Abstract
In searching for remedial etiologies for congenital heart disease (CHD), we have focused on identifying interactive signaling pathways or “hubs” in which mutations disrupt fundamental cell biological functions in cardiac progenitor cells in a lineage-specific manner. Based on the frequency of heart defects seen in a clinical setting, we emphasize two signaling hubs – nodal kinases activated by extracellular ligands (e.g., periostin) and the cytoskeletal regulatory protein, filamin A (FLNA). We discuss them in the context of valve and septal development and the lineages which give origin to their progenitor cells. We also explore developmental windows that are potentially amenable to remedial therapy using homeostatic mechanisms like those revealed by a chimeric mice model, i.e., irradiated animals whose bone marrow had been reconstituted with GFP+ hematopoietic stem cells, that shows bone marrow-derived cells track to the heart, engraft, and give rise to bona fide fibroblasts. We propose to use this model to deliver genetic payloads or protein cargos during the neonatal period to override biochemical or structural deficits of CHD associated with valve and septal signaling hubs or fibroblast/myocyte interactions. Preliminary tests of the model indicate remedial potential for cardiac injuries.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Heart defects
- Valves
- Septa
- Matricellular proteins
- Periostin
- Cell signaling
- Kinases
- Filamin A
- Hematopoietic stem cells
- Fibroblast
- Lineage
- Genetic engineering
1 Introduction
1.1 Emerging Concepts
Etiologies of congenital heart defects have been difficult to identify and explain, often invoking discussions of genetics and environment or both. After 40+ years of searching for etiologies, especially ones that might be potentially remedial, a few conceptual observations are emerging (although by no means is there consensus). These include suppositions that:
-
1.
Genetics more than environment is responsible for CHD based on the ability to simulate human heart congenital defects in animal genetic models (see also Bhattacharya [1]).
-
2.
The evolution of genetic thought is toward trans-heterozygous, multigenetic interactions vs. single gene hits.
-
3.
Mutations in the downstream intracellular signaling targets of growth factor, transcription factors, or matricellular proteins are more likely to be the root cause of heart defects seen in children by pediatric cardiologists (as the loss of both alleles for upstream early regulatory genes is usually lethal).
-
4.
Intersecting regulatory networks or signaling hubs coordinate fundamental biological processes in cardiac progenitor cells in a lineage-restricted manner.
-
5.
Based on the frequency of different types of heart defects, the lineages most likely to be modified by genetic mutations or environmental stressors in CHD are the non-cardiomyocyte lineages, primarily fibroblasts.
1.2 Hub Hypothesis
In trying to provide answers to the questions most often asked by parents of children born with heart defects – “why did it happen?” and “what can be done about it?” – we have endeavored to integrate these five emerging concepts around two examples of central signaling hubs or platforms that intersect with multiple gene regulatory networks (like the spokes of a wheel) to regulate or “tune” behavioral changes in the progenitor cells engaged in valvuloseptal morphogenesis (or myocardial remodeling). In so doing, we propose that intersecting signaling hubs explain why so many different genes, if mutated or deleted, can engender similar anatomical dysmorphic phenotypes, e.g., ventricular septal defects. This suggests that there are only so many ways progenitor cells can respond to normal or abnormal signaling inputs: e.g., they can proliferate, activate, or suppress apoptotic pathways; transport ions; secrete, endocytose, adhere/migrate, and generate contractile forces; change polarity or shape; and differentiate. Thus, the conceptual appeal of common final pathways or intersecting signaling hubs/nodes is that they provide potential for exploring “shared” remedial therapies for CHD that do not require an individual approach for correcting each abnormal gene.
