusions of this work.
13.2 Models and Methods
In this section, we introduce the disease models we shall employ which, although well-known and widely studied already, are here introduced in the context of stochastic dynamics in finite populations, a formulation that has received less attention than the standard continuous model formulation in terms of coupled Ordinary Differential Equations (ODEs). Furthermore, we introduce and discuss in detail the temporal contact network model.
13.2.1 Disease Spreading Models in Finite Populations
Here we introduce three standard models of disease transmission that we shall employ throughout the manuscript, using this section at profit to introduce also the appropriate notation associated with stochastic dynamics of finite populations and the Markov chain techniques that we shall also employ in the remainder of this chapter. We shall start by discussing the models in the context of well-mixed populations, which will serve as a reference scenario for the disease dynamics, leaving for the next section the coupling of these disease models with the temporal network model described below. We investigate the popular Susceptible-Infected-Susceptible (SIS) model [2, 4], the Susceptible-Infected (SI) model [2] used to study, e.g., AIDS [2, 64], and the Susceptible-Infected-Recovered (SIR) model [2, 65], more appropriate to model, for instance, single season flu outbreaks [2] or computer virus spreading [7]. It is also worth pointing out that variations of these models have been used to successfully model virus dynamics and the interplay between virus dynamics and the response of the immune system [66].
13.2.1.1 The SIS Model
In the SIS model individuals can be in one of two epidemiological states: Infected (I) or Susceptible (S). Each disease is characterized by a recovery rate (δ) and an infection rate (λ). In an infinite, well-mixed population, the fraction of infected individuals (x) changes in time according to the following differential equation
where y = 1 − x is the fraction of susceptible individuals and 〈k〉 the average number of contacts of each individual [4]. There are two possible equilibria (\( \dot{x}=0 \)): x = 0 and \( x=1-{R}_0^{-1} \), where R 0 = λ〈k〉/δ denotes the basic reproductive ratio. The value of R 0 determines the stability of these two equilibria: \( x=1-{R}_0^{-1} \) is stable when R 0 > 1 and unstable when R 0 < 1.
Let us now move to finite populations, and consider the well-mixed case where the population size is fixed and equal to N. We define a discrete stochastic Markov process describing the disease dynamics associated with the SIS model. Each configuration of the population, which is defined by the number of infected individuals i, corresponds to one state of the Markov chain. Time evolves in discrete steps and two types of events may occur which change the composition of the population: infection events and recovery events. This means that, similar to computer simulations of the SIS model on networked populations, at most one infection or recovery event will take place in each (discrete) time step. Thus, the dynamics can be represented as a Markov chain M with N+1 states [67, 68] — as many as the number of possible configurations — illustrated in the following Fig. 13.1.
In a finite, well-mixed population, the number i of infected will decrease at a rate given by
where τ 0 denotes the recovery time scale, \( \frac{i}{N} \) the probability that a randomly selected individual is infected and δ the probability that this individual recovers. Adopting τ 0 as a reference, we assume that the higher the average number of contacts 〈k〉, the smaller the time scale τ INF at which infection update events occur (τ INF = τ 0/〈k〉) [4]. Consequently, the number of infected will also increase at a rate given by
Equations (13.1) and (13.2) define the transitions between different states. This way, we obtain the following transition matrix for M:
where each element p kj of P represents the probability of moving from state k to state j during one time step. The state without any infected individual (i=0) is an absorbing state of M. In other words, the disease always dies out and will never re-appear, once this happens.
13.2.1.1.1 Average Times to Absorption
At this level of approximation, it is possible to derive an analytical expression for the average time t i it takes to reach the single absorbing state of the SIS Markov chain (i.e., the average time to absorption) starting from a configuration in which there are i infected individuals. Denoting by P i (t) the probability that the disease disappears at time t when starting with i infected individuals at time 0, we may write [69]
Using the properties of P i (t) we obtain the following recurrence relation for t i
whereas for t N we may write
Defining the auxiliary variables \( {\gamma}_i=\frac{T_i^{-}}{T_i^{+}} \) and \( {q}_i=\prod\limits_{l=1}^i{\gamma}_l \), a little algebra allows us to write, for t 1
such that t i can be written as a function of t 1 as follows
The intrinsic stochasticity of the model, resulting from the finiteness of the population, makes the disease disappear from the population after a certain amount of time. As such, the population size plays an important role in the average time to absorption associated with a certain disease, a feature we shall return to below.
13.2.1.1.2 Quasi-Stationary Distributions in Finite Populations
Equations (13.1) and (13.2) define the Markov chain M just characterized. The fraction of time the population spends in each state is given by the stationary distribution of M, which is defined as the eigenvector associated with eigenvalue 1 of the transition matrix of M [67, 68]. The fact that in the SIS model the state without infected (i=0) is an absorbing state of the Markov chain, implies that the standard stationary distribution will be completely dominated by this absorbing state, which precludes one to gather information on the relative importance of other configurations. This makes the so-called quasi-stationary distribution of M [70] the quantity of interest. This quantity allows us to estimate the relative prevalence of the population in configurations other than the absorbing state, by computing the stationary distribution of the Markov chain obtained from M by excluding the absorbing state i=0 [70]. It provides information on the fraction of time the population spends in each state, assuming the disease does not go extinct.
13.2.1.1.2.1 The Infinite, Well-Mixed Populations as a Limiting Case
The Markov process M defined before provides a finite population analogue of the well-known mean-field equations written at the beginning of Sect. 2.1.1. Indeed, in the limit of large populations, \( {\tau}_0G(i)={T}^{+}(i)-{T}^{-}(i) \) provides the rate of change of infected individuals. For large N, replacing \( \frac{i}{N} \) by x and \( \frac{N-i}{N} \) by y, the gradients of infection which characterize the rate at which the number of infected are changing in the population, are given by
Again, we obtain two roots: τ 0 G(i) = 0 for i = 0 and \( {i}_{r_0}^{\ast }=N-\frac{\left(N-1\right)\kern0.2em \delta }{\left\langle k\right\rangle \lambda } \). Moreover, \( {i}_{r_0}^{\ast } \) becomes the finite population equivalent of an interior equilibrium for \( {R}_0\equiv \frac{\lambda }{\delta}\left\langle k\right\rangle \frac{N}{N-1}>1 \) (note that, for large N we have that \( \frac{N}{N-1}\approx 1 \)). The disease will most likely expand whenever \( i<{i}_{r_0}^{\ast } \), the opposite happening otherwise.
