Abstract
Huntington’s disease (HD) is a fatal, dominantly inherited neurodegenerative disorder caused by a CAG trinucleotide expansion in the Huntingtin (HTT) gene, leading to an expanded polyglutamine (polyQ) region in the encoded protein HTT. We have used homologous recombination (HR) to genetically correct HD patient-derived induced pluripotent stem cells (iPSCs) and found that this reversed HD disease phenotypes. We have utilized exploited genome editing tools including TALENs (Transcription like activator effectors) and CRISPR (Clustered Regulatory Interspaced Short Palindromic Repeats)/Cas9 technology to carry out genetic correction or expansion, and we were able to detect HR without selection in human cells. The overall goal is to use this technology to model HD-relevant cell types and better understand disease progression by leveraging system biology approaches. To understand the disease progression, isogenic iPSC lines were created. We found that the disease phenotypes only manifested in the differentiated neural stem cell (NSC) stage, not in iPSCs. Transcriptomic analysis of HD iPSCs and HD NSCs compared to isogenic controls was utilized to understand the molecular basis for the CAG repeat expansion-dependent disease phenotypes in NSCs. Differential gene expression and pathway analysis identified transforming growth factor β (TGF-β) signaling, netrin-1 signaling and medium spiny neuron (MSNs) maturation and maintenance as the top dysregulated pathways in HD NSCs. The ability to create additional isogenic cell lines through CRISPR-mediated HR will further enhance our understanding of HD progression. These lines can be manipulated with CRISPR to understand the effects of common SNPs (single nucleotide polymorphism) that modulate disease onset in HD, allowing the identification of new pathways and helping to elucidate potential therapeutic targets for HD. Beyond drug discovery, the CRISPR system could eventually be optimized to use in vivo, correcting a patient’s disease-causing mutation, in the asymptomatic stages of HD.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Huntington’ Disease
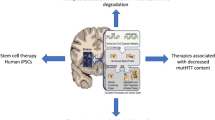
Huntington’s disease (HD) is a devastating, dominantly inherited movement and psychiatric disorder that is caused by expansion of a CAG trinucleotide repeat in the first exon of the Huntingtin gene (HTT), resulting in translation of an expanded polyQ repeat in the HTT protein. The production of the abnormal expanded polyQ-containing HTT protein leads to a dramatic loss of striatal and cortical neurons and pro-survival growth factors such as BDNF (brain derived neurotrophic factor) in HD patients. The polyQ expansion in the HTT protein leads to disrupted cellular homeostasis and activation of cellular death pathways (Fig. 1). Since the disease is inherited in an autosomal dominant fashion, each child of an affected parent has a 50% chance of being affected. HD generally manifests in mid-life, with a mean age of onset of 35–45 years of age. The disease begins with cognitive disturbances and progresses to severe and debilitating motor symptoms (chorea) usually accompanied by psychiatric disturbances, with death following in about 15–20 years (Landles and Bates 2004). The current therapeutic approaches in HD focus on normalizing molecular pathways disturbed in HD or on lowering the levels of the mutant HTT protein (Canals et al. 2004; Conforti et al. 2008; Zuccato et al. 2008). To date none of these approaches are approved for use outside of clinical trials and they will not cure the disease.
Illustration on the neuronal changes occurring in the striatum of a Huntington's disease patient. The exon 1 CAG expansion in the HTT allele results in a mutant protein being formed; the mutant protein aggregates and is also cleaved into toxic fragments. The aggregates and the toxic fragments result in a disrupted cellular homeostasis and eventual neuronal cellular death in the striatum
In this chapter, we discuss the use of gene editing tools to model neurological diseases such as HD as well as the potential to use this technology to treat genetic neurological diseases.
Gene Editing Enzymes
Targeted gene editing has evolved dramatically in the last 25 years. While originally a technique that a handful of laboratories had mastered, it is now a common tool used in hundreds of laboratories around the world. One family of gene editing proteins is the customized zinc finger proteins (Segal and Barbas 2000; Wolfe et al. 2000; Pabo et al. 2001; Nagaoka and Sugiura 2000). These proteins were adapted for targeted use in the late 1990s (Liu et al. 1997; Segal et al. 1999; Dreier et al. 2001). Each zinc finger protein could be designed to recognize three different base pairs on DNA through various interactions between the proteins alpha helix amino acids and the DNA base pairs (Segal and Barbas 2001). To recognize a specific sequence of DNA, the zinc fingers could be attached to each other, with six zinc fingers recognizing a unique 18-base pair sequence in an organism’s genome. The zinc finger proteins could have effector or nuclease domains attached, allowing for gene regulation or gene replacement. The effector domains included VP64 for gene activation, KRAB for gene silencing and DNMT1 for methylation (Beerli et al. 1998; Rivenbark et al. 2012). The nuclease domain could cut targeted genomic sites and allow for mutagenesis or homologous recombination at enhanced efficiency. Zinc finger proteins have been successfully used in human cells, animal organs and have reached Phase II human clinical trials (Geurts et al. 2009; Urnov et al. 2005; SangamoBiosciences 2001; Eisenstein 2012). Although promising, zinc fingers presented several challenges for researchers. Their targeting ability was limited, they required specialized design techniques and they exhibited a frequent incidence of off-target events (Cornu and Cathomen 2010; Gupta et al. 2010; Gabriel et al. 2011). Some advances have been made to reduce the off-target potential and increase detection of these events (Zykovich et al. 2009; Cornu et al. 2008). The therapeutic potential of zinc fingers for a variety of diseases, including HD, continues to be explored by the biotechnology company Sangamo (Cornu et al. 2008; Wolffe 2016).