2 Searching for Candidate Signaling Hubs in Heart Development
2.1 Nodal Signaling Kinases
In patients with Down, Marfan, or Noonan Syndromes, there is increasing evidence that genes encoding nodal signaling kinases like FAK/AKTkt/PI3K, RAS, MEKK/ERK1/2, PTPN11, etc., are likely candidates for CHD if they are mutated or overexpressed [2–4]. Such genes are not usually lethal (as there exist molecular or functional redundancies), yet as indicated for syndromic heart defects, they have potential to change functional behaviors in progenitor cells that normally mold and remodel the simple tubular heart into a four-chambered organ. As indicated in Fig. 5.1, intracellular signaling kinases (and small regulatory GTPases) are the direct and indirect downstream targets of growth factors (e.g., TGFβ and BMP 2&4) that are normally secreted by the embryonic endocardium or myocardium [5], or in the case of extracellular ligands like matricellular proteins (periostin, the CCN family), they are secreted by the mesenchymal progenitor cells of valves and connective septa which are derived from both endocardium and epicardium [6]. In the case of matricellular and other extracellular proteins (Fig. 5.2), binding to integrin receptors triggers integrin-dependent, downstream signaling kinases/GTPases (FAK/AKT/PI3k) which activate effector mechanisms of growth, survival, and differentiation into the fibrous structures (valves and septa) that assure coordinated and unidirectional blood flow through the right and left sides of the developing heart. Epicardial-derived mesenchymal cells also express periostin as they invade the ventricular myocardium and, like endothelial-derived mesenchyme, secrete collagen and differentiate into ventricular connective tissue but also contribute to the parietal leaflets of the AV valves [7, 8]. Disruption of these signaling pathways by either silencing one or more of the kinases shown in Fig. 5.1, inhibiting β-integrin functions, or deleting the periostin gene itself resulted in septal defects, abnormal (poorly differentiated, hypertrophic) valves, arrhythmias associated with a reduction in the AV fibrous connective tissue, and a reduction in ventricular elastic modulus due to loss of interstitial collagen [9, 10]. These findings are consistent with the relevance of central or nodal signaling kinases to heart development and how different genes, if mutated or inhibited, could produce similar abnormal anatomical phenotypes as a result of either binding to, activating, or encoding a kinase or GTPase component of an interactive signaling pathway. Conceivably, these same interactive signaling pathways could also be used to explore remedial therapies for CHD. Any one or combination of the signaling kinases or effector proteins shown in Fig. 5.1 could be a candidate therapeutic target that could be used to bypass or circumvent a genetic or biochemical block associated with a particular CHD, if a way could be found to administer drugs or small molecules that silence, simulate, or activate them. Identifying an in utero approach using orally administered or injected signaling inhibitors, lithium, or retinoids is appealing but lacks target specificity (potentially engendering a broad spectrum of side effects), or it can create a catch-22 in the sense that treatments may have to begin too early, i.e., before it can be actually determined that a development defect is going to happen. Genetic rescues can identify and test candidate target mechanisms (e.g., breeding Noonan mice into an Erk null background) but have little practical remedial potential for preventing CHD.
(Left): Generalized model proposing the candidate genes for CHD. Based on syndromic mutations or overexpression, the model proposes that the most likely candidates are genes that encode components of a signaling hub such as the one shown for an extracellular ligand and membrane receptors. Their interaction activates a cascade of nodal kinases within valvuloseptal progenitor cells that regulate biological functions in progenitor cells that are normally associated with remodeling the primary heart into a four-chambered organ. (Right): Periostin-integrin model depicting downstream targets activated by periostin binding to specific β-integrin heterodimeric receptors expressed in valve and septal progenitor cells. Based on published results [6], periostin binding to integrin induces phosphorylation of p-FAK (a nodal kinase) which, in turn, activates different signaling pathways, p-AKT/PI3K and/or p-Erk, each with distinct biological activities; e.g., p-AKT/PI3K promotes hyaluronan (HA) secretion and enhances periostin expression, whereas Erk activates collagen secretion and survival (anti-apoptotic) mechanisms
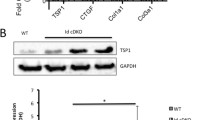
Coordinated activation of Filamin and p21-activated kinase 1 (PAK1) by periostin/β1-integrin signaling in E16.5 mouse AV cells. (a) Representative Western blot for periostin (PN), p-filamin A (ser 2152), and filamin A from lysates prepared from isolated mouse E16.5 AV valve cells, transfected with vector controls, PN cDNA, PN-siRNA, and CD44 siRNA for 48 h or treated with 5 μg/ml β1-integrin-antibody for 24 h. (b) Same protocol as A but lysates were immunoprecipitated with filamin A and binding proteins in the immunoprecipitate were immunoblotted. The figure shows that upon activation, phospho-PAK1 (ser144) binds to Filamin (and transfers its phosphate to activate the FLNA protein). Because silencing CD44 prevented binding of PAK1 to FLNA, we have suggested that CD44 forms a signaling complex with periostin-integrin-FAK that is required for activating FLNA through PAK1
Another plus for interactive signaling hubs is their potential to reveal a new or unexpected “spoke” whereby genetic, protein, or biochemical connections to signaling pathways are not readily apparent or known, e.g., hyaluronan synthetase 2 (Has2). As shown by Misra et al. [6], Has2 is activated by nodal kinases that are regulated by periostin-integrin-linked signaling [6]. Phosphorylation of Has2 leads to secretion of hyaluronan (HA) (Fig. 5.1, right) which, in addition to its osmotic properties, also binds to CD44, a tyrosine kinase receptor for HA that is expressed by valvular-septal mesenchymal progenitor cells [11]. Binding of HA to CD44 activates some of the same kinases also activated by periostin-integrin signaling, thereby triggering positive feedback loops that can amplify or sustain shared biological effects of periostin including formation, migration, and survival of progenitor valvuloseptal mesenchyme as well as a positive feedback on the secretion of periostin itself (Fig. 5.1) [6, 11]. CD44 signaling also appears to be part of a signaling complex revealed by immunoprecipitation that is required for periostin to activate another candidate signaling hub – filamin A (Fig. 5.2).
