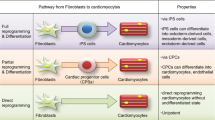
Abstract
Heart disease is a leading cause of death in adults and children. We, and others, have described complex signaling, transcriptional, and translational networks that guide early differentiation of cardiac progenitors and later morphogenetic events during cardiogenesis. We found that networks of transcription factors and miRNAs function through intersecting positive and negative feedback loops to reinforce differentiation and proliferation decisions. We have utilized a combination of major cardiac regulatory factors to induce direct reprogramming of cardiac fibroblasts into cardiomyocyte-like cells with global gene expression and electrical activity similar to cardiomyocytes. The in vivo efficiency of reprogramming into cells that are more fully reprogrammed was greater than in vitro and resulted in improved cardiac function after injury. We have also identified a unique cocktail of transcription factors and small molecules that reprogram human fibroblasts into cardiomyocyte-like cells and are testing these in large animals. Knowledge regarding the early steps of cardiac differentiation in vivo has led to effective strategies to generate necessary cardiac cell types for regenerative approaches and may lead to new strategies for human heart disease.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
1 Introduction
Heart disease remains the leading cause of death worldwide, despite improved treatments that have decreased death rates from cardiovascular diseases. Congenital heart malformations, the most common of all human birth defects, occur in nearly 1 % of the population worldwide, regardless of race. With more than one million survivors of congenital heart disease (CHD) in the United States, it is becoming apparent that genetic disruptions that predispose to developmental defects can have ongoing consequences in maintenance of specific cell types and cellular processes over decades. A more precise understanding of the causes of CHD is imperative for the recognition and potential intervention of progressive degenerative conditions among survivors of CHD.
Because cardiomyocytes (CMs) rarely regenerate in postnatal hearts, survivors of congenital and acquired heart disease often develop chronic heart failure. Unfortunately, end-stage heart failure can only be addressed by heart transplantation , which is limited by the number of donor organs available, particularly in children. Alternative solutions include cellular therapy that replaces lost CMs either by transplanting CMs or inducing new CMs in situ at areas affected by the infarction.
In recent years it has become apparent that adult somatic cells can be converted into other types of cells through epigenetic reprogramming. With this technology, for example, somatic cells, such as fibroblasts, can be dedifferentiated into pluripotent stem cells by nuclear transfer [1] or with defined transcription factors [2]. Direct reprogramming of fibroblasts into the chief functional cells of different organs, including CMs, neurons, hepatocytes, hematopoietic cells, and endothelial cells , has been accomplished and holds great promise for regenerative medicine [3]. In particular, cardiac fibroblasts represent a large pool of cells in the adult heart [4] and thus may provide a reservoir of cells from which to generate new CMs through epigenetic reprogramming.
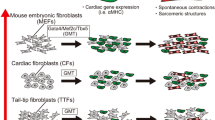
In 2010, we reported that mouse cardiac and dermal fibroblasts could be directly reprogrammed into induced CM-like cells (iCMs) in vitro by a combination of three developmental cardiac transcription factors, Gata4, Mef2c, and Tbx5 (GMT) [5]. Since then, other labs around the world have reported success in reprogramming mouse fibroblasts into iCMs with similar cocktails of reprogramming factors [6–10]. We and others have also directly converted human cardiac and dermal fibroblasts into cardiac cells [11–13]. Several review papers published from different labs illustrate the potential and challenges of this new avenue for cardiac regenerative medicine [14–19].
Here, we discuss the cardiac developmental biology discoveries that underpin direct reprogramming approaches and consider the opportunities and challenges of this technology for addressing cardiac regeneration .
2 Molecular Networks Regulate Cardiac Cell Fate
In vertebrate embryos, the heart is the first functional organ to form. The morphological development of the heart has been summarized in several reviews. During development, numerous signaling and transcriptional cascades regulate cell fate decisions in distinct heart fields . In addition, the Gata, Mef2, Hand, Nkx, and T-box family of transcription factors control expression of cardiac genes and direct the specification and differentiation of cardiac myocytes. These transcription factors have been frequently included in studies that directly convert fibroblasts into cardiac cells.
