Abstract
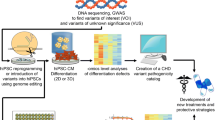
Congenital heart disease (CHD) is the most common cause of neonatal mortality related to birth defects. Etiology is multifactorial including genetic and/or environmental causes. The genetic etiology is known in less than 20 % cases. Animal studies have identified genes involved in cardiac development. However, generating cardiac phenotypes usually requires complete gene knockdown in animal models which does not reflect the haplo-insufficient model commonly seen in human CHD. Human pluripotent stem cells which include human embryonic stem cells (hESC) and human-induced pluripotent stem cells (hiPSC) provide a unique in vitro platform to study human “disease in a dish” by providing a renewable resource of cells that can be differentiated into virtually any somatic cell type in the body. This chapter will discuss the use of human pluripotent stem cells to model human CHD.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
- Human embryonic stem cells
- Induced pluripotent stem cells
- Williams syndrome
- Hypoplastic left heart syndrome
- Fetal reprogramming
1 Introduction
Human embryoni c stem cells (hESC) can give rise to all three germ layers – ectoderm, endoderm, and mesoderm – and can be used to generate differentiated cells of different lineages [1]. The Nobel prize-winning discovery by Yamanaka of the ability to reprogram somatic cells to induced pluripotent stem cells (iPSC) using specific reprogramming factors [2] uncovered a whole new field of research focused on the use of iPSCs to model human disease, perform drug screens, and explore strategies for autologous cell-based therapies in the future. Reprogramming protocols include ectopic expression of four transcription factors [2, 3] that induce reprogramming of somatic cells into an embryonic state. Viral integration-free protocols are also used albeit are less efficient. These cells can then be expanded and differentiated into several somatic cell types including cardiac lineages such as cardiomyocytes, vascular smooth muscles cells (SMCs), and endothelial cells. The process of cardiac differentiation of hESCs and hiPSCs recapitulates cardiac embryogenesis thereby providing a unique opportunity to explore the impact of gene or environmental defects on early cardiac development and gain novel insights into disease mechanisms [4]. Strategies for modeling cardiac malformations are discussed.
2 Modeling Fetal Cardiac Reprogramming in Hypoplastic Left Heart Syndrome (HLHS)
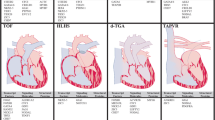
HLHS is one of the m ost severe cardiac malformations characterized by poor growth of left-sided cardiac structures. This is commonly associated with endocardial fibroelastosis (EFE). The mechanism of LV growth failure and fibrosis is poorly understood. We studied 29 normal and 30 HLHS fetal hearts during second trimester [5]. We found increased nuclear expression of hypoxia-inducible factor 1α (HIF1α) in fetal HLHS compared to normal LVs, a central hypoxia-responsive gene that promotes activation of angiogenic, metabolic, and other genes to facilitate cardiac adaptation to hypoxia. However, expression of vascular endothelial growth factor (VEGF) was downregulated. The failure of hypoxia-induced angiogenesis was likely related to cell senescence as shown by DNA damage (nuclear γH2AX activation and p53 upregulation) and of cell senescence (β-galactosidase upregulation). Senescent cells, although functional, do not produce growth factors essential for the survival and proliferation of stem/progenitor cells thereby compromising tissue renewal capacity. Not surprisingly, HLHS hearts showed fewer cardiac progenitor markers, as well as reduced differentiated cardiomyocyte and endothelial cells. DNA damage was most prominent in endothelial cells followed by myocytes, with SMCs being least susceptible. Additionally, there was increased transforming growth factor (TGFβ1) expression, increased myofibroblast transformation, and increased interstitial and perivascular fibrosis in fetal HLHS compared to controls. Together this suggested that the fetal LV may be susceptible to chronic hypoxia or reduced blood flow (a phenomenon that occurs in HLHS due to reduced antegrade flow through the diminutive ascending aorta) resulting in DNA damage and cell senescenc e and consequent loss of cell replication and growth capacity as well as fibrosis.