2.2 Filamin A
Filamin A (FLNA) is a homo- or heterodimeric Y-shaped cytoplasmic protein in which each main chain consists of an actin-binding region at the amino terminus, a core of 24 highly homologous Ig-like repeats followed by a carboxyl integrin-binding domain. The most widely studied function of FLNA is its ability, upon activation, to self-assemble into signaling scaffolds or to cross-link cortical actin filaments into a dynamic three-dimensional structure [12]. In the developing heart, FLNA can promote or suppress cell processes important for heart development as evident that when the gene is deleted, lethality ensues. Through its role as an actin-binding protein that anchors various transmembrane proteins to the cytoskeleton, it can generate contractile forces to support migration and fusion of the paired midline AV mesenchymalized “cushions” to form the AV septum [13]. Upon activation, the scaffolds assembled by FLNA form docking sites for second messengers like smad 2/3 which are critical for transmitting TGFβ signaling in heart development [14, 15]. While FLNA is also expressed in the brain and other organs, it is specifically expressed in the heart in valvuloseptal mesenchyme derived from the endocardium and epicardium [16]. The null phenotype, although lethal at ED15.0, is consistent with the pattern of FLNA expression in valvuloseptal primordia, i.e., there are septal and valve defects at all axial levels: atrial, atrioventricular, ventricular, and outlet [15–17].
FLNA itself responds to many signaling “inputs” that can activate FLNA or modify its binding to actin [18], including Ras and Rho kinase. Immunoprecipitation studies confirmed that FLNA is also a downstream target of kinases induced by periostin-integrin signaling that promote the binding of PAK-1 to FLNA, resulting in its activation (Fig. 5.2). A large primary atrial septal defect is seen in periostin null mice [19] that is also seen in FLNA nulls, suggesting an interaction between “signaling hubs” that (normally) promote: (1) fusion of paired mesenchymal primordia (often called “cushions”) to form septal and valvular primordia and (2) remodeling of the fused cushion primordia during fetal and early postnatal life into mature leaflets or tendon-like septal structures.
Loss-of-function, point mutations in FLNA have also been found in patients with mitral valve disease (e.g., prolapse), all of which occurred in the actin-binding domains of FLNA and similarly effected cell signaling pathways or cytoskeletal organization [20]. For example, a G288R and P637Q point disrupted a signaling network that balanced RhoA and Rac1 GTPases activities and correlated directly with inhibited cell spreading, migration, and contractile force generation [21]. The G288R and P637Q mutation also interfered with FLNA’s normal capabilities to promote the expression and transport of beta integrins to the cell surface or bind a variety of intracellular target proteins including vimentin and transglutaminase 2 (TG2) [16, 22]. We found that TG2 homeostatically functions to covalently link serotonin to FLNA enhancing its potential to bind cytoskeletal F-actin and generate contractile forces related to or required for remodeling of valve primordia into compacted, sculpted leaflets [16].
FLNA expression, like periostin, peaks in the postnatal period but is barely detectable in adult heart tissues unless there is an injury ([17, 19, 23]). These findings have three implications: (1) heart development is not over at birth; (2) the postnatal period may still be an open window to explore remedial therapies for CHD, particularly for valve and septal defects [24, 41]; and (3) mutations in genes that cause abnormal structural or functional changes in heart tissues may not always be immediately visible at birth but progressively appear over time [25]. Thus, some “adult cardiovascular degenerative diseases” may actually have an embryonic developmental etiology [26].