MicroRNAs (miRs) —small single-stranded noncoding RNAs that negatively regulate the stability of gene transcripts—also regulate cardiac gene expression [20, 21]. Transcription factors regulate miR expression, and in turn, miRs can modulate the activities of transcription factors through positive and negative feedback loops. One of the major regulators of cardiac lineage determination during heart development is miR-1. Expression of miR-1 in either mouse or human embryonic stem cells (ESCs) caused them to favor the muscle cell fate. In contrast, miR-133 promoted muscle progenitor expansion and prevented terminal differentiation, while another miRNA, miR-499, promoted the ventricular cell fate in human ESC-differentiated CMs and caused cardiac hypertrophy and enlarged hearts in miR-499 transgenic mice. These studies demonstrate that miRs cooperate with transcription factors to form an intertwined network that reinforces specific cell fate decisions and differentiation during cardiac development .
3 Cardiac Fibroblasts in the Normal and Remodeling Heart
Fibroblasts are mesenchymal cells that produce many extracellular matrix components in organs. Fibroblasts show heterogeneity based on morphology, glycogen pools, collagen production, cell surface markers, and global gene-expression profiles. Although the percentage of fibroblasts among the total cells in the heart varies between species, the large population of fibroblasts is quiescent and abundantly distributed in the interstitial and perivascular matrix in the normal heart.
Cardiac fibroblasts synthesize extracellular matrix to provide a 3D network for myocytes and other cells of the heart; they also regulate the biological and electrophysiological response of CMs during physiological and pathological development. In embryonic mouse hearts, cardiac fibroblasts induced proliferation of CMs via paracrine signals of fibronectin, collagen, and heparin-binding EGF-like growth factor ; however, in adult mouse hearts, cardiac fibroblasts promoted hypertrophic maturation of CMs via beta1-integrin signaling [22]. Cardiac fibroblasts were also found to form intracellular electrical coupling and communicate with myocytes through gap junctions , suggesting that cardiac fibroblasts could conduct electric signaling between different regions of myocytes that are electrically isolated by connective tissue in the normal heart.
4 Direct Cardiac Reprogramming In Vitro
To identify the combination of reprogramming factors that could convert a cardiac fibroblast toward a cardiomyocyte-like state, we generated a transgenic mouse in which enhanced green fluorescent protein (EGFP) was driven by the alpha myosin heavy chain (αMHC) promoter [5, 22], providing a tool to screen through many potential regulators. We ultimately found that a combination of three transcription factors—Gata4, Mef2c, and Tbx5 (GMT)—could convert ~15 % of cardiac fibroblasts into αMHC-EGFP-positive cells, which we called induced CM-like cells (iCMs) [5]. iCMs formed sarcomere structures and displayed whole-transcriptome expression profiles that were shifted significantly toward the profile of CMs, even at the single-cell level. Although most in vitro iCMs were only partially reprogrammed, many of them could generate Ca2+ transients, and some started beating spontaneously 4–6 weeks after reprogramming. Using a lineage-tracing strategy in mice (e.g., Isl1-Cre-YFP and Mesp1-Cre-YFP), we did not observe activation of cardiac progenitor markers during GMT cardiac reprogramming [5], suggesting that GMT directly converted fibroblasts toward the cardiac cell fate without dedifferentiation back into a progenitor status. Song et al. [8] found that a basic helix-loop-helix transcription factor, Hand2, could help GMT to convert TTFs into functional beating iCMs.
5 Direct Cardiac Reprogramming In Vivo
The ultimate goal in generating new iCMs is to improve systolic function of damaged hearts and restore its normal structure and function. Therefore, we tested direct cardiac reprogramming in vivo with the hypothesis that the heart’s native microenvironment would promote direct reprogramming of fibroblasts to CMs. We expected that direct cardiac reprogramming would be enhanced in the native heart and could improve the function of the damaged heart.