To clarify the role of hypoxia in fetal cardiac differentiation, we exposed hESC-derived cardiac lineages to 1 % hypoxia for 72 h. This was associated with recapitulation of the fetal HLHS phenotype including increased HIF1α; reduced VEGF; TGFβ1 upregulation; DNA damage (highest in endothelial cells followed by myocytes followed by SMCs); cell senescence; reduced cell proliferation, resulting in a reduction in myocyte and endothelial lineages but increase in SMC lineages; and reduced contractility (Fig. 45.1). Treatment with TGFβ1 inhibitor reversed this abnormal phenotype. This suggests that immature cardiac lineages may be susceptible to hypoxic injury and that this may be mediated in part by TGFβ1 activation. This may contribute to the phenotype of LV growth failure and fibrosis in cardiac malformations like fetal HLHS.
Effect of hypoxia on DNA damage and oncogene upregulation in hPSC-derived cardiac lineages. (a) DNA damage-related marker γH2AX (green nuclear foci) and senescence marker β-gal (blue) are increased in hypoxic cells. Blue represents nuclear staining with DAPI. (b) qPCR results revealed higher mRNA expression of the tumor suppressor oncogene p53 and the G1 cell cycle inhibitors p16 and p18 in hypoxic (gray bars) compared with control cells (black bars). (c) Double immunostaining revealed co-localization of ph-p53 (green) with cTnT+ myocytes (red), CD31+ endothelial cells (red), and SMA+ SMCs (red), indicating DNA damage in all three lineages. Blue represents nuclear staining with TO-PRO-3. (d) Cellomics quantification confirmed the higher number of phospho-p53+ cardiac lineages in hypoxic cells (gray bars) compared with controls (black bars), with most severe injury in ECs followed by myocytes and then SMCs. ∗P < 0.01 versus controls; †P < 0.05 versus SMA+ cells; ‡P < 0.05 versus cTnT+ cells (n = 3 experiments in each group). Original magnification: ×1,000 (γH2AX); ×600 (β-gal) (a); ×1,000 (c) (Reproduced with permission) [5]. Reprinted from Gaber et al. [5], Copyright (2013), with permission from Elsevier [5])
These findings have several implications. They suggest that antenatal intervention for HLHS may be more effective in promoting LV growth if performed before irreversible tissue injury. However, complementary strategies to provide missing growth factors and/or inhibit TGFβ1 either pre- or postnatally may be needed to promote LV growth and ameliorate progressive fibrosis.
3 hiPSCs to Model Williams-Beuren Syndrome (WBS)
WBS is a genetic disorder caused by deletion of 26–28 genes in the 7q11.23 region. Cardiac manifestations are common and are related primarily to haploinsufficiency of the elastin gene in the deleted region. Elastin insufficiency causes vascular SMC proliferation resulting in either generalized arteriopathy or discrete arterial stenoses including supravalvar aortic stenosis, coronary stenosis, pulmonary stenosis, and renal artery stenosis [6]. Surgical correction is often associated with recurrence of stenosis, and there are no medical therapies to prevent or reduce vascular stenoses. Mouse models require complete elastin gene knockdown to reproduce supravalvar aortic stenosis. We therefore generated iPSCs from a patient with WBS with supravalvar aortic stenosis to provide a more human-relevant model for study. Skin fibroblasts obtained at the time of surgery were reprogrammed using four factor retroviral reprogramming. Four iPSC lines were characterized for pluripotency and subjected to SMC differentiation using a published protocol [7]. SMCs generated from iPSCs from normal BJ fibroblasts showed high elastin expression, with 90 % positive for SM22α (a marker of SMC differentiation). These cells showed a good contractile response (Ca2+ flux) to a vasoactive agonist like endothelin and tube-forming capacity on Matrigel assay. In contrast, WBS iPSC-derived SMCs showed low elastin expression, had fewer SM22α-positive cells, were highly proliferative, showed poor tube-forming capacity on Matrigel, and did not show a contractile response to endothelin (Fig. 45.2) [8]. Treatment with rapamycin, a mTOR inhibitor and antiproliferative agent, showed partial rescue of the abnormal phenotype in WBS-SMCs by enhancing differentiation, reducing proliferation, and improving tube-forming capacity. However, it did not restore contractile response to endothelin. Ge et al. used a similar approach to generate and study SMCs from a patient with supravalvar aortic stenosis with WBS and another with elastin loss-of-function mutation that showed a similar phenotype that was rescued by ERK1/2 inhibition [9]. To identify additional compounds that not only improve SMC differentiation but also promote functional maturation and vasoactive responsiveness, we are developing a high-throughput high-content screening assay to facilitate screening of drug libraries using WBS-SMCs. Compounds that fully rescue the abnormal SMC phenotype in WBS may guide the development of new drugs to relieve vascular stenoses in WBS and, by extension, in other vascular disorders including atherosclerosis, stent restenosis, and transplant graft vasculopathy.