2.3 Relevance of Signaling Hubs to CHD
Shared (interconnected) protein signaling hubs point to a multiplicity of ways by which seemingly unrelated genes (including ones yet to be identified) can converge to engender anatomically similar developmental heart defects [27, 28]. Understanding these mechanistic relationships will be important as they have realistic remedial potential for identifying candidate approaches for preventing, modifying, or even reversing some of the clinical consequences of CHD, particularly if they are recognized early in the neonatal period. For example, AV valves in which there is diminished function of FLNA are enlarged at birth (and mesenchymal-like) but have elevated phosphorylation of ERK1/2 (16, unpublished data) which progressively leads to a more pronounced degenerative, myxomatous-like phenotype. A potential remedial therapy would be to find a way to attenuate ERK signaling and assess whether progression to a degenerative phenotype could be delayed or even reversed. A precedent for this would be the fibrillin-1 knockin model of Marfan syndrome in which the valves become myxomatous or the Loeys-Dietz mice with mutated TGFβ receptors in which the aortic root is dilated and prone to aneurysm [29, 30]. ERK 1/2 pathways are used in both syndromes to transduce elevated, noncanonical TGFβ signaling. Pharmacological approaches (e.g., Losartan) to blunt ERK signaling in both syndromes have improved their cardiovascular function and tissue structural integrity [30]. In a related fashion, understanding that TG2 promotes serotonylation of FLNA (and its binding affinity for actin), altering serotonin uptake, and/or synthesis could also prove to be new remedial approaches for treating valve and septal structural defects during the neonatal period when they normally complete their maturation. The key would be to identify a means or route for delivering these potential remedial therapies to the “right cell targets” at the “right time”.
3 Lineage Is a Key to Remedial Therapy
Finding the “right cell targets” for remedial therapy is a question of lineage. Any hope of efficacious remedial treatments would seemingly require that treatments be directed to those cell populations that are normally involved in valve and septal formation and, if mutated, result in CHD. In heart development, it is the non-myocyte – mostly fibroblast – populations of the heart that appear to be the critical players in valvuloseptal morphogenesis. They are the only heart progenitor cells to express the extracellular signaling ligands like periostin or the cytoskeletal regulatory protein like FLNA. The progenitors of fibroblasts, as noted above, are the mesenchymal stem cells (or “cushion” cells) derived from the transformation of two epithelia: endocardium and epicardium. Cells derived from either lineage have the potential to differentiate into fibroblasts by autonomously secreting periostin [31, 32]. Thus, cardiac fibroblasts are derived from at least two origins which are carried over from intrauterine to postnatal and adult life [32]. This raises two questions: (1) Are new fibroblast progenitor cells added to the heart after birth? (2) If so, are the same signals (e.g., periostin) used to direct their progression into a cardiac fibroblast lineage during embryonic life also used postnatally? The answers to both questions appear to be “yes.”
3.1 Postnatal Origin of Cardiac Fibroblasts
Recent single cell engraftment experiments indicate that in postnatal and adult life, cardiac fibroblasts are also derived from bone marrow or from pericytes that express the hematopoietic stem cell marker – CD45 [33–38]. In these experiments, adult mice were lethally irradiated and a single (or clone) of a rigorously isolated wild-type CD45+ hematopoietic stem cell (HSC) carrying a green fluorescent protein (GFP) marker was injected into their tail vein. Mice which survived clearly had received a true multipotential stem cell capable of restoring the blood cell lineages. We then asked if any of the original clones of CD45+/GFP+ HSCs left the bone marrow and engrafted elsewhere. They did. We found that CD45+/GFP+ cells migrate and engraft in several organs including the heart. Specifically they engrafted into the inlet and outlet valves, ventricular fibrous interstitium, and as pericytes surrounding coronary microvasculature [36, 37]. In addition to the CD45 marker and GFP, they also expressed fibroblast markers, e.g., collagen-1, HSP47, vimentin, discoid domain receptor 2 (DDR2), and, importantly, periostin [33, 37]. This suggested to us that cardiac fibroblasts are a renewable cell population that can be replenished homeostatically by circulating progenitor cells of bone marrow, HSC origin. Based on marker expression, monocytes are probably the immediate circulating progenitors of the CD45+/collagen I+ cells that engraft into the heart, probably as “blank” cells that differentiate into fibroblasts [38, 39]. Importantly, their numbers increase in the valves and ventricular interstitium significantly if the heart was injured by coronary ligation or cryoablation, indicating that they are also a population of fibrogenic precursors that can respond dynamically to injury or inflammatory signals [38, 40]. In quantitative terms, bone marrow-derived cells accounted for 20–30 % of fibroblasts in normal adult myocardial tissue [36, 37, 39]. In contrast, we found GFP+ label in myocytes at exceedingly low frequency [36] making this approach suitable for a targeted assessment of non-myocyte contributions to cardiac structure and function. It is important to recall, in this context, that not all lineage markers of hematopoietic cell sources (e.g., CD34) are applicable across species.