We delivered GMT into mouse hearts after acute MI via retroviruses. These viruses can only infect actively dividing cells and, thus, could deliver the reprogramming factors into non-myocytes (mostly fibroblasts ), but not into CMs that exit the cell cycle after differentiation. Four weeks after introducing GMT, we found numerous genetically labeled reprogrammed cells within the scar area of mouse hearts, indicating newly born iCMs reprogrammed from cardiac fibroblasts (Fig. 1.1). We found that more than 50 % of in vivo-derived iCMs closely resembled endogenous ventricular CMs, had a rod shape, were binucleate, assembled sarcomeres, generated Ca2+ transients, and elicited ventricular-like action potentials and beating activity [23]. By microarray analysis, in vivo mouse GMT-iCMs showed similar global gene-expression profiles with mouse adult CMs, such that they were clustered as one type of cells [11]. The iCMs reprogrammed in vivo were electrically coupled with endogenous CMs, and no arrhythmias were observed in mice that received GMT reprogramming factors. Most importantly, introducing GMT in vivo reduced scar size and cardiac dysfunction up to 12 weeks after coronary ligation [23]. Similarly, Inagawa et al. [24] successfully reprogrammed cardiac fibroblasts into iCMs in vivo by introducing GMT into the heart of immunosuppressed mice with single-polycistronic retrovirus, which contains GMT with self-cleaving 2A peptides. Furthermore, Song et al. [8] found that introducing GMT and Hand2 in vivo could directly convert cardiac fibroblasts into iCMs and also improved function and decreased scar.
6 Direct Cardiac Reprogramming in Human Fibroblasts
Establishing the technology of cardiac reprogramming in human cells was a necessary step toward considering clinical application. However, neither GMT nor GHMT, which reprogrammed iCMs from mouse fibroblasts, was able to reprogram human fibroblasts into iCMs in vitro [11–13]; however, inclusion of additional reprogramming factors resulted in successful reprogramming. Nam et al. [12] found that GHT, without MEF2C, but with another transcription factor, myocardin, and two muscle-specific miRNAs, miR-1 and miR-133, could reprogram human fibroblasts into iCMs. These reprogrammed iCMs expressed multiple cardiac genes, developed sarcomere-like structures, and generated Ca2+ transients with a small subset of the cells exhibiting spontaneous contractility after 11 weeks in culture. Wada R. et al. [13] reported that GMT with MESP1 and myocardin could activate cardiac gene expression in human neonatal and adult cardiac fibroblasts. In our study, pairing GMT with ESRRG and MESP1 induced global expression of cardiac genes and shifted the phenotype of human fibroblasts toward the CM-like state. Reprogrammed human iCMs were epigenetically stable and formed sarcomere structures, and some could generate Ca2+ transients and action potentials [11]. By comparing whole-transcriptome expression of 4-week and 12-week iCMs, we found that reprogramming human cells takes longer than mouse cells because of their progressive repression of fibroblast genes. Nonetheless, our analysis of orthologous gene expression indicated that at the global gene-expression level, human iCMs were reprogrammed at a level similar to mouse iCMs reprogrammed by GMT in vitro [11].
7 Challenges and Future Directions
The studies summarized here demonstrate that forced expression of three or four developmentally critical transcription factors can directly convert mouse cardiac fibroblasts into CM-like cells in vitro, although most of these iCMs were partially reprogrammed. These same factors generated more mature CM-like cells in the native heart, improved heart function, and reduced scar size in the mouse heart post-MI. Similarly, expression of five to seven reprogramming factors converted human fibroblasts into non-perfect iCMs in vitro. By comparing in vitro versus in vivo iCMs in mice [8, 23] and considering the similarities between human and mouse iCMs in vitro [11], we speculate that the cocktails recently identified in human fibroblasts may be sufficient to reprogram adult CM-like cells that are fully functional in the in vivo environment, such as in the pig or nonhuman primate heart, as is the case in mice. These studies have been driven by the deep knowledge of cardiac development gained over the last two decades and represent a valuable application of this knowledge for potential clinical development postnatally.