Functional characteriza tion of BJ-smooth muscle cells (SMCs) and WBS-SMCs. (a) BJ and WBS-SMCs (line B shown) were treated with 10 mM carbachol, a muscarinic agonist, and phase-contrast live-cell imaging was done every 30 s. Change in cell surface area (white arrows) was calculated from 0 min (top panel) to 30 min (bottom panel). (b) BJ-SMCs showed a 25 % reduction in cell surface area compared with 14 % reduction in WBS-SMCs (average of all four lines). *, p < 0.05 BJ versus WBS. (c) Calcium flux [Ca2+]i was measured in response to endothelin-1 treatment (arrow) in BJ and WBS-SMCs (five cells each). The fluorescence intensity ratio (F340 nm/F380 nm) showed a transient rise in [Ca2+]i after activation by endothelin-1 in BJ-SMCs but not in WBS-B SMCs. (d) Graph showing the ch anges in [Ca2+]i following endothelin-1 treatment in BJ and all the WBS lines. Changes of [Ca2+]i = peak [Ca2+]i − resting [Ca2+]i. *, p < 0.01 BJ versus WBS. WBS Williams-Beuren syndrome (Reprinted from Kinnear et al. [8], Copyright (2013), with permission from Alpha Med Press [8] (pending))
4 Future Directions and Clinical Applications
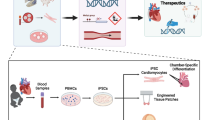
These studies provide proof of principle that hESCs and iPSCs can generate in vitro models to study CHD. However, the cardiac lineages generated using this approach are relatively immature, i.e., fetal stage. While fetal stage cells may be well suited to study developmental cardiac disorders [10], maturation protocols that generate more functionally mature lineages may be more useful to study late-onset disease phenotypes and accurately evaluate drug responses [11–14]. Our study further suggests that the technology can be expanded to study not just genetic influences, particularly in the rapidly emerging era of genome editing [15], but also environ mental teratogens (toxins, chemicals, drugs, infections) to define the mechanisms by which they impact fetal cardiac development or differentiation. This may facilitate delineating the combined role of genetic and environmental factors in CHD causation in the near future [16]. The ability to differentiate pluripotent stem cells into many different organ or cell types may allow the study not only of cardiac but also of extracardiac phenotypes particularly in syndromic disorders as recently shown in a patient with Timothy syndrome [17, 18].
In summary, pluripotent stem cell-derived models are revolutionizing our understanding of disease pathogenesis and are positioned to expedite drug screening and discovery particularly for rare cardiac disorders with a genetic basis for which no therapies are available and where clinical studies are challenging. The technology provides a renewable source of functional cardiomyocytes and other cardiac lineages with genetic and epigenetic variation that are likely to be more human relevant. While the use of these cells for in vivo therapies is several years away, this platform is well positioned to study the molecular underpinnings of genetic cardiac disorders and help identify new therapies for personalized care of the affected child.
References
Thomson JA, Itskovitz-Eldor J, Shapiro SS, et al. Embryonic Stem Cell Lines Derived from Human Blastocysts. Science. 1998;282:1145–1147.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–676.
Yu J. Induced pluripotent stem cell lines derived from human somatic cells. Science. 2007;318:1917–1920.