3.2 A Strategy to Use Fibroblast Progenitors to Carry Genetic Payloads
The bone marrow origin of postnatal cardiac fibroblasts does not in any way exclude new fibroblasts arising by proliferation from lineages carried over from intrauterine life. However, those fibroblasts would be expected to continue to carry forward any mutations or other functional or biochemical deficits from embryonic life, whereas those derived from bone marrow provide an opportunity to be isolated and genetically or pharmacologically manipulated (or reengineered) and then returned to the marrow. Thus, we propose to use these fibrogenic CD45+ progenitors to carry genetic payloads (or protein cargos) to sites where fibrous valvuloseptal or interstitial tissues are underdeveloped or hyperplastic or where maturation (e.g., valves) has been delayed. Genetic reengineering theoretically could be done to benefit the health of the newborn with a cardiac developmental defect at any time or age, but given its potential for regeneration, earlier in the postnatal period would appear to be the “right time” to implement remedial therapies before “wet cement becomes hardened” [41].
3.2.1 This Strategy Calls for a Conceptual Revision in Our Thinking About Fibroblasts
This strategy calls for a conceptual revision in our thinking about fibroblasts that recognizes that they are a renewable cell source that can form a community of active modulators of cell behaviors that can change cardiac function. The full story of cardiac fibroblasts is yet to be realized as to their function in health and disease including CHD. Already much is being learned about their potential for repair through homotypic interactions or heterotypic interactions with myocytes. Modes of contact between fibroblasts or fibroblasts and myocytes include connexin-based, gap junctions through which ions or small molecules ( microRNAs) can be transferred [42], tunneling nanotubes [43] through which mitochondria can be exchanged, or by paracrine secretions of cytokines, growth factors, or matricellular proteins that facilitate their engraftment, proliferation, and/ or differentiation [44].
4 Remedial Therapies: Delivering Genetic “Payloads”
Our basic premise is that bone marrow-derived fibroblasts make homeostatic contributions to the non-myocyte cells of adult heart valve and septal connective tissues. We propose to exploit our findings using the single cell engraftment model to deliver relevant payloads to connective tissues of hearts with visible congenital malformations, e.g., valves, septa, hypoplastic ventricles, or non-compacted myocardium in which growth has been compromised. We include myocardial growth malformations because there is emerging evidence that fibroblasts establish heterotypic contacts in vivo with myocytes that can affect myocyte growth or electrophysiology [45]. For example, isolated CD45+ bone marrow cells could be genetically engineered to secrete growth factors like neuregulin which has been shown to reactivate cell cycling in mononucleated cardiomyocytes [46]. For hypoplastic valve or septal tissues, we have already suggested (see above) several genetic or small molecule pharmacological remedial approaches that would be amenable to using CD45 cells for their delivery; for example, viral transduction of CD45+ cells could be used to overexpress or suppress any of kinase or integrin genes known to promote or inhibit fibroblast differentiation [47, 48], focusing on Cre-promoters that allow sharp delineation from myocytes lineages [49, 50]. Antennapedia internalization sequences could also be used to introduce a wide range of “cargo” peptides into isolated CD45 cells.
4.1 Preliminary Studies
While we have shown using chimeric mice, i.e., irradiated animals whose bone marrow had been reconstituted with GFP+ hematopoietic stem cells, that it is possible to track bone marrow-derived cells that give rise to bona fide fibroblasts in the heart, it remains to be determined whether they can effectively be transduced to carry a cargo that can have a remedial effect in injured heart tissues. As proof of concept, we prepared a lentivirus periostin-silencing (shRNA) vector directed against the 3′ end of the periostin transcript that was shown to block periostin protein expression (>95 %) in vitro [6, 19]. These silencing or empty control vectors were injected directly into the bone marrow minutes prior to an acute myocardial “cryoinjury” administered by a liquid nitrogen cooled microprobe directly to the left ventricle of anesthetized mice. This procedure allows for a consistent injury of known size and location within 4 days. As shown in Fig. 5.3, an externally visible scar (blue-white in color) developed within a week in control cryoinjured mice, whereas in experimental cryoinjured mice in which periostin-silencing vectors were injected into the bone marrow, the scar was barely visible and the tissue appeared healthy and vascularized. Echocardiographic analyses were performed at weekly intervals. Cryoinjured mice that received bone marrow injections of the periostin-silencing vector exhibited performance metrics (e.g., LV vol., ejection fraction) comparable to normal mouse baseline values, whereas those that received the empty control vector exhibited significant reduction of all baseline values. While preliminary, we expect that bone marrow transduction will target cells that potentially can engraft into the heart and modify connective tissue or myocardial remodeling to benefit patient health.