In 2009, a new gene editing protein was described, transcription activator-like effectors (TALEs; Boch et al. 2009; Moscou and Bogdanove 2009). These proteins were originally characterized in Xanthomonas bacteria and represented a major advance for DNA regulating proteins. TALEs, unlike zinc fingers, made contact with individual DNA base pairs, which greatly expanded the sequences that could be targeted in the genome (Moscou and Bogdanove 2009). They were also much easier to design and assemble. Much like zinc fingers, TALEs could have effector or nuclease domains attached to the DNA binding domain, allowing for the DNA to be cut or for genes to be regulated (Christian et al. 2010; Maeder et al. 2013a, b; Cong et al. 2012). Promising experiments in a variety of organisms have validated the efficacy of TALEs, although no human clinical trials have begun. A recent publication has shown the ability of TALEs to specifically silence the mutant HTT allele in cell culture models or to engineer an allelic series into the HTT locus (Fink et al. 2016; Wang et al. 2013). The TALEs still exhibit off-target effects and may have potential immune issues (Guilinger et al. 2014).
Gene editing became a widely accessible technology in 2012 with the characterization of the CRISPR system and its implications for targeted gene editing and regulation. The CRISPR system is composed of a Cas9 nuclease and a gRNA complex. To cut the DNA, Cas9 attaches to the guide RNA (gRNA), which targets a specific site in the organism’s DNA (Jinek et al. 2012; Wiedenheft et al. 2012). This system is found in archea and bacteria and is used as a natural defense mechanism against bacteriophages. The system has been characterized and adapted for mammalian-targeted genome editing. The gRNA has one targeting requirement, a PAM motif (typically a NGG) at the 3′ end of the DNA targeting site; this sequence is common in DNA and thus almost any gene can be targeted with the CRISPR system (Gilbert et al. 2013; Qi et al. 2013). As with previous gene editing proteins, the Cas9 can be modified to either silence or activate gene transcription (Fig. 2; Sander and Joung 2014; Larson et al. 2013). Due to some initial off-target cleavage events, the Cas9 nuclease was modified to become a Cas9 nickase (Cas9n; Ran et al. 2013). This modification drastically increased targeting specificity, as the binding of two Cas9n proteins targeting two different DNA sites was required to make a double strand break in the DNA and encouraged homologous recombination (HR) with a potential donor DNA strand. Overall the off-target effects of Cas9n could be reduced to background levels (O’Geen et al. 2015; Wu et al. 2014). The modified Cas9n was found to have similar cleavage efficiency when two gRNAs were used, one targeted on each strand of the DNA, resulting in a double strand break. The technique has been widely adopted to create disease-modeling cell lines, rodent and non-human primate models and in non-viable human embryos (Liang et al. 2015; Chen et al. 2015). What has made the CRISPR system so accessible is that, unlike the zinc fingers and TALEs, the same core protein, Cas9, is used to target any sequence, whereas the targeting portion of the CRISPR system, the gRNA, is what varies. The gRNA can be designed and synthesized either in a standard lab or by an outside company. This separation of the targeting portion (gRNA) of the CRISPR system from the modifying portion (Cas9 or other effectors) allows for targeting multiple genes in one experiment (Wang et al. 2013). The ability to target multiple genes in a single experiment drastically reduces the time needed to model complex genetic disorders in which more than one gene is involved. All of these unique characteristics have resulted in a rapid popularization of the CRISPR system in research labs, with thousands of papers having been published in the last five years.
Illustrations of different CRISPR/Cas9 uses with variable effector domains. The wild-type Cas9 nuclease can be used to initiate double strand breaks, encouraging homologous recombination. The inactive Cas9 (dCas9) attached to a DNMT3 can be used for site-specific methylation, resulting in semi-permanent gene repression. A dCas9 can have a KRAB domain attached for temporary gene repression or a VP64 domain for activation
Uses for Gene Editing to Understand Human Diseases
Due to their ability to precisely target a gene or regulatory element, genome editing tools have been widely utilized to model human diseases both in cells and in animals. Neurodegenerative diseases such as Parkinson’s disease and HD have been modeled by introducing disease-causing mutations into human induced pluripotent stem cells (iPSCs) facilitated by genome editing tools (O’Brien et al. 2015; Soldner et al. 2011). CRISPR/Cas9 or TALENs can also be injected into zygotes or embryos to get genetically modified animals. Researchers have injected TALEN-expressing mRNAs into zebrafish embryos to target the gene glucocerebrosidase 1, which is mutated in the lysosomal storage disorder Gaucher’s disease. The introduction of these TALENs caused a deletion mutation of the protein Glucocerebrosidase 1, and characteristics of the Gaucher’s disease were present in this zebrafish model (Keatinge et al. 2015). Duchenne muscular dystrophy (DMD) is a neuromuscular disorder caused by a loss-of-function mutation of the gene dmd. A DMD rat model was generated by delivering CRISPR system into rat zygotes to target the dmd gene (Nakamura et al. 2014). These disease models are valuable tools for the exploration of disease mechanisms and for the pursuit of therapeutics.