13.2.1.2 The SI Model
The SI model is mathematically equivalent to the SIS model with δ = 0, and has been employed to study for instance the dynamics of AIDS. The Markov Chain representing the disease dynamics is therefore defined by transition matrix Eq. (13.3), with \( {T}_i^{-}=0 \) for all i. The remaining transition probabilities \( {T}_i^{+} \) (0 < i < N) are exactly the same as for the SIS model. Since all \( {T}_i^{-} \) equal zero, the Markov Chain has two absorbing states: the canonical one without any infected (i=0) and the one without any susceptible (i=N). The disease will expand monotonically as soon as one individual in the population gets infected, ultimately leading to a fully infected population. The average amount of time after which this happens, which we refer to as the average infection time, constitutes the main quantity of interest. This quantity can be calculated analytically [28]: The average number of time steps needed to reach 100% infection, starting from i infected individuals is given by
13.2.1.3 The SIR Model
With SIR one models diseases in which individuals acquire immunity after recovering from infection. We distinguish three epidemiological states to model the dynamics of such diseases: susceptible (S), infected (I) and recovered (R), indicating those who have become immune to further infection.
The SIR model in infinite, well-mixed populations is defined by a recovery rate δ and an infection rate λ. The fraction of infected individuals x changes in time according to the following differential equation
where y denotes the fraction of susceptible individuals, which in turn changes according to
Finally, the fraction of individuals z in the recovered class changes according to
To address the SIR model in finite, well-mixed populations, we proceed in a way similar to what we have done so far with SIS and SI models. The Markov Chain describing the disease dynamics becomes slightly more complicated and has states (i, r), where i is the number of infected individuals in the population and r the number of recovered (and immune) individuals (i + r ≤ N). A schematic representation of the Markov Chain is given in Fig. 13.2.
Note that the states (0, r), with 0 ≤ r ≤ N, are absorbing states. Each of these states corresponds to the number of individuals that are (or have become) immune at the time the disease goes extinct.
Consider a population of size N with average degree 〈k〉. The number of infected will increase with a rate
and decrease with a rate
where τ 0 denotes the recovery time scale. As before, the gradient of infection G(i), such that \( {\tau}_0G(i)={T}^{+}(i)-{T}^{-}(i) \), measures the likelihood for the disease to either expand or shrink in a given state, and is given by
Note that we recover Eq. (13.10) in the limit N → ∞. For a fixed number of recovered individuals r 0, we have that τ 0 G(i, r 0) = 0 for i = 0 and for \( {i}_{r_0}^{\ast }=N-\frac{\left(N-1\right)\delta }{\left\langle k\right\rangle \lambda }-{r}_0 \). For \( {R}_0^{r_0}=\left\langle k\right\rangle \frac{\lambda }{\delta}\frac{N-{r}_0}{N-1}>1 \), \( {i}_{r_0}^{\ast } \) becomes the finite population analogue of an interior equilibrium. Furthermore, one can show that the partial derivative \( \frac{\partial G\left(i,r\right)}{\partial i} \) has at most one single root in (0, 1), possibly located at \( {\overline{i}}_{r_0}=\frac{i_{r_0}^{\ast }}{2}\le {i}_{r_0}^{\ast } \). Hence, G(i, r 0) reaches a local maximum at \( {\overline{i}}_{r_0} \)(given that at that point \( {\left.\frac{\partial^2G\left(i,r\right)}{\partial {i}^2}\right|}_{{\overline{i}}_{r_0}}=-\frac{2\left\langle k\right\rangle \lambda }{N\left(N-1\right)}<0 \)). The number of infected will therefore most likely increase for \( i<{i}_{r_0}^{\ast } \) (assuming r 0 immune individuals), and most likely decrease otherwise.
The gradient of infection also determines the probability to end up in each of the different absorbing states of the Markov chain. These probabilities can be calculated analytically [28]. To this end, let us use \( {y}_{i,r}^a \) to denote the probability that the population ends up in the absorbing state with a recovered individuals, starting from a state with i infected and r recovered. We obtain the following recurrence relationship for \( {y}_{i,r}^a \)
which reduces to
The following boundary conditions
allow us to compute \( {y}_{i,r}^a \) for every a, i and r.
13.2.2 Network Model
Our network model explicitly considers a finite and constant population of N individuals. Its temporal contact structure allows, however, for a variable number of overall links between individuals, which in turn will depend on the incidence of disease in the population. This way, infection proceeds along the links of a contact network whose structure may change based on each individual’s health status and the availability of information regarding the health status of others. We shall assume the existence of some form of local information about the health status of social contacts. Information is local, in the sense that individual behavior will rely on the nature of their links in the contact network. Moreover, this will influence the way in which individuals may be more or less effective in avoiding contact with those infected while remaining in touch with the healthy.
Suppose all individuals seek to establish links at the same rate c. For simplicity, we assume that new links are established and removed randomly, a feature which usually does not always apply in real cases, where the limited social horizon of individuals or the nature of their social ties may constrain part of their neighborhood structure (see below). Let us further assume that links may be broken off at different rates, based on the nature of the links and the information available about the individuals they connect: Let us denote these rates by b pq for links of type pq (p , q ∈ {S, I, R}. We assume that links are bidirectional, which means that we have links of pq types SI, SR, and IR. Let L pq denote the number of links of type pq and \( {L}_{pq}^M \) the maximum possible number of links of that type, given the number of individuals of type S, I and R in the population. This allows us to write down (at a mean-field level) a system of ODEs [57, 58] for the time evolution of the number of links of pq-type (L pq ) [57, 58]
which depends on the number of individuals in states p and q (\( {L}_{pp}^M=p\left(p-1\right)/2 \) and \( {L}_{pq}^M= pq \) for p ≠ q) and thereby couples the network dynamics to the disease dynamics. In the steady state of the linking dynamics (\( {\dot{L}}_{pq}=0 \)), the number of links of each type is given by \( {L}_{pq}^{\ast }={\varphi}_{pq}{L}_{pq}^M \), with φ pq = c/(c + b pq ) the fractions of active pq-links, compared to the maximum possible number of links \( {L}_{pq}^M \), for a given number of S, I and R. In the absence of disease only SS links exist, and hence ϕ SS determines the average connectivity of the network under disease free conditions, which one can use to characterize the type of the population under study. In the presence of I individuals, to the extent that S individuals manage to avoid contact with I, they succeed in escaping infection. Thus, to the extent that individuals are capable of reshaping the contact network based on available information of the health status of other individuals, disease progression will be inhibited. In the extreme limit of perfect information and individual capacity to immediately break up contacts with infected, we are isolating all infected, and as such containing disease progression. Our goal here, however, is to understand how and in which way local information, leading to a temporal reshaping of the network structure, affects overall disease dynamics.