Cryoinjured adult hearts treated with lentiviral siRNA vectors to silence periostin injected into the bone marrow. (a) arrows denote a typical injury site 4 days after cryoablation; after 8 weeks, a large scar (white color) is visible on the left ventricular surface which when examined histologically reveals extensive fibrosis (blue in Masson stain, muscle is red). (b) left panel, empty vector control showing developing scar is turning bluish due to lost vascularity and fibrosis; right panel is a heart after bone marrow injection with periostin siRNA vector. The injury site is barely detectable and well vascularized. Performance metrics (bottom) indicate that silencing periostin gave results similar to those of baseline controls
Finally, as developmental biologists searching for remedial etiologies and therapies, we believe the crux of the issue lies in shifting our focus toward understanding that:
-
1.
The later periods of fetal and neonatal cardiovascular development present a window to initiate remedial therapies.
-
2.
The interactions between multiple signaling hubs will become a priority rather than continuing to investigate early events or single genes.
-
3.
The heart is an integrated organ in which lineages can provide insight for improving therapeutic outcomes for CHD through genetic or pharmacological manipulation.
-
4.
The complexity and balance of signals that affect processes controlling the formation of the heart may also come into play after birth or into adult life, where their engineered re-expression can be used to enhance or awaken the ability of heart cells to adapt to pathophysiological stimuli engendered during development.
References
Bentham J, Bhattacharya S. Genetic mechanisms controlling cardiovascular development. Ann N Y Acad Sci. 2008;1123:10–9.
Wu H, Kao SC, Barrientos T, Baldwin SH, Olson EN, Crabtree GR, Zhou B, Chang CP. Down syndrome critical region-1 is a transcriptional target of nuclear factor of activated T cells-c1 within the endocardium during heart development. J Biol Chem. 2007;282(42):30673–9.
Holm TM, Habashi JP, Doyle JJ, et al. Noncanonical TGFbeta signaling contributes to aortic aneurysm progression in Marfan syndrome mice. Science. 2011;332:358–61.
Nava C, Hanna N, Michot C, et al. Cardio-facio-cutaneous and Noonan syndromes due to mutations in the RAS/MAPK signalling pathway: genotype-phenotype relationships and overlap with Costello syndrome. J Med Genet. 2007;44:763–71.
Person AD, Klewer SE, Runyan RB. Cell biology of cardiac cushion development. Int Rev Cytol. 2005;243:287–335.
Ghatak S, Misra S, Norris R, Moreno-Rodriquez RA, Levine RA, Hascall VC, Markwald RR. Periostin induces intracellular cross talk between kinases and hyaluronan in atrioventricular valvulogenesis. J Biol Chem. 2014;289(12):8545–61.
Lie-Venema H, Eralp I, Markwald RR, et al. Periostin expression by epicardium-derived cells is involved in the development of the atrioventricular valves and fibrous heart skeleton. Differentiation. 2008;76:809–19.
Wessels A, van den Hoff MJ, Adamo RF, Phelps AL, Lockhart MM, Sauls K, Briggs LE, Norris RA, van Wijk B, Perez-Pomares JM, et al. Epicardially derived fibroblasts preferentially contribute to the parietal leaflets of the atrioventricular valves in the murine heart. Dev Biol. 2012;366:111–24.
Snider P, Hinton RB, Moreno-Rodriguez RA, et al. Periostin is required for maturation and extracellular matrix stabilization of noncardiomyocyte lineages of the heart. Circ Res. 2008;102:752–60.
Norris RA, Moreno-Rodriguez R, Hoffman S, Markwald RR. The many facets of the matricellular protein periostin during cardiac development, remodeling, and pathophysiology. J Cell Commun Signal. 2009;3:275–86.
Nakamura T, Colbert M, Krenz M, et al. Mediating ERK 1/2 signaling rescues congenital heart defects in a mouse model of Noonan syndrome. J Clin Invest. 2007;117:2123–32.
Camenisch TD, Spicer AP, Brehm-Gibson T, et al. Disruption of hyaluronan synthase-2 abrogates normal cardiac morphogenesis and hyaluronan-mediated transformation of epithelium to mesenchyme. J Clin Invest. 2000;106:349–60.
Feng Y, Walsh CA. The many faces of filamin: a versatile molecular scaffold for cell motility and signalling. Nat Cell Biol. 2004;6:1034–8.
Sasaki A, Masuda Y, Ohta Y, Ikeda K, Watanabe K. Filamin associates with Smads and regulates transforming growth factor-beta signaling. J Biol Chem. 2001;276:17871–7.
Feng Y, Chen MH, Moskowitz IP, et al. Filamin A (FLNA) is required for cell-cell contact in vascular development and cardiac morphogenesis. Proc Natl Acad Sci U S A. 2006;103:19836–41.