Successfully achieving cardiac reprogramming requires high expression and proper stoichiometry of reprogramming factors, healthy and non-senescent fibroblasts , and optimal conditions for cell culture. In addition, culture conditions, such as electric stimulation, might help facilitate and maintain the functional maturation of iCMs at late stages (i.e., after cardiac cell fate conversion), and the process may require some small-molecule compounds and growth factors to overcome epigenetic barriers at the early stages of in vitro reprogramming. The epigenetic barriers that prevent cardiac reprogramming in vitro remain unknown; however, the in vivo environment of the heart appears to overcome these epigenetic blocks. We speculate that secreted factors and direct cell-cell interactions, including mechanical and electrical, from myocytes and non-myocytes may work together to improve direct cardiac reprogramming.
Another elusive concept is the molecular mechanism that underlies direct cardiac reprogramming. What are the DNA targets of those reprogramming factors? How are those transcriptional changes epigenetically stabilized during reprogramming? By combining mechanism assays at whole-population and single-cell levels, we can gain a more integral and comprehensive understanding of how core transcription factors establish a self-reinforcing molecular network that controls cardiac cell fate.
While many challenges and hurdles remain in this blossoming research field, the high demand for regenerative medicine strategies for the heart emphasizes the significance of these efforts in discovering new therapeutic strategies. Observing the functional benefits of in vivo reprogramming in mouse heart and the promising and similar degree of reprogramming in mouse and human in vitro iCMs, we are endeavoring to translate direct cardiac reprogramming for future clinical applications.
References
Gurdon JB. The developmental capacity of nuclei taken from intestinal epithelium cells of feeding tadpoles. J Embryol Exp Morphol. 1962;10:622–40.
Takahashi K, Yamanaka S. Induction of pluripotent stem cells from mouse embryonic and adult fibroblast cultures by defined factors. Cell. 2006;126:663–76.
Sancho-Martinez I, Baek SH, Izpisua Belmonte JC. Lineage conversion methodologies meet the reprogramming toolbox. Nat Cell Biol. 2012;14:892–9.
Snider P, Standley KN, Wang J, Azhar M, Doetschman T, Conway SJ. Origin of cardiac fibroblasts and the role of periostin. Circ Res. 2009;105:934–47.
Ieda M, Fu JD, Delgado-Olguin P, Vedantham V, Hayashi Y, Bruneau BG, et al. Direct reprogramming of fibroblasts into functional cardiomyocytes by defined factors. Cell. 2010;142:375–86.
Jayawardena TM, Egemnazarov B, Finch EA, Zhang L, Payne JA, Pandya K, et al. MicroRNA-mediated in vitro and in vivo direct reprogramming of cardiac fibroblasts to cardiomyocytes. Circ Res. 2012;110:1465–73.
Protze S, Khattak S, Poulet C, Lindemann D, Tanaka EM, Ravens U. A new approach to transcription factor screening for reprogramming of fibroblasts to cardiomyocyte-like cells. J Mol Cell Cardiol. 2012;53:323–32.
Song K, Nam YJ, Luo X, Qi X, Tan W, Huang GN, et al. Heart repair by reprogramming non-myocytes with cardiac transcription factors. Nature. 2012;485:599–604.
Addis RC, Ifkovits JL, Pinto F, Kellam LD, Esteso P, Rentschler S, et al. Optimization of direct fibroblast reprogramming to cardiomyocytes using calcium activity as a functional measure of success. J Mol Cell Cardiol. 2013;60:97–106.
Christoforou N, Chellappan M, Adler AF, Kirkton RD, Wu T, Addis RC, et al. Transcription factors MYOCD, SRF, Mesp1 and SMARCD3 enhance the cardio-inducing effect of GATA4, TBX5, and MEF2C during direct cellular reprogramming. PLoS One. 2013;8:e63577.