13.2.3 Computer Simulations
We investigate the validity of the approximations made to derive analytical results as well as their robustness by means of computer simulations. All individual-based simulations start from a complete network of size N=100. Disease spreading and network evolution proceed together under asynchronous updating. Disease update events take place with probability (1 + τ)−1, where τ = τ NET /τ DIS . We define τ DIS as the time-scale of disease progression, whereas τ NET is the time scale of network change. The parameter τ = τ NET /τ DIS provides the relative time scale in terms of which we may interpolate between the limits when network adaptation is much slower than disease progression (τ → 0) and the opposite limit when network adaptation is much faster than disease progression (τ → ∞). Since τ = τ NET /τ DIS is the only relevant parameter, we can make, without loss of generality, τ DIS = 1. For network update events, we randomly draw two nodes from the population. If connected, then the link disappears with probability given by the respective b pq . Otherwise, a new link appears with probability c. When a disease update event occurs, a recovery event takes place with probability (1 + 〈k〉)−1, an infection event otherwise. In both cases, an individual j is drawn randomly from the population. If j is infected and a recovery event has been selected then j will become susceptible (or recovered, model dependent) with probability δ. If j is susceptible and an infection event occurs, then j will get infected with probability λ if a randomly chosen neighbor of j is infected. The quasi-stationary distributions are computed (in the case of the SIS model) as the fraction of time the population spends in each configuration (i.e., number of infected individuals) during 109 disease event updates (107 generations; under asynchronous updating, one generation corresponds to N update events, where N is the population size; this means that in one generation, every individual has one chance, on average, to update her epidemic state). The average number of infected 〈I〉 and the mean average degree of the network 〈k〉∗ observed during these 107 generations are kept for further plotting. We have checked that the results reported are independent of the initial number of infected in the network. Finally, for the SIR and SI models, the disease progression in time, shown in the following sections, is calculated from 104 independent simulations, each simulation starting with 1 infected individual. The reported results correspond to the average amount of time at which i individuals become infected.
13.3 Results and Discussion
In this section we start by (i) showing that a quickly adapting community induces profound changes in the dynamics of disease spreading, irrespective of the underlying epidemic model; then, (ii) we resort to computer simulations to study the robustness of these results for intermediate time-scales of network adaptation; finally, (iii) we profit from the framework introduced above to analyze the impact of information on average time for absorption and disease progression in adaptive networks.
13.3.1 Disease Spreading in a Quickly Adaptive Network Structure
Empirically, it is well-known that often individuals prevent infection by avoiding contact with infected once they know the state of their contacts or are aware of the potential risks of such infection [31, 33, 42,43,44,45,46,47,48,49,50]: such is the case of many sexually transmitted diseases [42, 71,72,73], for example, and, more recently, the voluntary use of face masks and the associated campaigns adopted by local authorities in response to the SARS outbreak [40, 43,44,45] or even the choice of contacting or not other individuals based on information on their health status gathered from social media [41, 74, 75]. In the present study, individual decision is based on available local information about the health state of one’s contacts. Thus, we can study analytically the limit in which the network dynamics — resulting from adaptation to the flow of local information — is much faster than disease dynamics, as in this case, one may separate the time scales between network adaptation and contact (disease) dynamics: The network has time to reach a steady state before the next contact takes place. Consequently, the probability of having an infected neighbor is modified by a neighborhood structure which will change in time depending on the impact of the disease in the population and the overall rates of severing links with infected.
Let us start with the SIR model. The amount of information available translates into differences mostly between the break-up rates of links that may involve a potential risk for further infection (b SI , b IR , b II ), and those that do not (b SS , b SR , b RR ). Therefore, we consider one particular rate b I for links involving infected individuals (b I ≡ b SI = b IR = b II ), and another one, b H , for links connecting healthy individuals (b H ≡ b SS = b SR = b RR ). In general, one expects b I to be maximal when each individual has perfect information about the state of her neighbors and to be (minimal and) equal to b H when no information is available, turning the ratio between these two rates into a quantitative measure of the efficiency with which links to infected are severed compared to other links. Note that we reduce the model to two break-up rates in order to facilitate the discussion of the results. Numerical simulations show that the general principles and conclusions remain valid when all break-up rates are incorporated explicitly. It is worth noticing that three out of these six rates are of particular importance for the overall disease dynamics: b SS , b SR and b SI . These three rates, combined with the rate c of creating new links, define the fraction of active SS, SR and SI links, and subsequent correlations between individuals [76], and therefore determine the probability for a susceptible to become infected (see Models and Methods). This probability will increase when considering higher values of c (assuming b I > b H ). In other words, when individuals create new links more often, therefore increasing the likelihood of establishing connections to infected individuals (when present), they need to be better informed about the health state of their contacts in order to escape infection. In the fast linking limit, the other three break-up rates (b II , b IR and b RR ) will also influence disease progression since they contribute to changing the average degree of the network.
When the time scale for network update (τ NET ) is much smaller than the one for disease spreading (τ DIS ), we can proceed analytically using at profit the separation of times scales. In practice, this means that the network has time to reach a steady state before the next disease event takes place. Consequently, the probability of having an infected neighbor is modified by a neighborhood structure which will change in time depending on the impact of the disease in the population and the overall rates of severing links with infected individuals. For a given configuration (i,r) of the population, the stationary state of the network is characterized by the parameters φ SS , φ SI and φ SR . Consequently, the number of infected increases at a rate [28]
where we made τ 0 = 1. The effect of the network dynamics becomes apparent in the third factor, which represents the probability that a randomly selected neighbor of a susceptible is infected. In addition, Eq. (13.14) remains valid, as the linking dynamics does not affect the rate at which the number of infected decreases. It is noteworthy that we can write Eq. (13.19) in the form
which is formally equivalent to Eq. (13.13) and shows that disease spreading in a temporal adaptive network is equivalent to that in a well-mixed population with (i) a frequency dependent average degree 〈k〉 and (ii) a transmission probability that is rescaled compared to the original λ according to λ A = η −1 λ, where
Note that this expression remains valid for both SIR, SIS (r = 0) and SI (δ = 0, r = 0) models. Since the lifetime of a link depends on its type, the average degree 〈k〉 of the network depends on the number of infected in the population, and hence becomes frequency (and time) dependent, as 〈k〉 depends on the number of infected (through \( {L}_{pq}^M \)) and changes in time. Note that η scales linearly with the frequency of infected in the population, decreasing as the number of infected increases (assuming ϕ SS /ϕ SI > 1); moreover, it depends implicitly (via the ratio ϕ SS /ϕ SI ) on the amount of information available.