Sauls K, de Vlaming A, Harris BS, Markwald RR, Norris RA, et al. Developmental basis for filamin-A-associated myxomatous mitral valve disease. Cardiovasc Res. 2012;96:109–19.
Norris RA, Moreno-Rodriguez R, Wessels A, et al. Expression of the familial cardiac valvular dystrophy gene, filamin-A, during heart morphogenesis. Dev Dyn. 2010;239:2118–27.
Vadlamudi RK, Li F, Adam L, et al. Filamin is essential in actin cytoskeletal assembly mediated by p21-activated kinase 1. Nat Cell Biol. 2002;4:681–90.
Norris RA, Moreno-Rodriguez RA, Sugi Y, et al. Periostin regulates atrioventricular valve maturation. Dev Biol. 2008;316:200–13.
Kyndt F, Gueffet JP, Probst V, et al. Mutations in the gene encoding filamin A as a cause for familial cardiac valvular dystrophy. Circulation. 2007;115:40–9.
Duval D, Lardeux A, Le Tourneau T, et al. Valvular dystrophy associated filamin A mutations reveal a new role of its first repeats in small-GTPase regulation. Biochim Biophys Acta. 2014;1843(2):234–44.
MacPherson M, Fagerholm SC. Filamin and filamin-binding proteins in integrin-regulation and adhesion. Focus on: “Filamin A is required for vimentin-mediated cell adhesion and spreading”. Am J Physiol: Cell Physiol. 2010;298:C206–8.
Norris RA, Borg TK, Butcher JT, Baudino TA, Banerjee I, Markwald RR. Neonatal and adult cardiovascular pathophysiological remodeling and repair: developmental role of periostin. Ann N Y Acad Sci. 2008;1123:30–40.
Porrello ER, Mahmoud AI, Simpson E, Hill JA, Richardson JA, Olson EN, Sadek HA. Transient regenerative potential of the neonatal mouse heart. Science. 2011;331(6020):1078–80.
Markwald RR, Norris RA, Moreno-Rodriguez R, Levine RA. Developmental basis of adult cardiovascular diseases: valvular heart diseases. Ann N Y Acad Sci. 2011;1188:177–83.
Judge DP, Markwald RR, Hagège AA, Levine RA. Translational research on the mitral valve: from developmental mechanisms to new therapies. J Cardiovasc Transl Res. 2011;4:699–701.
Chen PC, Wakimoto H, Conner D, Araki T, Yuan T, Roberts A, Seidman C, Bronson R, Neel B, Seidman JG, Kucherlapati R. Activation of multiple signaling pathways causes developmental defects in mice with a Noonan syndrome–associated Sos1 mutation. J Clin Invest. 2010;120(12):4353–65.
Lage K, Greenway SC, Rosenfeld JA, Wakimoto H, Gorham JM, Segrè AV, Roberts AE, Smoot LB, Pu WT, Pereira AC, Mesquita SM, Tommerup N, Brunak S, Ballif BC, Shaffer LG, Donahoe PK, Daly MJ, Seidman JG, Seidman CE, Larsen LA. Genetic and environmental risk factors in congenital heart disease functionally converge in protein networks driving heart development. Proc Natl Acad Sci U S A. 2012;109(35):14035–40.
Habashi JP, Doyle JJ, Holm TM, Aziz H, Schoenhoff F, Bedja D, Chen Y, Modiri AN, Judge DP, Dietz HC. Angiotensin II type 2 receptor signaling attenuates aortic aneurysm in mice through ERK antagonism. Science. 2011;332:361–5.
Davis F, Rateri DL, Daugherty A. Aortic aneurysms in Loeys-Dietz syndrome – a tale of two pathways? J Clin Invest. 2014;124(1):79–81. See comment in PubMed Commons below.
Norris RA, Potts JD, Yost MJ, et al. Periostin promotes a fibroblastic lineage pathway in atrioventricular valve progenitor cells. Dev Dyn. 2009;238:1052–63.
Snider P, Standley KN, Wang J, Azhar M, Doetschman T, Conway SJ. Origin of cardiac fibroblasts and the role of periostin. Circ Res. 2009;105:934–47.
Ebihara Y, Masuya M, Larue AC, et al. Hematopoietic origins of fibroblasts: II. In vitro studies of fibroblasts, CFU-F, and fibrocytes. Exp Hematol. 2006;34:219–29.
Ogawa M, LaRue AC, Mehrotra M. Hematopoietic stem cells are pluripotent and not just “hematopoietic”. Blood Cells Mol Dis. 2013;51(1):3–8.