When combined with human pluripotent stem cells, genome editing tools can provide some unique advantages in disease modeling and mechanism study. Human pluripotent stem cells, including iPSCs and embryonic stem cells, can be directed to any cell types of the human body with the correct differentiation conditions. Thus relevant cell types for the disease and changes in this development can be studied in these models. When genome editing tools are used to add or remove a mutation at the pluripotent stem cell stage, isogenic cell lines with an almost identical genetic background are obtained. As cells are differentiated into more restricted stem cells and terminally differentiated cells, the isogenic background will persist. Phenotypic changes of these cells are most likely a result of the mutation, as they have an identical genetic background. However, one may still have to consider epigenetic changes and mitochondrial mutations that may remain harbored in the patient’s iPSCs’ background (Chinnery et al. 2012; Calvanese et al. 2009). These isogenic cell lines can be subjected to systematic approaches including DNA microarray, RNA-seq and mass spectrometry for transcriptomic and proteomic information. Bioinformatic analysis can identify interesting gene/protein targets or signaling pathways that have distinct disease-associated patterns. The cleaner background of isogenic cell models should result in more relevant and reliable hits. After proper validation, these potentially important disease targets may lead to discovery of new mechanisms or drugs.
Recent advances in stem cell research suggest that iPSCs may provide novel models of disease and new treatments for diseases. An isogenic iPSC line was established in the Ellerby lab through traditional means of HR on a human HD patient iPSC line. This isogenic line introduced a corrected donor strand for the CAG expansion and corrected the disease allele to a wild type allele (An et al. 2012). The isogenic corrected line had the exact same genetic background as the patient, reducing the genetic variables that are present when one compares disease phenotypes across multiple different patients to matched wild type individuals. One of the first questions we addressed was whether we could take HD patient-derived iPSCs and, through genetic correction of the disease allele, reverse disease phenotypes. Interestingly, we did not detect phenotypes in the undifferentiated HD iPSCs but only observed disease phenotypes in the differentiated neural stem cell (NSC) state, and these phenotypes were reversible upon genetic correction of the patient mutation.
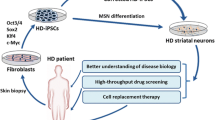
To understand the molecular basis for the CAG repeat expansion-dependent disease phenotypes in iPSCs and NSCs, RNA-Seq was performed comparing the isogenic corrected lines to HD iPSCs and HD NSCs. We observed that there were few phenotypic differences between HD and wild type iPSCs, but there were substantial differences—over 2000 dysregulated genes—in the NSCs. Some of the key pathways that were dysregulated included TGF-β, netrin-1 signaling and development of the striatum (Fig. 3; Ring et al. 2015). Particularly important, our isogenic HD-iPSCs with corrected alleles identified the maturation or maintenance of medium spiny neurons (MSNs) as being dysregulated (Ring et al. 2015). We showed that the pathways or factors that were involved in this process were therapeutic targets for HD (Ring et al. 2015). A subsequent publication from another group emphasized the de-differentiation of MSNs or loss of MSN identity in HD is a major source of dysfunction (Langfelder et al. 2016). These pathways offer new options for therapeutic treatments and drug targets. Using genetic engineering, we generated an isogenic allelic HD iPSC series for HD modeling (CAG repeat of 21, 45, 72, 100). By creating additional isogenic lines, the contribution of the CAG expansion to the disease phenotypes can be elucidated from background variation; this information can help guide researchers towards additional treatment targets (O’Brien et al. 2015).
A flow chart comparing the donor Huntington disease (HD) and genetically corrected isogenic iPSCs and NSCs. Transcriptomic analysis was performed on the cell lines in which significant differences were found in multiple signaling pathways. These newly identified pathways could result in additional drug targets
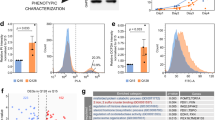
Besides directly modifying the disease gene, genome editing tools can also be used to engineer cells to facilitate disease research by making reporter cell lines. In an effort to investigate the roles of a gene encoding a sodium channel subunit in epilepsy, a tdTomato fluorescence protein gene cassette was inserted into iPSCs under a GABAergic neuron-specific promoter with CRISPR/Cas9. When differentiated into GABAergic neurons, these cells were red fluorescently labeled and could be readily followed for electrophysiological studies (Liu et al. 2016). Another example is in the peripheral neuropathy Charcot–Marie–Tooth disease, type 1A. With TALENs a bioluminescent reporter was integrated under the regulation of the disease causing gene pmp22, which allowed high throughput screening for reagents that can decrease expression of this gene (Inglese et al. 2014). In an effort to better track the recombination repair efficiency in HD cells, the Ellerby lab has designed a myc-tagged donor strand that, when incorporated into the cell, is detectable by both Western blot and PCR amplification; these methods are so sensitive that recombination efficiencies can be detected at levels as low as 5% (Fig. 4). For polyglutamine disease, it is also possible to detect the prevalence of the polyglutamine expansion through the use of specific antibodies, which detect the expanded polyglutamine region (Fig. 4; An et al. 2014). The ability to qualitatively assess how many cells have been corrected will increase the field’s understanding of what may be a therapeutic level of correction for the disease. Having specific tags to monitor genetic correction rates and resulting phenotypic improvements will advance the field’s understanding toward designing genetic correction and optimize treatment conditions.