It is important to stress the distinction between the description of the disease dynamics at the local level (in the vicinity of an infected individual) and that at the population wide level. Strictly speaking, a dynamical network does not change the disease dynamics at the local level, meaning that infected individuals pass the disease to their neighbors with probability intrinsic to the disease itself. At the population level, on the other hand, disease progression proceeds as if the infectiousness of the disease effectively changes, as a result of the network dynamics. Consequently, analyzing a temporal network scenario at a population level can be achieved via a renormalization of the transmission probability, keeping the (mathematically more attractive) well-mixed scenario. In this sense, from a well-mixed perspective, dynamical networks contribute to changing the effective infectiousness of the disease, which becomes frequency and information dependent. Note further that this information dependence is a consequence of using a single temporal network for spreading the disease and information. Interestingly, adaptive networks have been shown to have a similar impact in social dilemmas [63]. From a global, population-wide perspective, it is as if the social dilemma at stake differs from the one every individual actually plays.
As in Sect. 2, one can define a gradient of infection G, which measures the tendency of the disease to either expand or shrink in a population with given configuration (defined by the number of individuals in each of the states S, I and R). To do so, we study the partial derivative \( \frac{\partial G\left(i,r\right)}{\partial i} \) at i = 0
This quantity exceeds zero whenever
Note that taking r = 0 yields the basic reproductive ratio \( {R}_0^A \) for both SIR and SIS: \( {R}_0^A\equiv N{\phi}_{SI}\frac{\lambda }{\delta }>1 \). On the other hand, whenever \( {R}_0^A<1 \), eradication of the disease is favored in the SIS model (G(i)<0), irrespective of the fraction of infected, indicating how the presence of information (b H < b I ) changes the basic reproductive ratio.
In Fig. 13.3 we illustrate the role of information in the SIS model by plotting G for different values of b I (assuming b H < b I ) and a fixed transmission probability λ. The corresponding quasi-stationary distributions are shown in the right panel and clearly reflect the sign of G. Whenever G(i) is positive (negative), the dynamics will act to increase (decrease), on average, the number of infected. Figure 13.3 indicates how the availability of local information hinders disease progression: For b I = 0.75 the interior root of G(i) disappears, making disease expansion unlikely in any configuration of the population.
Disease spreading under fast linking dynamics in the SIS model. The left panel shows the gradient of infection G as a function of the fraction of infected for different values of the rate b 1 at which links with infected disappear (b I ≡ b SI = b II ): b I = 0.8 (dotted line), b I = 0.4 (dashed line) and b I = 0.2 (solid line). The right panel shows the corresponding quasi-stationary distributions, obtained analytically (lines) and via individual-based computer simulations (circles for b I = 0.8, squares for b I = 0.4 and crosses for b I = 0.2). We use b H ≡ b SS = 0.2, c = 0.25, N = 100, \( {N\lambda }\left/{\delta}\right.=4 \) and τ = 10−2
The analysis of the gradient of infection of the SIS model has the advantage of showing the effect of adaptive networks in a one-dimensional simplex (the fraction of infected). Yet, an analogous result holds for the SIR model. The gradient of infection now also depends on the number of recovered (r) individuals in the population and, once again, allows us to identify when disease expansion will be favored or not. Figure 13.4 gives a complete picture of the gradient of infection, using the appropriate simplex structure in which all points satisfy the relation i+r+s=N. The dashed line indicates the boundary G(i, r) = 0 in case individuals do not have any information about the health status of their contacts, i.e., links that involve infected individuals disappear at the same rate as those that do not (b I = b H ). Disease expansion is more likely than disease contraction (G(i, r) > 0) when the population is in a configuration above the line, and less likely otherwise. Similarly, the solid line indicates the boundary G(i, r) = 0 when individuals share information about their health status, and use it to avoid contact with infected. Once again, the availability of information modifies the disease dynamics, inhibiting disease progression for a broad range of configurations.
Gradient of infection in the SIR model in a network with information (solid black line, b I = 0.8, b H = 0.2), and without information (dashed white line, b I = b H = 0.2). Each point in the triangle (the so-called simplex) satisfies that population size is conserved, i.e., i+r+s=N. Vertices of the simplex represent populations with only one class of individuals present. Lines in the interior of the simplex indicate configurations in which G(i, r) = 0. For each case, disease expansion is more likely than disease contraction in configurations above the line, and less likely otherwise, showing that availability of information greatly reduces the regions of state space in which disease may progress We use the following parameter values: (\( c=0.25, N=100,\ {N\lambda}\left/{\delta}\right.=10 \))
13.3.2 Analysis of Intermediate Time-Scales Through Computer Simulations
Up to now we have assumed that the network dynamics proceeds much faster than disease spreading (the limit τ → 0). This may not always be the case, and hence it is important to assess the domain of validity of this limit. In the following, we use computer simulations to verify to which extent these results, obtained analytically via time scale separation, remain valid for intermediate values of the relative timescaleτ for the linking dynamics. We start with a complete network of size N, in which initially one individual is infected, the rest being susceptible. As stated before, disease spreading and network evolution proceed simultaneously under asynchronous updating. Network update events take place with probability (1 + τ)−1, whereas a disease model (SI, SIS or SIR) state update event occurs otherwise. For each value of τ, we run 104 simulations. For the SI model, the quantity of interest to calculate is the average number of generations after which the population becomes completely infected. These values are depicted in Fig. 13.5.
Disease spreading in the SI model for variable time scales τ of the linking dynamics. Solid circles show the average number of generations to reach a fully infected population, starting from one single infected individual, obtained in simulation. Dashed lines indicate the analytical predictions for these values, either in the limit τ → 0 (upper dashed line), or in the limit τ → ∞ (lower dashed line). We use the following parameter values: b I = 0.8, b H = 0.2, c = 0.25, N = 100 and λ = 0.001
The lower dashed line indicates the analytical prediction of the infection time in the limit τ → ∞ (the limit when networks remain static), which we already recover in the simulations for τ > 102. When τ is smaller than 102, the average infection time significantly increases, and already reaches the analytical prediction for the limit τ → 0 (indicated by the upper dashed line) when τ < 1. Hence, the validity of the time scale separation does again extend well beyond the limits one might expect.
For the SIR model, we let the simulations run until the disease goes extinct, and computed the average final fraction of individuals that have been affected by the disease, which corresponds to the final fraction of individuals in the recovered class. These results are depicted in Fig. 13.6.
