Abstract
Our body possesses three body axes, anteroposterior, dorsoventral, and left-right (L-R) axes. L-R asymmetry is achieved by three consecutive steps: symmetry breaking at the node, differential patterning of the lateral plate by a signaling molecule Nodal, and finally situs-specific organogenesis. Breaking of L-R symmetry in the mouse embryo takes place in the ventral node, where two types of cilia are found. Whereas centrally located motile cilia generate a leftward fluid flow, peripherally located immotile cilia sense a flow-dependent signal. Although Ca2+ signaling is implicated in flow sensing, it is still not clear what triggers Ca2+ signaling, a determinant molecule transported by the flow or mechanical force induced by the flow.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Most of visceral organs in vertebrates including the human are left-right (L-R) asymmetric in their position or shape. The process by which L-R asymmetry is generated can be divided into three steps (Fig. 7.1):
Three steps underlying the generation of L-R asymmetry. Three steps that contribute to the generation of L-R asymmetry are shown: (1) symmetry breaking, (2) molecular patterning of the LPM, and (3) asymmetric organogenesis. The developmental stage (E, embryonic day) corresponding to each step in the mouse is indicated on the left
-
1.
The initial breaking of L-R symmetry, which occurs in or near the node and at the late neural-fold stage
-
2.
Transfer of an L-R-biased signal(s) from the node to the lateral plate mesoderm (LPM) , which leads to L-R asymmetric expression of signaling molecules such as the transforming growth factor-β (TGF-β)-related proteins Nodal and Lefty on the left side of the LPM
-
3.
L-R asymmetric morphogenesis of visceral organs induced by these signaling molecules
2 Symmetry Breaking by Motile Cilia and Fluid Flow
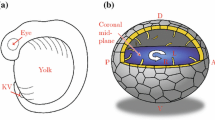
The breaking of L-R symmetry takes place in the node, an embryonic midline structure located at the anterior tip of the primitive streak in mouse embryos (Fig. 7.2). At the central region of the node, there are about 200 motile cilia that protrude from the ventral side of the node into the node cavity [1] (Fig. 7.2) and rotate in the clockwise direction (when viewed from the ventral side) at a speed of 600 rpm [2]. This rotational movement of the cilia generates the leftward laminar flow of extraembryonic fluid in the node cavity [2], occurs at a speed of ~15–20 μm/s. This leftward fluid flow in the node, referred to as nodal flow, is responsible for symmetry breaking. Many mutant mice that lack nodal flow because the node cilia are either missing or immotile have been identified, all of which exhibit aberrant L-R patterning of the LPM. Furthermore, L-R patterning of the embryo can be reversed when the direction of the flow was experimentally reversed by imposing the rightward artificial flow [3], establishing that the direction of the flow determines L-R.
Cilia and fluid flow in the node. A lateral view of the embryonic day 8.0 mouse embryo (a). Note that the node is located at the midline. Left-right (L-R) and anteroposterior (A-P) orientations are indicated. A ventral view of the mouse node at lower magnification (b). The red arrow denotes the leftward flow of extraembryonic fluid. A scanning electron micrograph showing that each cell on the ventral side of the mouse node has a monocilium (c)
How is the unidirectional fluid flow generated by rotational movement of the cilia? Hydrodynamic principles predict that the cilia can generate a unidirectional flow if they are tilted toward a specific direction. When the cilia move closer to the surface, the movement of fluid near the surface will be restricted as a result of the “no-slip boundary effect.” Conversely, when the cilia move away from the surface, they move the neighboring fluid more effectively. If cilia are tilted toward the posterior side, they will be moving toward the right when they come close to the surface and toward the left when they are far from the surface, thus generating a leftward flow. Observation of these rotating cilia by high-speed video microscopy revealed that they are indeed tilted posteriorly at an average angle of 30° [4, 5]. Recent evidence [6] suggests that, in addition to the “no-slip boundary effect,” intrinsic asymmetry in rotational stroke may also help generating the unidirectional flow.