(a) Use of myc tag in corrected donor plasmid allows for both insertion screening at the DNA level by PCR (left) and at the protein level by Western blot (right); red triangles indicate expected band size. (b) Use of 1C2 antibody screening with an expanded donor plasmid, a rapid method to optimize different gRNA combinations for homologous recombination efficiency
Gene Editing In Vivo to Treat Genetic Diseases
With its extreme ease of use and targeting, the CRISPR system is being studied extensively with a goal of in vivo correction of genetic mutations. Recent advances have shown that it takes about 15 h for Cas9-mediated double strand breaks to be repaired; this is potentially due to Cas9 remaining bound to the DNA for an extended period of time and because it asymmetrically releases the target strand (Richardson et al. 2016). This asymmetric release of the strand has given researchers the ability to rationally design the donor strands in an effort to increase gene correction percentages; it also provides additional insight as to how to target and design the donor strands. The guide RNAs have also continued to evolve since the first characterization of the CRISPR system. Initially there were two components to the guide RNA, a crRNA and a gRNA, and these were able to be fused creating a simpler method in which the gRNA could be delivered already assembled. Multiple assembled gRNAs could be placed on the same plasmid, allowing for multiple gene targeting with minimal plasmids (Wang et al. 2013; Hsu et al. 2013). A couple of new CRISPR variants have been characterized that offer even lower off-target binding levels and are smaller (Ran et al. 2015). Both of these new characteristics may be useful in eventual patient treatment, as a smaller CRISPR protein could be more easily packaged for delivery and lower off-target binding increases the specificity of the CRISPR protein, restricting the effects to the target site.
The most exciting application of genome editing tools in human genetic diseases is genetic correction and normalization of those disease mutations. These have been done in cells. For example, in Myotonic dystrophy type 1, a genetic modification has been introduced by TALEN in a NSC model and this modification has shown some restoration of disease phenotypes (Xia et al. 2015). More encouragingly, genetic correction has been achieved in adult animals. Recently several groups published genetic correction in a mouse DMD model. Adeno-associated virus-delivered CRISPR/Cas9 was used to remove a mutation from the gene dmd. Partial phenotypic recovery has been observed in these studies (Xu et al. 2016; Nelson et al. 2016; Tabebordbar et al. 2016). The use of CRISPR in vivo to ablate the rhodopsin gene carrying the dominant S334ter mutation in rats with severe autosomal dominant retinitis pigmentosa also highlights the use of genetic correction in disease (Bakondi et al. 2016). These proof-of-principle experiments may be the first steps towards overcoming many currently incurable genetic diseases. CRISPR technology is already being used in human cells and disease models with the eventual goal of patient treatment. A recent study conducted in China has even used CRISPR technology on non-viable human embryos (Liang et al. 2015). As this technology has advanced so rapidly, the scientific community has held a summit meeting to discuss the potential future of CRISPR technology, much in the same way the Asilomar Conference discussed recombinant DNA over 40 years ago (Baltimore et al. 2015; Berg et al. 1975a, b).
In HD, it is possible that a variety of CRISPR tools could prove beneficial for treatment. Previous studies have shown that a reduction in mutant HTT levels can ameliorate symptoms of the disease (Canals et al. 2004; Conforti et al. 2008; Zuccato et al. 2008). A recent study has shown reduction of mutant Huntingtin in cells by using TALE-ATFs (artificial transcription factors) to specifically target the mutant allele by targeting SNPs common on that allele. The TALE-ATF has a KRAB domain attached that represses transcription of the mutant Huntingtin allele (Fink et al. 2016). This technique has yet to be tried in Huntington model mice; however, previous studies have used ATFs to repress transcription in the brains of mice (Bailus et al. 2016). Another approach using CRISPR would involve increasing transcription of genes that could be neuroprotective in HD; BDNF could be a potential target for this type of therapy (Pollock et al. 2016). As screening studies are further refined using more genetically engineered isogenic cell lines, it will be possible to uncover additional gene regulation targets.
The ideal therapy for HD would involve gene replacement therapy, where the mutant allele would be replaced by a corrected donor allele. Using the CRISPR system, it will eventually be possible to do this correction in vivo. When designing the donor strand, it is possible to detect site-specific insertion by PCR if a small tag is added to the donor strand, allowing for optimization of different CRISPR components (Fig. 4). After design and condition optimization, there are still several issues that need to be addressed to develop CRISPR into an in vivo therapy. One area to examine is the immune response, as Cas9 is not an endogenous protein in mammals, although there are mouse models that constitutively express Cas9 from birth (Platt et al. 2014). Previous studies in humans with zinc finger proteins have shown minimal immune response. Cas9 is not endogenous to animals and may elicit an immune response if given over an extended period of time. A second major concern for gene correction in vivo is the delivery of the CRISPR system to the desired organ or tissue. For certain diseases, it may be possible to directly inject the organ and correct only a subpopulation of the cells; for other diseases, especially those that effect the brain, delivery is more difficult (Liu et al. 2016; Yin et al. 2016). Direct injection into the brain is possible, and packaging the CRISPR system into an appropriately pseudotyped viral vector could allow for additional coverage beyond the injection point. The CRISPR system has been packaged into both AAV and lentivirus and used successfully in several mouse studies (Yin et al. 2016; Senis et al. 2014; Wang et al. 2015; Graham 2016). Nanoparticles and purified proteins are additional methods that have been used to successfully deliver CRISPR into cells and tissues (Wang et al. 2016; Ramakrishna et al. 2014). Each of these delivery methods has advantages and disadvantages, but with additional optimization successful gene replacement therapy in vivo should be possible. Since early HD diagnosis is possible, genetic correction therapy could be performed during the asymptomatic stage, potentially preventing onset of the disease.