Disease spreading in the SIR model for variable time scales τ of the linking dynamics. Solid circles show the final fraction of recovered individuals as a function of τ in populations with initially one infected. The upper (lower) dashed line shows the corresponding analytical prediction in the limit τ → ∞ (τ → 0). We use the following parameter values: b I = 0.8, b H = 0.2, c = 0.25, λ = 0.01, δ = 0.15 and N = 100
The upper dashed line indicates the expected fraction of recovered individuals in a static network (τ → ∞). This value is obtained by calculating \( \sum\limits_{i=0}^Ni\kern0.1em {y}_{1,0}^i \), where \( {y}_{1,0}^i \) is given by Eqs. (13.17) and (13.18). One observes that linking dynamics does not affect disease dynamics for τ > 10. Once τ drops below ten, a significantly smaller fraction of individuals is affected by the disease. This fraction reaches the analytical prediction for τ → 0 as soon as τ < 0.1. Hence, and again, results obtained via separation of time scales remain valid for a wide range of intermediate time scales.
We finally investigate the role of intermediate time scales in the SIS model. We performed computer simulations in the conditions discussed already, and computed several quantities that we plot in Fig. 13.7.
Disease spreading under linking dynamics in the SIS model. Circles show results of individual-based simulations for the quasi-stationary average fraction of infected 〈I〉 as function of τ. The lower (upper) dashed line shows the analytical prediction of 〈I〉 for τ → 0 (τ → ∞), calculated as the average of the quasi-stationary distribution. The analytical prediction in the fast linking limit (τ → 0) remains valid as long as τ < 1, whereas the prediction in the limit of static networks (τ → ∞) remains valid as long as τ > 105. The solid line depicts the analytical prediction of 〈I〉 in static networks whose average degree equals the value obtained computationally for the average connectivity of the network at each givenτ. Results show that for τ > 102, the network dynamics influences disease progression only by controlling 〈k〉∗. We use b I = 0.8, b H = 0.2, c = 0.25, N = 100 and \( {N\lambda}\left/{\delta}\right.=4 \)
Figure 13.7 shows the average 〈I〉 of the quasi-stationary distributions obtained via computer simulations (circles) as a function of the relative time scale τ of network update. Whenever τ → ∞, we can characterize the disease dynamics analytically, assuming a well-mixed population (complete graph), whereas for τ → 0 we recover the analytical results obtained in the fast linking limit. At intermediate time scales, Fig. 13.7 shows that as long as τ is smaller than ten, network dynamics contributes to inhibit disease spreading by effectively increasing the critical infection rate. Overall, the validity of the time scale separation extends well beyond the limits one might anticipate based solely on the time separation ansatz. As long as the time scale for network update is smaller than the one for disease spreading (τ < 1), the analytical prediction for the limit τ → 0, indicated by the lower dashed line in Fig. 13.7, remains valid. The analytical result in the extreme opposite limit (τ → ∞), indicated by the upper dashed line in Fig. 13.7, holds as long as τ > 105. Moreover, it is noteworthy that the network dynamics influences the disease dynamics both by reducing the frequency of interactions between susceptible and infected, and by reducing the average degree of the network. These complementary effects are disentangled in intermediate regimes, in which the network dynamics is too slow to warrant sustained protection of susceptible individuals from contacts with infected, despite managing to reduce the average degree (not shown). In fact, for τ > 10 the disease dynamics is mostly controlled by the average degree, as shown by the solid lines in Fig. 13.7. Here, the average stationary distribution was determined by replacing, in the analytic expression for static networks, 〈k〉 by the time-dependent average connectivity 〈k〉∗ computed numerically. This, in turn, results from the frequency dependence of 〈k〉. When b I > b H , the network will reshape into a configuration with smaller 〈k〉 as soon as the disease expansion occurs. For τ < 1, 〈k〉∗ reflects the lifetime of SS links, as there are hardly any infected in the population. For 100 < τ < 103, the network dynamics proceeds fast enough to reduce 〈k〉, but too slowly to reach its full potential in hindering disease progression. Given the higher fraction of infected, and the fact that SI and II links have a shorter lifetime than SS links, the average degree drops when increasing τ from 1 to 103. Any further increase in τ leads to a higher average degree, as the network approaches its static limit.
13.3.3 Average Time to Absorption in Adaptive Networks
Contrary to the deterministic SIS model, the stochastic nature of disease spreading in finite populations ensures that the disease disappears after some time. However, this result is of little relevance given the times required to reach the absorbing state (except, possibly, in very small communities). Indeed, the characteristic time scale of the dynamics plays a determinant role in the overall epidemiological process and constitutes a central issue in disease spreading.
Figure 13.8 shows the average time to absorption t 1 in adaptive networks for different levels of information, illustrating the spectacular effect brought about by the network dynamics on t 1 . While on networks without information (b I = b H ) t 1 rapidly increases with the rate of infection λ, adding information moves the fraction of infected individuals rapidly to the absorbing state, and, therefore, to the disappearance of the disease.
Impact of information on times to absorption. Average number of generations required for disease eradication in an adaptive contact network for different rates b I , using the SIS model. The remaining parameters are b H = 0.2, c = 0.25 and N = 100. The availability of information drastically reduces the time for disease eradication
Moreover, the size of the population can have a profound effect on t 1 . With increasing population size, the population spends most of the time in the vicinity of the state associated with the interior root of G(i). For large populations, this acts to reduce the intrinsic stochasticity of the dynamics, dictating a very slow extinction of the disease, as shown in Fig. 13.9.
Impact of population size on the average times to absorption t 1 . (a) Average number of generations required for disease eradication in the SIS model in static networks of different size N, while keeping the average degree 〈k〉 constant (〈k〉 = 49). (b) Quasi-stationary distribution of the number of infected for the same values of N and 〈k〉. The disease parameters satisfy 〈k〉λ/δ = 2
When recovery from the disease is impossible, a situation captured by the SI model, the population will never become disease-free again once it acquires at least one infected individual. The time to reach absorbing state in which all individuals are infected, again depends on the presence of information. When information prevails, susceptible individuals manage to resist infection for a long time, thereby delaying the rapid progression of the disease, as shown in the inset of Fig. 13.10. Naturally, the average number of generations needed to reach a fully infected population increases with the availability of information, as illustrated in the main panel of Fig. 13.10.
Impact of information on infection times. The main plot shows the average number of generations after which a disease infects the entire population in the SI model, using the same parameters as in Fig. 13.8. The inset shows how, starting from one infected individual, the fraction of infected changes in time for the same rates b I and λ = 10−3. The results obtained via individual-based computer simulations (circles, τ = 10−1) fit perfectly with those calculated analytically (lines)
13.4 Conclusions
Making use of three standard models of epidemics involving a finite population in which infection takes place along the links of a temporal graph, the nodes of which are occupied by individuals, we have shown analytically that the bias introduced into the graph dynamics resulting from the availability of information about the health status of others in the population induces fundamental changes in the overall dynamics of disease progression.