Visconti RP, Markwald RR. Recruitment of new cells into the postnatal heart: potential modification of phenotype by periostin. Ann N Y Acad Sci. 2006;1080:19–33.
Visconti RP, Ebihara Y, LaRue AC, Markwald R, et al. An in vivo analysis of hematopoietic stem cell potential: hematopoietic origin of cardiac valve interstitial cells. Circ Res. 2006;98:690–6.
Hajdu Z, Romeo SJ, Fleming PA, Markwald RR, Visconti RP, Drake CJ. Recruitment of bone marrow-derived valve interstitial cells is a normal homeostatic process. J Mol Cell Cardiol. 2011;51:955–65.
Lee R, Perry B, Heywood J, Reese C, Bonner M, Hatfield CM, Silver RM, Visconti RP, Hoffman S, Tourkina E. Caveolin-1 regulates chemokine receptor 5-mediated contribution of bone marrow-derived cells to dermal fibrosis. Front Pharmacol. 2014;5:140.
Mollmann H, Nef HM, Kostin S, von Kalle C, Pilz I, Weber M, Schaper J, Hamm CW, Elsasser A. Bone marrow-derived cells contribute to infarct remodelling. Cardiovasc Res. 2006;71:661–71.
Haudek SB, Ying X, Huebener P, Lee JM, Signe C, Crawford JR, Pilling D, Gomer RH, Trial JA, Frangogiannis NG, Entman ML. Bone marrow-derived fibroblast precursors mediate ischemic cardiomyopathy in mice. Proc Natl Acad Sci U S A. 2006;103(48):18284–1828.
Porrello ER, Olson EN. A neonatal blueprint for cardiac regeneration. Stem Cell Res. 2014;13(3 Pt B):556–70.
Ongstad EL, Gourdie RG. Myocyte-fibroblast electrical coupling: the basis of a stable relationship? Cardiovasc Res. 2012;93:215–7.
Ma Z, Yang H, Liu H, Xu M, Runyan RB, Eisenberg CA, Markwald RR, Borg TK, Gao BZ. Mesenchymal stem cell-cardiomyocyte interactions under defined contact modes on laser-patterned biochips. PLoS One. 2013;8(2):e56554.
Frangogiannis NG. Matricellular proteins in cardiac adaptation and disease. Physiol Rev. 2012;92:635–88.
Kohl P, Gourdie RG. Fibroblast-myocyte electrotonic coupling: does it occur in native cardiac tissue? J Mol Cell Cardiol. 2014;70:37–46.
Bersell K, Arab S, Haring B, Kuhn B. ErbB4 signaling induces cardiomyocyte proliferation and repair of heart injury. Cell. 2009;138:257–70.
Haudek SB, Gupta D, Dewald O, Schwartz RJ, Wei L, Trial J, Entman ML. Rho kinase-1 mediates cardiac fibrosis by regulating fibroblast precursor cell differentiation. Cardiovasc Res. 2009;83(3):511–8.
Ieda M, Tsuchihashi T, Ivey KN, Ross RS, Hong TT, Shaw RM, Srivastava D. Cardiac fibroblasts regulate myocardial proliferation through beta1 integrin signaling. Dev Cell. 2009;16:233–24.
Misra S, Hascall VC, Karamanos NK, Markwald RR, Ghatak S. Delivery systems targeting cancer at the level of ECM. In: Karamanos N, editor. Extracellular matrix: pathobiology and signaling. Berlin: DeGruyter; 2012. p. 865–83.
Lindsley A, Snider P, Zhou H, et al. Identification and characterization of a novel Schwann and outflow tract endocardial cushion lineage-restricted periostin enhancer. Dev Biol. 2007;307:340–55.
Acknowledgments
Funding for this work was provided by the National Institutes (HL 33756; NIGMS103342, NIGMS 103444-06), the National Science Foundation (EPS0903795), the American Heart Association (11SDG5270006), and the Leducq Foundation, Paris, France (07CVD04).
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited. The images or other third party material in this chapter are included in the work's Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work's Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Markwald, R.R., Ghatak, S., Misra, S., Moreno-Rodríguez, R.A., Sugi, Y., Norris, R.A. (2016). Congenital Heart Disease: In Search of Remedial Etiologies. In: Nakanishi, T., Markwald, R., Baldwin, H., Keller, B., Srivastava, D., Yamagishi, H. (eds) Etiology and Morphogenesis of Congenital Heart Disease. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54628-3_5
Download citation
DOI: https://doi.org/10.1007/978-4-431-54628-3_5
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54627-6
Online ISBN: 978-4-431-54628-3
eBook Packages: MedicineMedicine (R0)