Conclusion
Genome engineering is providing neuroscientists with new methods to address critical questions in the field and offers the hope for new treatments of neurological genetic diseases. The application of genetic engineering to disease modeling is accelerating efforts to understand the molecular mechanism of these diseases and offers new approaches to identifying therapeutic targets and drugs. The recent advances in genetic engineering allow for better modeling and understanding the role of SNPs in diseases with complex genetic alterations. These new genomic engineering technologies, which precisely alter the genome, are already offering insights into the complexity of the nervous system, its normal function and alterations in disease. Eventually these genome engineering technologies may correct the disease allele in human patients (in vivo) before symptoms manifest, resulting in therapy at the DNA level.
References
An MC, Zhang N, Scott G, Montoro D, Wittkop T, Mooney S, Melov S, Ellerby LM (2012) Genetic correction of Huntington’s disease phenotypes in induced pluripotent stem cells. Cell Stem Cell 11:253–263
An MC, O’Brien RN, Zhang N, Patra BN, De La Cruz M, Ray A, Ellerby LM (2014) Polyglutamine disease modeling: epitope based screen for homologous recombination using CRISPR/Cas9 System. PLoS Curr 6. doi: 10.1371/currents.hd.0242d2e7ad72225efa72f6964589369a
Bailus BJ, Pyles B, McAlister MM, O’Geen H, Lockwood SH, Adams AN, Nguyen JTT, Yu A, Berman RF, Segal DJ (2016) Protein delivery of an artificial transcription factor restores widespread Ube3a expression in an Angelman syndrome mouse brain. Mol Ther 24:548–555
Bakondi B, Lv W, Lu B, Jones MK, Tsai Y, Kim KJ, Levy R, Akhtar AA, Breunig JJ, Svendsen CN, Wang S (2016) In vivo CRISPR/Cas9 gene editing corrects retinal dystrophy in the S334ter-3 rat model of autosomal dominant retinitis pigmentosa. Mol Ther 24:556–563
Baltimore D, Berg P, Botchan M, Carroll D, Charo RA, Church G, Corn JE, Daley GQ, Doudna JA, Fenner JA, Greely HT, Jinek M, Martin GS, Penhoet E, Puck J, Sternberg SH, Weissman JS, Yamamoto KR (2015) Biotechnology. A prudent path forward for genomic engineering and germline gene modification. Science 348:36–38
Beerli RR, Segal DJ, Dreier B, Barbas CF III (1998) Toward controlling gene expression at will: specific regulation of the erbB-2/HER-2 promoter by using polydactyl zinc finger proteins constructed from modular building blocks. Proc Natl Acad Sci USA 95:14628–14633
Berg P, Baltimore D, Brenner S, Roblin RO 3rd, Singer MF (1975a) Asilomar conference on recombinant DNA molecules. Science 188:991–994
Berg P, Baltimore D, Brenner S, Roblin RO, Singer MF (1975b) Summary statement of the Asilomar conference on recombinant DNA molecules. Proc Natl Acad Sci USA 72:1981–1984
Boch J, Scholze H, Schornack S, Landgraf A, Hahn S, Kay S, Lahaye T, Nickstadt A, Bonas U (2009) Breaking the code of DNA binding specificity of TAL-type III effectors. Science 326:1509–1512
Calvanese V, Lara E, Kahn A, Fraga MF (2009) The role of epigenetics in aging and age-related diseases. Ageing Res Rev 8:268–276
Canals JM, Pineda JR, Torres-Peraza JF, Bosch M, Martín-Ibañez R, Muñoz MT, Mengod G, Ernfors P, Alberch J (2004) Brain-derived neurotrophic factor regulates the onset and severity of motor dysfunction associated with enkephalinergic neuronal degeneration in Huntington’s disease. J Neurosci 24:7727–7739
Chen Y, Zheng Y, Kang Y, Yang W, Niu Y, Guo X, Tu Z, Si C, Wang H, Xing R, Pu X, Yang SH, Li S, Ji W, Li XJ (2015) Functional disruption of the dystrophin gene in rhesus monkey using CRISPR/Cas9. Hum Mol Genet 24:3764–3774
Chinnery PF, Elliott HR, Hudson G, Samuels DC, Relton CL (2012) Epigenetics, epidemiology and mitochondrial DNA diseases. Int J Epidemiol 41:177–187
Christian M, Cermak T, Doyle EL, Schmidt C, Zhang F, Hummel A, Bogdanove AJ, Voytas DF (2010) Targeting DNA double-strand breaks with TAL effector nucleases. Genetics 186:757–761