The network dynamics employed here differs from those used in most other studies [29, 32,33,34,35,36, 51,52,53,54,55]. We argue, however, that the differences obtained stem mostly from the temporal aspect of the network, and not so much from the detailed dynamics that is implemented. Importantly, temporal network dynamics leads to additional changes in R 0 compared to those already obtained when moving from the well-mixed assumption to static networks [77]. An important ingredient of our model, however, is that the average degree of the network results from the self-organization of the network structure, and co-evolves with the disease dynamics. A population suffering from high disease prevalence where individuals avoid contact in order to escape infection will therefore exhibit a lower average degree than a population with hardly any infected individuals. Such a frequency-dependent average degree further prevents that containment of infected individuals would result in the formation of cliques of susceptible individuals, which are extremely vulnerable to future infection, as reported before [36, 51, 54].
The description of disease spreading as a stochastic contact process embedded in a Markov chain constitutes a second important ingredient of the present model. This approach allows for a direct comparison between analytical predictions and individual-based computer simulations, and for a detailed analysis of finite-size effects and convergence times, whose exponential growth will signal possible bistable disease scenarios. In such a framework, we were able to show that temporal adaptive networks in which individuals may be informed about the health status of others lead to a disease whose effective infectiousness depends on the overall number of infected in the population. In other words, disease propagation on temporal adaptive networks can be seen as mathematically equivalent to disease spreading on a well-mixed population, but with a rescaled effective infectiousness. In accord with the intuition advanced in the introduction, as long as individuals react promptly and consistently to accurate available information on whether their acquaintances are infected or not, network dynamics effectively weakens the disease burden the population suffers. Last but not least, if recovery from the disease is possible, the time for disease eradication drastically reduces whenever individuals have access to accurate information about the health state of their acquaintances and use it to avoid contact with those infected. If recovery or immunity is impossible, the average time needed for a disease to spread increases significantly when such information is being used. In both cases, our model clearly shows how availability of information hinders disease progression (by means of quick action on infected, e.g., their containment via link removal), which constitutes a crucial factor to control the development of global pandemics.
Finally, it is also worth mentioning that knowledge about the health state of others may not always be accurate or available in time. This is for instance the case for diseases where recently infected individuals remain asymptomatic for a substantial period. The longer the incubation period associated with the disease, the less successful individuals will be in escaping infection, which in our model translates into a lower effective rate of breaking SI links, with the above mentioned consequences. Moreover, different (social) networks through which awareness of the health status of others proceeds may lead to different rates of information spread. One may take these features into account by modeling explicitly the spread of information through a coupled dynamics between disease expansion and individuals’ awareness of the disease [31, 33].
Creation and destruction of links may for instance not always occur randomly, as we assumed here, but in a way that is biased by a variety of factors such as social and genetic distance, geographical proximity, family ties, etc. The resulting contact network may therefore become organized in a specific way, promoting the formation of particular structures, such as networks characterized by long-tailed degree distributions or with strong topological correlations among nodes [3, 78,79,80] which, in turn, may influence the disease dynamics. The impact of combining such effects, resulting from specific disease scenarios, with those reported here will depend on the prevalence of such additional effects when compared to link-rewiring dynamics. A small fraction of non-random links, or of ties which cannot be broken, will likely induce small modifications on the average connectivity of the contact network, which can be incorporated in our analytic expressions without compromising their validity regarding population wide dynamics. On the other hand, when the contact network is highly heterogeneous (e.g., exhibiting pervasive long-tail degree distributions), non-random events may have very distinct effects, from being almost irrelevant (and hence can be ignored) to inducing hierarchical cascades of infection [81], in which case our results will not apply.
References
Keeling, M.J., Rohani, P.: Modeling Infectious Diseases in Humans and Animals. Princeton University Press, Princeton (2008)
Anderson, R.M., May, R.M.: Infectious Diseases in Humans. Oxford University Press, Oxford (1992)
Dorogovtsev, S.N., Mendes, J.F.F.: Evolution of Networks: From Biological Nets to the Internet and WWW. Oxford University Press, Oxford (2003)
Barrat, A., Barthelemy, M., Vespignani, A.: Dynamical Processes in Complex Networks. Cambridge University Press, Cambridge (2008)
Pastor-Satorras, R., Castellano, C., Van Mieghem, P., Vespignani, A.: Epidemic processes in complex networks. Rev. Mod. Phys. 87(3), 925–979 (2015)
Watts, D.J.: Small Worlds: The Dynamics of Networks Between Order and Randomness. Princeton University Press, Princeton/Oxford (1999)
Lloyd, A.L., May, R.M.: Epidemiology. How viruses spread among computers and people. Science. 292(5520), 1316–1317 (2001)
Santos, F.C., Rodrigues, J.F., Pacheco, J.M.: Epidemic spreading and cooperation dynamics on homogeneous small-world networks. Phys. Rev. E. 72(5 Pt 2), 056128 (2005)