Fu JD, Stone NR, Liu L, Spencer CI, Qian L, Hayashi Y, et al. Direct reprogramming of human fibroblasts toward a cardiomyocyte-like state. Stem Cell Rep. 2013;1:235–47.
Nam YJ, Song K, Luo X, Daniel E, Lambeth K, West K, et al. Reprogramming of human fibroblasts toward a cardiac fate. Proc Natl Acad Sci U S A. 2013;110:5588–93.
Wada R, Muraoka N, Inagawa K, Yamakawa H, Miyamoto K, Sadahiro T, et al. Induction of human cardiomyocyte-like cells from fibroblasts by defined factors. Proc Natl Acad Sci U S A. 2013;110:12667–72.
Yi BA, Mummery CL, Chien KR. Direct cardiomyocyte reprogramming: a new direction for cardiovascular regenerative medicine. Cold Spring Harb Perspect Med. 2013;3:a014050.
Srivastava D, Berry EC. Cardiac reprogramming: from mouse toward man. Curr Opin Genet Dev. 2013;23:574–8.
Qian L, Srivastava D. Direct cardiac reprogramming: from developmental biology to cardiac regeneration. Circ Res. 2013;113:915–21.
Nam YJ, Song K, Olson EN. Heart repair by cardiac reprogramming. Nat Med. 2013;19:413–5.
Addis RC, Epstein JA. Induced regeneration – the progress and promise of direct reprogramming for heart repair. Nat Med. 2013;19:829–36.
Muraoka N, Ieda M. Direct reprogramming of fibroblasts into myocytes to reverse fibrosis. Annu Rev Physiol. 2014;76:21–37.
Ivey KN, Srivastava D. MicroRNAs as regulators of differentiation and cell fate decisions. Cell Stem Cell. 2010;7:36–41.
Cordes KR, Srivastava D. MicroRNA regulation of cardiovascular development. Circ Res. 2009;104:724–32.
Ieda M, Tsuchihashi T, Ivey KN, Ross RS, Hong TT, Shaw RM, et al. Cardiac fibroblasts regulate myocardial proliferation through beta1 integrin signaling. Dev Cell. 2009;16:233–44.
Qian L, Huang Y, Spencer CI, Foley A, Vedantham V, Liu L, et al. In vivo reprogramming of murine cardiac fibroblasts into induced cardiomyocytes. Nature. 2012;485:593–8.
Inagawa K, Miyamoto K, Yamakawa H, Muraoka N, Sadahiro T, Umei T, et al. Induction of cardiomyocyte-like cells in infarct hearts by gene transfer of gata4, mef2c, and tbx5. Circ Res. 2012;111:1147–56.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is distributed under the terms of the Creative Commons Attribution-Noncommercial 2.5 License (http://creativecommons.org/licenses/by-nc/2.5/), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited. The images or other third party material in this chapter are included in the work's Creative Commons license, unless indicated otherwise in the credit line; if such material is not included in the work's Creative Commons license and the respective action is not permitted by statutory regulation, users will need to obtain permission from the license holder to duplicate, adapt or reproduce the material.
Copyright information
© 2016 The Author(s)
About this chapter
Cite this chapter
Srivastava, D. (2016). Reprogramming Approaches to Cardiovascular Disease: From Developmental Biology to Regenerative Medicine. In: Nakanishi, T., Markwald, R., Baldwin, H., Keller, B., Srivastava, D., Yamagishi, H. (eds) Etiology and Morphogenesis of Congenital Heart Disease. Springer, Tokyo. https://doi.org/10.1007/978-4-431-54628-3_1
Download citation
DOI: https://doi.org/10.1007/978-4-431-54628-3_1
Published:
Publisher Name: Springer, Tokyo
Print ISBN: 978-4-431-54627-6
Online ISBN: 978-4-431-54628-3
eBook Packages: MedicineMedicine (R0)