Conforti P, Ramos C, Apostol BL, Simmons DA, Nguyen HP, Riess O, Thompson LM, Zuccato C, Cattaneo E (2008) Blood level of brain-derived neurotrophic factor mRNA is progressively reduced in rodent models of Huntington’s disease: restoration by the neuroprotective compound CEP-1347. Mol Cell Neurosci 39:1–7
Cong L, Zhou R, Kuo YC, Cunniff M, Zhang F (2012) Comprehensive interrogation of natural TALE DNA-binding modules and transcriptional repressor domains. Nat Commun 3:968
Cornu TI, Cathomen T (2010) Quantification of zinc finger nuclease-associated toxicity. Methods Mol Biol 649:237–245
Cornu TI, Thibodeau-Beganny S, Guhl E, Alwin S, Eichtinger M, Joung JK, Cathomen T (2008) DNA-binding specificity is a major determinant of the activity and toxicity of zinc-finger nucleases. Mol Ther 16:352–358
Dreier B, Beerli RR, Segal DJ, Flippin JD, Barbas CF 3rd (2001) Development of zinc finger domains for recognition of the 5′-ANN-3′ family of DNA sequences and their use in the construction of artificial transcription factors. J Biol Chem 276:29466–29478
Eisenstein M (2012) Sangamo’s lead zinc-finger therapy flops in diabetic neuropathy. Nat Biotechnol 30:121–123
Fink KD, Deng P, Gutierrez J, Anderson JS, Torrest A, Komarla A, Kalomoiris S, Cary W, Anderson JD, Gruenloh W, Duffy A, Tempkin T, Annett G, Wheelock V, Segal DJ, Nolta JA (2016) Allele-specific reduction of the mutant Huntingtin allele using transcription activator-like effectors in human Huntington’s disease fibroblasts. Cell Transplant 25:677–686
Gabriel R, Lombardo A, Arens A, Miller JC, Genovese P, Kaeppel C, Nowrouzi A, Bartholomae CC, Wang J, Friedman G, Holmes MC, Gregory PD, Glimm H, Schmidt M, Naldini L, von Kalle C (2011) An unbiased genome-wide analysis of zinc-finger nuclease specificity. Nat Biotechnol 29:816–823
Geurts AM, Cost GJ, Freyvert Y, Zeitler B, Miller JC, Choi VM, Jenkins SS, Wood A, Cui X, Meng X, Vincent A, Lam S, Michalkiewicz M, Schilling R, Foeckler J, Kalloway S, Weiler H, Ménoret S, Anegon I, Davis GD, Zhang L, Rebar EJ, Gregory PD, Urnov FD, Jacob HJ, Buelow R (2009) Knockout rats via embryo microinjection of zinc-finger nucleases. Science 325:433
Gilbert LA, Larson MH, Morsut L, Liu Z, Brar GA, Torres SE, Stern-Ginossar N, Brandman O, Whitehead EH, Doudna JA, Lim WA, Weissman JS, Qi LS (2013) CRISPR-mediated modular RNA-guided regulation of transcription in eukaryotes. Cell 154:442–451
Graham DM (2016) CRISPR/Cas9 corrects retinal dystrophy in rats. Lab Anim (NY) 45:85
Guilinger JP, Pattanayak V, Reyon D, Tsai SQ, Sander JD, Joung JK, Liu DR (2014) Broad specificity profiling of TALENs results in engineered nucleases with improved DNA-cleavage specificity. Nat Methods 11:429–435
Gupta A, Meng X, Zhu LJ, Lawson ND, Wolfe SA (2010) Zinc finger protein-dependent and -independent contributions to the in vivo off-target activity of zinc finger nucleases. Nucleic Acids Res 39:381–392
Hsu PD, Scott DA, Weinstein JA, Ran FA, Konermann S, Agarwala V, Li Y, Fine EJ, Wu X, Shalem O, Cradick TJ, Marraffini LA, Bao G, Zhang F (2013) DNA targeting specificity of RNA-guided Cas9 nucleases. Nat Biotechnol 31:827–832
Inglese J, Dranchak P, Moran JJ, Jang SW, Srinivasan R, Santiago Y, Zhang L, Guha R, Martinez N, MacArthur R, Cost GJ, Svaren J (2014) Genome editing-enabled HTS assays expand drug target pathways for Charcot-Marie-tooth disease. ACS Chem Biol 9:2594–2602
Jinek M, Chylinski K, Fonfara I, Hauer M, Doudna JA, Charpentier E (2012) A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 337:816–821
Keatinge M, Bui H, Menke A, Chen YC, Sokol AM, Bai Q, Ellett F, Da Costa M, Burke D, Gegg M, Trollope L, Payne T, McTighe A, Mortiboys H, de Jager S, Nuthall H, Kuo MS, Fleming A, Schapira AH, Renshaw SA, Highley JR, Chacinska A, Panula P, Burton EA, O’Neill MJ, Bandmann O (2015) Glucocerebrosidase 1 deficient Danio rerio mirror key pathological aspects of human Gaucher disease and provide evidence of early microglial activation preceding alpha-synuclein-independent neuronal cell death. Hum Mol Genet 24:6640–6652
Landles C, Bates GP (2004) Huntingtin and the molecular pathogenesis of Huntington’s disease. Fourth in molecular medicine review series. EMBO Rep 5:958–963