Since the L-R axis is the last axis to be determined, symmetry breaking of L-R axis must be achieved by utilizing preexisting positional cues. In fact, two preexisting positional cues are reflected in the cilia: The A-P and D-V axes are thus represented by the posterior tilt and ventral protrusion of the cilia, respectively (Fig. 7.3). The node cilia thus generate the leftward flow by making use both of the preexisting positional cues and their structural chirality.
L-R symmetry breaking by preexisting information. Each node cilium (red bars on left) is posteriorly tilted, likely because the basal body (green) is posteriorly shifted within the cell (blue). The cilium protrudes from the cell toward the ventral side of the embryo and rotates in a clockwise direction when viewed from the ventral side. Anteroposterior (A-P) and dorsoventral (D-V) orientations are indicated. A schematic representation of a transverse section of a cilium, revealing its chiral structure, is shown on the right. The cilium contains nine pairs of microtubules (green) as well as inner and outer arms of dynein (pink)
How is A-P information translated into the posterior tilt of the node cilia? Given its similarity to positioning of the hair in the Drosophila wing, a mechanism resembling the planar cell polarity (PCP) pathway involving noncanonical Wnt signaling [7] seems to underlie positioning of the node cilia. Thus, some of the PCP core proteins such as Prickle2 and Vangl1 are localized to the anterior side of node cells [8, 9], whereas Dvl protein is localized to the posterior side [10] (Fig. 7.3). However, it remains unknown what positional cue is responsible for the polarized localization of these PCP core proteins.
3 Sensing of the Fluid Flow by Immotile Cilia
In addition to the motile cilia, there are immotile cilia in the node [11, 12] (Fig. 7.4). Cells located at the central region of the node (pit cells) possess motile cilia, which generate the fluid flow. On the other hand, most cells located at the edge of the node (crown cells) possess immotile cilia [13]. Immotile cilia act as sensors of the fluid flow [13]. Mutant mouse embryo that lack all cilia including those at the node, such as Kif3a−/−mouse embryos, fail to develop nodal flow and show L-R defects [14]. Such cilium-less embryos are also unable to respond to the artificial flow. However, when immotile cilia are restored in crown cells, the resulting embryo can now respond to the artificial flow [13], demonstrating that immotile cilia sense the flow.
Two types of cilia in the node, motile and immotile. Two types of ciliated cell are present in the node: Those located centrally (green) have motile cilia that generate nodal flow, whereas those located peripherally (pink) possess immotile cilia that sense the flow. Sensing of the flow requires ciliary localization of a Ca2+ channel, the Pkd2-Pkd1l1 complex. The flow-mediated signal results in degradation of Cerl2 mRNA. In this model, the flow would bend an immotile cilium on the left side
Sensing of the fluid flow by immotile cilia requires a Ca2+ channel composed of Pkd2 [15] and Pkd1l1 [16, 17]. Indeed, several Ca2+ signaling blockers have been shown to disrupt asymmetric gene expression in crown cells [13]. In particular, the effects of GdCl3 [an inhibitor of stretch-sensitive transient receptor potential (TRP) channels], 2-ABP [an inhibitor of the inositol 1,4,5-trisphosphate (IP3) receptor], and thapsigargin (an inhibitor of Ca2+-dependent ATPase activity in the endoplasmic reticulum) suggest involvement of a TRP-type channel such as Pkd2 and the IP3 receptor in the sensing of nodal flow. A mutation in Pkd2 that disrupts the ciliary localization of the encoded protein results in L-R defects similar to those of Pkd2−/−embryos [13, 16], suggesting that Pkd2, together with Pkd1l1, functions in the ciliary compartment of crown cells. Whereas Pkd2 encodes a Ca2+ channel with a short extracellular domain, Pkd1l1 possesses a much larger extracellular domain at its amino terminus. Pkd1l1 may be responsible for sensing of the flow signal and regulating Ca2+ channel activity of Pkd2. While oscillations of Ca2+ signaling with subtle L>R asymmetry were detected in the node [18], direct observation of L-R asymmetric Ca2+ signaling in crown cells has not been successful [13].