Yang L, Soonpaa MH, Adler ED, et al. Human cardiovascular progenitor cells develop from a KDR+ embryonic-stem-cell-derived population. Nature. 2008;453:524.
Gaber N, Gagliardi M, Patel P, Kinnear C, Zhang C, Chitayat D, Shannon P, Jaeggi E, Tabori U, Keller G, Mital S. Fetal reprogramming and senescence in hypoplastic left heart syndrome and in human pluripotent stem cells during cardiac differentiation. Am J Pathol. 2013;183(3):720–34.
Pober BR. Williams–Beuren Syndrome. N Engl J Med. 2010;362:239–252.
Xie C-Q, Zhang J, Villacorta L, Cui T, Huang H, Chen YE. Arterioscler Thromb Vasc Biol. 2007;27:e311–e312.
Kinnear C, Chang WY, Khattak S, Hinek A, Thompson T, de Carvalho Rodrigues D, Kennedy K, Mahmut N, Pasceri P, Stanford WL, Ellis J, Mital S. Modeling and rescue of the vascular phenotype of Williams-Beuren syndrome in patient induced pluripotent stem cells. Stem Cells Transl Med. 2013;2(1):2–15.
Ge X, Ren Y, Bartulos O, et al. Modeling Supravalvular Aortic Stenosis Syndrome With Human Induced Pluripotent Stem Cells. Circulation. 2012;126:1695–1704.
Zhang J, Wilson GF, Soerens AG, et al. Functional Cardiomyocytes Derived From Human Induced Pluripotent Stem Cells. Circ Res. 2009;104:e30–e41.
Ng K-M, Lee Y-K, Lai W-H, et al. Exogenous Expression of Human apoA-I Enhances Cardiac Differentiation of Pluripotent Stem Cells. PLoS ONE. 2011;6:e19787.
Paige SL, Osugi T, Afanasiev OK, Pabon L, Reinecke H, Murry CE. Endogenous Wnt/Î2-Catenin Signaling Is Required for Cardiac Differentiation in Human Embryonic Stem Cells. PLoS ONE. 2010;5:e11134.
Otsuji TG, Minami I, Kurose Y, Yamauchi K, Tada M, Nakatsuji N. Progressive maturation in contracting cardiomyocytes derived from human embryonic stem cells: Qualitative effects on electrophysiological responses to drugs. Stem Cell Res. 2010;4:201–213.
Gerecht S, Burdick JA, Ferreira LS, Townsend SA, Langer R, Vunjak-Novakovic G. Hyaluronic acid hydrogel for controlled self-renewal and differentiation of human embryonic stem cells. Proc Natl Acad Sci. 2007;104:11298–11303.
Musunuru K. Genome editing of human pluripotent stem cells to generate human cellular disease models. Dis Model Mech. 2013;6:896–904.
Patel P, Mital S. Stem cells in pediatric cardiology. Eur J Pediatr. 2013;172:1287–1292.
Yazawa M, Hsueh B, Jia X, et al. Using induced pluripotent stem cells to investigate cardiac phenotypes in Timothy syndrome. Nature. 2011;471:230–234.
Pasca SP, Portmann T, Voineagu I, et al. Using iPSC-derived neurons to uncover cellular phenotypes associated with Timothy syndrome. Nat Med. 2011;17:1657–1662.
Acknowledgments
The work was funded by the Canadian Institute of Health Research (MOP 126146), SickKids Labatt Family Heart Centre Innovations fund, and the Ontario Ministry of Research and Innovation GL2 award.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited. The images or other third party material in this chapter are included in the work's Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work's Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Mital, S. (2016). Human Pluripotent Stem Cells to Model Congenital Heart Disease. In: Nakanishi, T., Markwald, R., Baldwin, H., Keller, B., Srivastava, D., Yamagishi, H. (eds) Etiology and Morphogenesis of Congenital Heart Disease. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54628-3_45
Download citation
DOI: https://doi.org/10.1007/978-4-431-54628-3_45
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54627-6
Online ISBN: 978-4-431-54628-3
eBook Packages: MedicineMedicine (R0)