Langfelder P, Cantle PJ, Chatzopoulou D, Wang N, Gao F, Al-Ramahi I, XH L, Ramos EM, El-Zein K, Zhao Y, Deverasetty S, Tebbe A, Schaab C, Lavery DJ, Howland D, Kwak S, Botas J, Aaronson JS, Rosinski J, Coppola G, Horvath S, William Yang X (2016) Integrated genomics and proteomics define Huntingtin CAG length-dependent networks in mice. Nat Neurosci 19:623–633
Larson MH, Gilbert LA, Wang X, Lim WA, Weissman JS, Qi LS (2013) CRISPR interference (CRISPRi) for sequence-specific control of gene expression. Nat Protoc 8:2180–2196
Liang P, Xu Y, Zhang X, Ding C, Huang R, Zhang Z, Lv J, Xie X, Chen Y, Li Y, Sun Y, Bai Y, Songyang Z, Ma W, Zhou C, Huang L (2015) CRISPR/Cas9-mediated gene editing in human tripronuclear zygotes. Protein Cell 6:363–372
Liu Q, Segal DJ, Ghiara JB, Barbas CF III (1997) Design of polydactyl zinc-finger proteins for unique addressing within complex genomes. Proc Natl Acad Sci USA 94:5525–5530
Liu J, Gao C, Chen W, Ma W, Li X, Shi Y, Zhang H, Zhang L, Long Y, Xu H, Guo X, Deng S, Yan X, Yu D, Pan G, Chen Y, Lai L, Liao W, Li Z (2016) CRISPR/Cas9 facilitates investigation of neural circuit disease using human iPSCs: mechanism of epilepsy caused by an SCN1A loss-of-function mutation. Transl Psychiatry 6:e703
Maeder ML, Angstman JF, Richardson ME, Linder SJ, Cascio VM, Tsai SQ, Ho QH, Sander JD, Reyon D, Bernstein BE, Costello JF, Wilkinson MF, Joung JK (2013a) Targeted DNA demethylation and activation of endogenous genes using programmable TALE-TET1 fusion proteins. Nat Biotechnol 31:1137–1142
Maeder ML, Linder SJ, Reyon D, Angstman JF, Fu Y, Sander JD, Joung JK (2013b) Robust, synergistic regulation of human gene expression using TALE activators. Nat Methods 10:243–245
Moscou MJ, Bogdanove AJ (2009) A simple cipher governs DNA recognition by TAL effectors. Science 326:1501
Nagaoka M, Sugiura Y (2000) Artificial zinc finger peptides: creation, DNA recognition, and gene regulation. J Inorg Biochem 82:57–63
Nakamura K, Fujii W, Tsuboi M, Tanihata J, Teramoto N, Takeuchi S, Naito K, Yamanouchi K, Nishihara M (2014) Generation of muscular dystrophy model rats with a CRISPR/Cas system. Sci Rep 4:5635
Nelson CE et al (2016) In vivo genome editing improves muscle function in a mouse model of Duchenne muscular dystrophy. Science 351:403–407
O’Brien R, DeGiacomo F, Holcomb J, Bonner A, Ring KL, Zhang N, Zafar K, Weiss A, Lager B, Schilling B, Gibson BW, Chen S, Kwak S, Ellerby LM (2015) Integration-independent transgenic Huntington Disease fragment mouse models reveal distinct phenotypes and life span in vivo. J Biol Chem 290:19287–19306
O’Geen H, Henry IM, Bhakta MS, Meckler JF, Segal DJ (2015) A genome-wide analysis of Cas9 binding specificity using ChIP-seq and targeted sequence capture. Nucleic Acids Res 43:3389–3404
Pabo CO, Peisach E, Grant RA (2001) Design and selection of novel Cys2his2 zinc finger proteins. Annu Rev Biochem 70:313–340
Platt RJ, Chen S, Zhou Y, Yim MJ, Swiech L, Kempton HR, Dahlman JE, Parnas O, Eisenhaure TM, Jovanovic M, Graham DB, Jhunjhunwala S, Heidenreich M, Xavier RJ, Langer R, Anderson DG, Hacohen N, Regev A, Feng G, Sharp PA, Zhang F (2014) CRISPR-Cas9 knockin mice for genome editing and cancer modeling. Cell 159:440–455
Pollock L, Dahlenburg H, Nelson H, Fink KD, Cary W, Hendrix K, Annett G, Torrest A, Deng P, Gutierrez J, Nacey C, Pepper K, Kalomoiris S, D Anderson J, McGee J, Gruenloh W, Fury B, Bauer G, Duffy A, Tempkin T, Wheelock V, Nolta JA (2016) Human mesenchymal stem cells genetically engineered to overexpress brain-derived neurotrophic factor improve outcomes in Huntington’s disease mouse models. Mol Ther 24:965–977
Qi LS, Larson MH, Gilbert LA, Doudna JA, Weissman JS, Arkin AP, Lim WA (2013) Repurposing CRISPR as an RNA-guided platform for sequence-specific control of gene expression. Cell 152:1173–1183
Ramakrishna S, Kwaku Dad AB, Beloor J, Gopalappa R, Lee SK, Kim H (2014) Gene disruption by cell-penetrating peptide-mediated delivery of Cas9 protein and guide RNA. Genome Res 24:1020–1027