A long-standing question since the discovery of nodal flow concerns the action of the flow. Two models have been proposed (Fig. 7.5). According to the chemosensor model (Fig. 7.5a), the flow would transport an unknown molecule toward the left side of the embryo, which will eventually act as the L-R determinant. In an alternative model (two-cilia model or mechanosensor model; Fig. 7.5b), the embryo would sense the mechanical force generated by the flow. Several molecules have been proposed to be the determinant transported by the flow. However, none of them fulfill the requirements for the determinant. On the other hand, many lines of circumstantial evidence, including the recent observation that as few as two rotating cilia are sufficient for the breaking of L-R symmetry [19], favor the latter model. However, it is still not clear what exactly the immotile cilia sense during the symmetry-breaking event.
4 Readouts of the Flow
Cerl2 is the most immediate readout of the flow signal [19, 20]. Cerl2 encodes a Nodal antagonist, although its precise action is not clear. It is asymmetrically (R > L) expressed in crown cells, and its absence results in randomization of L-R decision making [21]. Whereas expression of Nodal is bilateral in crown cells, the R > L expression of Cerl2 renders Nodal activity in crown cells higher on the left side (Fig. 7.6). The Cerl2-generated asymmetry (R < L) of Nodal activity at the node closely correlates with the asymmetric pattern of Nodal expression in LPM [22]. Expression of Cerl2 is initially symmetric (R = L) at the early headfold stage, but it becomes R > L as the velocity of nodal flow increases, with expression on the left side being downregulated [19, 22]. Finally, Pkd2−/−Cerl2−/−double-mutant embryos manifest randomized Nodal expression in LPM, resembling the Cerl2 single mutant [13]. Therefore, Cerl2 is the main target of the flow signal.
L-R asymmetry of Cerl2 expression is generated at a posttranscriptional level [23], by degradation of Cerl2 mRNA via its 3′ untranslated region. Preferential decay of Cerl2 mRNA on the left is initiated by the leftward flow and further enhanced by the operation of Wnt-Cerl2 interlinked feedback loops, in which Wnt3 upregulates Wnt3 expression and promotes Cerl2 mRNA decay, whereas Cerl2 promotes Wnt degradation. Mathematical modeling and experimental data suggest that these feedback loops behave as a bistable switch that is able to amplify in a noise-resistant manner a small bias conferred by fluid flow.
5 Future Directions
Although rapid progress has been made in the last 20 years, many important questions remain unanswered. Firstly, how is A-P information translated into the posterior tilt of node cilia? Namely, what is the nature of the A-P information that polarizes node cells along the A-P axis? Secondly, how is the direction of rotation determined for node cilia? Thirdly, how does the nodal flow work? How do motile cilia sense the flow? Do they sense a signaling molecule that is transported by the flow or sense mechanical force? Fourthly, what is the precise role of Ca2+ signaling? How does Ca2+ signaling induce degradation of Cerl2 mRNA? Finally, to what extent is the mechanism for breaking of L-R symmetry conserved among species? L-R symmetry breaking does not appear to depend on cilia in Drosophila and snail [24]. Further development of various approaches (including genetic, cellular, biophysical, and mathematical) will be necessary to answer these questions.
References
Sulik K, et al. Morphogenesis of the murine node and notochordal plate. Dev Dyn. 1994;201(3):260–78.
Nonaka S, et al. Randomization of left-right asymmetry due to loss of nodal cilia generating leftward flow of extraembryonic fluid in mice lacking KIF3B motor protein. Cell. 1998;95(6):829–37.
Nonaka S, et al. Determination of left-right patterning of the mouse embryo by artificial nodal flow. Nature. 2002;418(6893):96–9.
Okada Y, et al. Mechanism of nodal flow: a conserved symmetry breaking event in left-right axis determination. Cell. 2005;121(4):633–44.
Nonaka S, et al. De novo formation of left-right asymmetry by posterior tilt of nodal cilia. PLoS Biol. 2005;3(8):e268.