May, R.M.: Network structure and the biology of populations. Trends Ecol. Evol. 21(7), 394–399 (2006)
Mastrandrea, R., Barrat, A.: How to estimate epidemic risk from incomplete contact diaries data? PLoS Comput. Biol. 12(6), e1005002 (2016)
Kiti, M.C., Tizzoni, M., Kinyanjui, T.M., Koech, D.C., Munywoki, P.K., Meriac, M., Cappa, L., Panisson, A., Barrat, A., Cattuto, C.: Quantifying social contacts in a household setting of rural Kenya using wearable proximity sensors. EPJ Data Sci. 5(1), 21 (2016)
Fournet, J., Barrat, A.: Epidemic risk from friendship network data: an equivalence with a non-uniform sampling of contact networks. Sci. Rep. 6, 24593 (2016)
Merler, S., Ajelli, M., Fumanelli, L., Gomes, M.F., y Piontti, A.P., Rossi, L., Chao, D.L., Longini, I.M., Halloran, M.E., Vespignani, A.: Spatiotemporal spread of the 2014 outbreak of Ebola virus disease in Liberia and the effectiveness of non-pharmaceutical interventions: a computational modelling analysis. Lancet Infect. Dis. 15(2), 204–211 (2015)
Holme, P., Masuda, N.: The basic reproduction number as a predictor for epidemic outbreaks in temporal networks. PLoS One. 10(3), e0120567 (2015)
Holme, P.: Information content of contact-pattern representations and predictability of epidemic outbreaks. Sci. Rep. 5, 14462 (2015)
Holme, P., Liljeros, F.: Birth and death of links control disease spreading in empirical contact networks. Sci. Rep. 4, 4999 (2014)
Salathé, M., Freifeld, C.C., Mekaru, S.R., Tomasulo, A.F., Brownstein, J.S.: Influenza A (H7N9) and the importance of digital epidemiology. N. Engl. J. Med. 369(5), 401 (2013)
Masuda, N., Holme, P.: Predicting and controlling infectious disease epidemics using temporal networks. F1000 prime reports. 5, 6 (2013)
Goltsev, A.V., Dorogovtsev, S.N., Oliveira, J.G., Mendes, J.F.F.: Localization and spreading of diseases in complex networks. Phys. Rev. Lett. 109(12), 128702 (2012)
Swinburn, B.A., Sacks, G., Hall, K.D., McPherson, K., Finegood, D.T., Moodie, M.L., Gortmaker, S.L.: The global obesity pandemic: shaped by global drivers and local environments. Lancet. 378(9793), 804–814 (2011)
Salathé, M., Kazandjieva, M., Lee, J.W., Levis, P., Feldman, M.W., Jones, J.H.: A high-resolution human contact network for infectious disease transmission. Proc. Natl. Acad. Sci. U. S. A. 107(51), 22020–22025 (2010)
Salathé, M., Jones, J.H.: Dynamics and control of diseases in networks with community structure. PLoS Comput. Biol. 6(4), e1000736 (2010)
Funk, S., Salathé, M., Jansen, V.A.: Modelling the influence of human behaviour on the spread of infectious diseases: a review. J. R. Soc. Interface. 7(50), 1247–1256 (2010)
Masuda, N., Lambiotte, R.: A Guide to Temporal Networks, vol. 4. World Scientific, London (2016)
Holme, P., Saramäki, J.: Temporal networks. Phys. Rep. 519(3), 97–125 (2012)
Barrat, A., Cattuto, C., Colizza, V., Gesualdo, F., Isella, L., Pandolfi, E., Pinton, J.-F., Ravà, L., Rizzo, C., Romano, M.: Empirical temporal networks of face-to-face human interactions. EPJ ST. 222(6), 1295–1309 (2013)
Lee, S., Rocha, L.E., Liljeros, F., Holme, P.: Exploiting temporal network structures of human interaction to effectively immunize populations. PLoS One. 7(5), e36439 (2012)
Van Segbroeck, S., Santos, F.C., Pacheco, J.M.: Adaptive contact networks change effective disease infectiousness and dynamics. PLoS Comput. Biol. 6(8), e1000895 (2010)
Schwartz, I.B., Shaw, L.B.: Rewiring for adaptation. Physics. 3(17), (2010)
Marceau, V., Noël, P.-A., Hébert-Dufresne, L., Allard, A., Dubé, L.J.: Adaptive networks: coevolution of disease and topology. Phys. Rev. E. 82(3), (2010)
Funk, S., Gilad, E., Jansen, V.A.A.: Endemic disease, awareness, and local behavioural response. J. Theor. Biol. 264, 501–509 (2010)
Risau-Gusman, S., Zanette, D.H.: Contact switching as a control strategy for epidemic outbreaks. J. Theor. Biol. 257, 52–60 (2009)
Funk, S., Gilad, E., Watkins, C., Jansen, V.A.: The spread of awareness and its impact on epidemic outbreaks. Proc. Natl. Acad. Sci. U. S. A. 106(16), 6872–6877 (2009)
Zanette, D.H., Risau-Gusman, S.: Infection spreading in a population with evolving contacts. J. Biol. Phys. 34, 135–148 (2008)
Shaw, L.B., Schwartz, I.B.: Fluctuating epidemics on adaptive networks. Phys. Rev. E. 77, 066101 (2008)
Gross, T., Blasius, B.: Adaptive coevolutionary networks: a review. J. R. Soc. Interface. 5(20), 259–271 (2008)
Galvani, A.P., Reluga, T.C., Chapman, G.B.: Long-standing influenza vaccination policy is in accord with individual self-interest but not with the utilitarian optimum. Proc. Natl. Acad. Sci. U. S. A. 104(13), 5692–5697 (2007)
Colizza, V., Barrat, A., Barthelemy, M., Valleron, A.J., Vespignani, A.: Modeling the worldwide spread of pandemic Influenza: baseline case and containment interventions. PLoS Med. 4(1), e13 (2007)
Hufnagel, L., Brockmann, D., Geisel, T.: Forecast and control of epidemics in a globalized world. Proc. Natl. Acad. Sci. U. S. A. 101(42), 15124–15129 (2004)
Svoboda, T., Henry, B., Shulman, L., Kennedy, E., Rea, E., Ng, W., Wallington, T., Yaffe, B., Gournis, E., Vicencio, E., Basrur, S., Glazier, R.H.: Public health measures to control the spread of the severe acute respiratory syndrome during the outbreak in Toronto. N. Engl. J. Med. 350(23), 2352–2361 (2004)
Salathé, M., Bengtsson, L., Bodnar, T.J., Brewer, D.D., Brownstein, J.S., Buckee, C., Campbell, E.M., Cattuto, C., Khandelwal, S., Mabry, P.L.: Digital epidemiology. PLoS Comput. Biol. 8(7), e1002616 (2012)
Ahituv, A., Hotz, V.J., Philipson, V.J.: The responsiveness of the demand for condoms to the local prevalence of AIDS. J. Hum. Resour. 31(4), 869–897 (1996)
Kristiansen, I.S., Halvorsen, P.A., Gyrd-Hansen, D.: Influenza pandemic: perception of risk and individual precautions in a general population. BMC Public Health. 7(48), (2007)
Lau, J.T., Yang, X., Tsui, H., Kim, J.H.: Impacts of SARS on health-seeking behaviors in general population in Hong Kong. Prev. Med. 41(2), 454–462 (2005)
Ferguson, N.: Capturing human behaviour. Nature. 446(7137), 733 (2007)