Takamatsu A, et al. Asymmetric rotational stroke in mouse node cilia during left-right determination. Phys Rev E Stat Nonlin Soft Matter Phys. 2013;87(5):050701.
Klein TJ, Mlodzik M. Planar cell polarization: an emerging model points in the right direction. Annu Rev Cell Dev Biol. 2005;21:155–76.
Antic D, et al. Planar cell polarity enables posterior localization of nodal cilia and left-right axis determination during mouse and Xenopus embryogenesis. PLoS One. 2010;5(2):e8999.
Song H, et al. Planar cell polarity breaks bilateral symmetry by controlling ciliary positioning. Nature. 2010;466(7304):378–82.
Hashimoto M, et al. Planar polarization of node cells determines the rotational axis of node cilia. Nat Cell Biol. 2010;12(2):170–6.
McGrath J, et al. Two populations of node monocilia initiate left-right asymmetry in the mouse. Cell. 2003;114(1):61–73.
Tabin CJ, Vogan KJ. A two-cilia model for vertebrate left-right axis specification. Genes Dev. 2003;17(1):1–6.
Yoshiba S, et al. Cilia at the node of mouse embryos sense fluid flow for left-right determination via Pkd2. Science. 2012;338(6104):226–31.
Takeda S, et al. Left-right asymmetry and kinesin superfamily protein KIF3A: new insights in determination of laterality and mesoderm induction by kif3A−/− mice analysis. J Cell Biol. 1999;145(4):825–36.
Pennekamp P, et al. The ion channel polycystin-2 is required for left-right axis determination in mice. Curr Biol. 2002;12(11):938–43.
Field S, et al. Pkd1l1 establishes left-right asymmetry and physically interacts with Pkd2. Development. 2011;138(6):1131–42.
Kamura K, et al. Pkd1l1 complexes with Pkd2 on motile cilia and functions to establish the left-right axis. Development. 2011;138(6):1121–9.
Takao D, et al. Asymmetric distribution of dynamic calcium signals in the node of mouse embryo during left-right axis formation. Dev Biol. 2013;376(1):23–30.
Shinohara K, et al. Two rotating cilia in the node cavity are sufficient to break left-right symmetry in the mouse embryo. Nat Commun. 2012;3:622.
Schweickert A, et al. The nodal inhibitor coco is a critical target of leftward flow in Xenopus. Curr Biol. 2010;20(8):738–43.
Marques S, et al. The activity of the Nodal antagonist Cerl-2 in the mouse node is required for correct L/R body axis. Genes Dev. 2004;18(19):2342–7.
Kawasumi A, et al. Left-right asymmetry in the level of active Nodal protein produced in the node is translated into left-right asymmetry in the lateral plate of mouse embryos. Dev Biol. 2011;353(2):321–30.
Nakamura T, et al. Fluid flow and interlinked feedback loops establish left-right asymmetric decay of Cerl2 mRNA. Nat Commun. 2012;3:1322.
Okumura T, et al. The development and evolution of left-right asymmetry in invertebrates: lessons from Drosophila and snails. Dev Dyn. 2008;237(12):3497–515.
Acknowledgments
I thank current and former members of my laboratory for discussion as well as for providing illustrations. The work performed in my laboratory has been supported by CREST, Japan Science and Technology Corporation (JST), and by grants from the Ministry of Education, Culture, Sports, Science, and Technology of Japan.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited. The images or other third party material in this chapter are included in the work's Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work's Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Hamada, H. (2016). Roles of Motile and Immotile Cilia in Left-Right Symmetry Breaking. In: Nakanishi, T., Markwald, R., Baldwin, H., Keller, B., Srivastava, D., Yamagishi, H. (eds) Etiology and Morphogenesis of Congenital Heart Disease. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54628-3_7
Download citation
DOI: https://doi.org/10.1007/978-4-431-54628-3_7
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54627-6
Online ISBN: 978-4-431-54628-3
eBook Packages: MedicineMedicine (R0)