Laver, S.M., Wetzels, J., Behrens, R.H.: Knowledge of malaria, risk perception, and compliance with prophylaxis and personal and environmental preventinve measures in travelers exiting Zimbabwe from Harare and Victoria Falls International Airport. J. Travel Med. 8(6), 298–303 (2001)
Brewer, N.T., Chapman, G.B., Gibbons, F.X., Gerrard, M., McCaul, K.D., Weinstein, N.D.: Meta-analysis of the relationship between risk perception and health behavior: the example of vaccination. Health Psychol. 26(2), 136–145 (2007)
Brewer, N.T., Cuite, C.L., Herrington, J.E., Weinstein, N.D.: Risk compensation and vaccination: can getting vaccinated cause people to engage in risky behaviors? Ann. Behav. Med. 34(1), 95–99 (2007)
Rubin, G.J., Amlot, R., Page, L., Wessely, S.: Public perceptions, anxiety, and behaviour change in relation to the swine flu outbreak: cross sectional telephone survey. BMJ. 339, b2651 (2009)
Jones, J.H., Salathe, M.: Early assessment of anxiety and behavioral response to novel swine-origin influenza A(H1N1). PLoS One. 4(12), e8032 (2009)
Gross, T., D'Lima, C.J., Blasius, B.: Epidemic dynamics on an adaptive network. Phys. Rev. Lett. 96(20), 208701 (2006)
Volz, E., Meyers, L.A.: Susceptible-infected-recovered epidemics in dynamic contact networks. Proc. R. Soc. B. 274, 2925–2933 (2007)
Han, X.-P.: Disease spreading with epidemic alert on small-world networks. Phys. Lett. A. 365, 1–5 (2008)
Gross, T., Kevrekidis, I.G.: Robust oscillations in SIS epidemics on adaptive networks: coarse graining by automated moment closure. Europhys. Lett. 82, 38004–38006 (2008)
Prado, F., Sheih, A., West, J.D., Kerr, B.: Coevolutionary cycling of host sociality and pathogen virulence in contact networks. J. Theor. Biol. 261(4), 561–569 (2009)
Santos, F.C., Pacheco, J.M., Lenaerts, T.: Cooperation prevails when individuals adjust their social ties. PLoS Comput. Biol. 2(10), e140 (2006)
Pacheco, J.M., Traulsen, A., Nowak, M.A.: Coevolution of strategy and structure in complex networks with dynamical linking. Phys. Rev. Lett. 97(25), 258103 (2006)
Pacheco, J.M., Traulsen, A., Nowak, M.A.: Active linking in evolutionary games. J. Theor. Biol. 243, 437–443 (2006)
Pacheco, J.M., Traulsen, A., Ohtsuki, H., Nowak, M.A.: Repeated games and direct reciprocity under active linking. J. Theor. Biol. 250(4), 723–731 (2008)
Van Segbroeck, S., Santos, F.C., Lenaerts, T., Pacheco, J.M.: Reacting differently to adverse ties promotes cooperation in social networks. Phys. Rev. Lett. 102, 058105 (2009)
Van Segbroeck, S., Santos, F.C., Lenaerts, T., Pacheco, J.M.: Selection pressure transforms the nature of social dilemmas in adaptive networks. New J. Phys. 13(1), 013007 (2011)
Pinheiro, F.L., Santos, M.D., Santos, F.C., Pacheco, J.M.: Origin of peer influence in social networks. Phys. Rev. Lett. 112(9), 098702 (2014)
Pinheiro, F.L., Santos, F.C., Pacheco, J.M.: Linking individual and collective behavior in adaptive social networks. Phys. Rev. Lett. 116(12), 128702 (2016)
May, R.M.: Uses and abuses of mathematics in biology. Science. 303, 790–793 (2004)
Kermack, W.O., McKendrick, A.G.: A contribution to the mathematical theory of epidemics. Proc. Roy. Soc. Lond. A. 115, 700–721 (1927)
Ribeiro, R.M., Bonhoeffer, S.: Production of resistant HIV mutants during antiretroviral therapy. Proc. Natl. Acad. Sci. U. S. A. 97(14), 7681–7686 (2000)
Karlin, S., Taylor, H.E.: A First Course in Stochastic Processes. Academic Press, New York (1975)
Van Kampen, N.G.: Stochastic Processes in Physics and Chemistry, 3rd edn. North Holland, Amsterdam (2007)
Antal, T., Scheuring, I.: Fixation of strategies for an evolutionary game in finite populations. Bull. Math. Biol. 68(8), 1923–1944 (2006)
Nåsel, I.: On the quasi-stationary distribution of the stochastic logistic epidemic. Math. Biosci. 156(1-2), 21–40 (1999)
Payn, B., Tanfer, K., Billy, J.O.G., Grady, W.R.: Men’s behavior change following infection with a sexually transmitted disease. Fam. Plan. Perspect. 29(4), 152–157 (1997)
Emlet, C.A.: An examination of the social networks and social isolation in older and younger adults living with HIV/AIDS. Health Soc. Work. 31(4), 299–308 (2006)
Zacks, S., Beavers, K., Theodore, D., Dougherty, K., Batey, B., Shumaker, J., Galanko, J., Shrestha, R., Fried, M.W.: Social stigmatization and Hepatitis C virus infection. J. Clin. Gastroenterol. 40(3), 220–224 (2006)
Salathé, M., Khandelwal, S.: Assessing vaccination sentiments with online social media: implications for infectious disease dynamics and control. PLoS Comput. Biol. 7(10), e1002199 (2011)
Chunara, R., Andrews, J.R., Brownstein, J.S.: Social and news media enable estimation of epidemiological patterns early in the 2010 Haitian cholera outbreak. Am.J.Trop. Med. Hyg. 86(1), 39–45 (2012)
Keeling, M.J.: The effects of local spatial structure on epidemiological invasions. Proc. Biol. Sci. 266(1421), 859–867 (1999)
Ogura, M., Preciado, V.M.: Epidemic processes over adaptive state-dependent networks. Phys. Rev. E. 93(6), 062316 (2016)
Amaral, L.A., Scala, A., Barthelemy, M., Stanley, H.E.: Classes of small-world networks. Proc. Natl. Acad. Sci. U. S. A. 97(21), 11149–11152 (2000)
Albert, R., Barabási, A.L.: Statistical mechanics of complex networks. Rev. Mod. Phys. 74, 47–98 (2002)
Newman, M.E.J.: The structure and function of complex networks. SIAM Rev. 45(2), 167–256 (2003)
Barthelemy, M., Barrat, A., Pastor-Satorras, R., Vespignani, A.: Velocity and hierarchical spread of epidemic outbreaks in scale-free networks. Phys. Rev. Lett. 92(17), 178701 (2004)
Acknowledgements
This research was supported by FCT-Portugal through grants PTDC/EEI-SII/5081/2014, PTDC/MAT/STA/3358/2014, UID/BIA/04050/2013 and UID/CEC/50021/2013.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Copyright information
© 2017 Springer Nature Singapore Pte Ltd.
About this chapter
Cite this chapter
Pacheco, J.M., Van Segbroeck, S., Santos, F.C. (2017). Disease Spreading in Time-Evolving Networked Communities. In: Masuda, N., Holme, P. (eds) Temporal Network Epidemiology. Theoretical Biology. Springer, Singapore. https://doi.org/10.1007/978-981-10-5287-3_13
Download citation
DOI: https://doi.org/10.1007/978-981-10-5287-3_13
Published:
Publisher Name: Springer, Singapore
Print ISBN: 978-981-10-5286-6
Online ISBN: 978-981-10-5287-3
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)