Ran FA, Hsu PD, Lin CY, Gootenberg JS, Konermann S, Trevino AE, Scott DA, Inoue A, Matoba S, Zhang Y, Zhang F (2013) Double nicking by RNA-guided CRISPR Cas9 for enhanced genome editing specificity. Cell 154:1380–1389
Ran FA, Cong L, Yan WX, Scott DA, Gootenberg JS, Kriz AJ, Zetsche B, Shalem O, Wu X, Makarova KS, Koonin EV, Sharp PA, Zhang F (2015) In vivo genome editing using Staphylococcus aureus Cas9. Nature 520:186–191
Richardson CD, Ray GJ, DeWitt MA, Curie GL, Corn JE (2016) Enhancing homology-directed genome editing by catalytically active and inactive CRISPR-Cas9 using asymmetric donor DNA. Nat Biotechnol 34:339–344
Ring KL, An MC, Zhang N, O’Brien RN, Ramos EM, Gao F, Atwood R, Bailus BJ, Melov S, Mooney SD, Coppola G, Ellerby LM (2015) Genomic analysis reveals disruption of striatal neuronal development and therapeutic targets in human Huntington’s disease neural stem cells. Stem Cell Rep 5:1023–1038
Rivenbark AG, Stolzenburg S, Beltran AS, Yuan X, Rots MG, Strahl BD, Blancafort P (2012) Epigenetic reprogramming of cancer cells via targeted DNA methylation. Epigenetics 7:350–360
Sander JD, Joung JK (2014) CRISPR-Cas systems for editing, regulating and targeting genomes. Nat Biotechnol 32:347–355
SangamoBioSciences (2001) Sangamo BioSciences announces presentation of first preclinical data using ZFP transcription factors. http://www.sangamo.com/053101giordano.htm
Segal DJ, Barbas CF III (2000) Design of novel sequence-specific DNA-binding proteins. Curr Opin Chem Biol 4:34–39
Segal DJ, Barbas CF III (2001) Custom DNA-binding proteins come of age: polydactyl zinc-finger proteins. Curr Opin Biotechnol 12:632–637
Segal DJ, Dreier B, Beerli RR, Barbas CF III (1999) Toward controlling gene expression at will: selection and design of zinc finger domains recognizing each of the 5′-GNN-3′ DNA target sequences. Proc Natl Acad Sci USA 96:2758–2763
Senis E, Fatouros C, Große S, Wiedtke E, Niopek D, Mueller AK, Börner K, Grimm D (2014) CRISPR/Cas9-mediated genome engineering: an adeno-associated viral (AAV) vector toolbox. Biotechnol J 9:1402–1412
Soldner F, Laganière J, Cheng AW, Hockemeyer D, Gao Q, Alagappan R, Khurana V, Golbe LI, Myers RH, Lindquist S, Zhang L, Guschin D, Fong LK, BJ V, Meng X, Urnov FD, Rebar EJ, Gregory PD, Zhang HS, Jaenisch R (2011) Generation of isogenic pluripotent stem cells differing exclusively at two early onset Parkinson point mutations. Cell 146:318–331
Tabebordbar M, Zhu K, Cheng JK, Chew WL, Widrick JJ, Yan WX, Maesner C, Wu EY, Xiao R, Ran FA, Cong L, Zhang F, Vandenberghe LH, Church GM, Wagers AJ (2016) In vivo gene editing in dystrophic mouse muscle and muscle stem cells. Science 351:407–411
Urnov FD, Miller JC, Lee YL, Beausejour CM, Rock JM, Augustus S, Jamieson AC, Porteus MH, Gregory PD, Holmes MC (2005) Highly efficient endogenous human gene correction using designed zinc-finger nucleases. Nature 435:646–651
Wang H, Yang H, Shivalila CS, Dawlaty MM, Cheng AW, Zhang F, Jaenisch R (2013) One-step generation of mice carrying mutations in multiple genes by CRISPR/Cas-mediated genome engineering. Cell 153:910–918
Wang X, Wang Y, Wu X, Wang J, Wang Y, Qiu Z, Chang T, Huang H, Lin RJ, Yee JK (2015) Unbiased detection of off-target cleavage by CRISPR-Cas9 and TALENs using integrase-defective lentiviral vectors. Nat Biotechnol 33:175–178
Wang M, Zuris JA, Meng F, Rees H, Sun S, Deng P, Han Y, Gao X, Pouli D, Wu Q, Georgakoudi I, Liu DR, Xu Q (2016) Efficient delivery of genome-editing proteins using bioreducible lipid nanoparticles. Proc Natl Acad Sci USA 113:2868–2873
Wiedenheft B, Sternberg SH, Doudna JA (2012) RNA-guided genetic silencing systems in bacteria and archaea. Nature 482:331–338
Wolfe SA, Ramm EI, Pabo CO (2000) Combining structure-based design with phage display to create new Cys(2)His(2) zinc finger dimers. Struct Fold Des 8:739–750
Wolffe E (2016) Corporate profile: Sangamo BioSciences, Inc. Regen Med 11:375–379
Wu X, Scott DA, Kriz AJ, Chiu AC, Hsu PD, Dadon DB, Cheng AW, Trevino AE, Konermann S, Chen S, Jaenisch R, Zhang F, Sharp PA (2014) Genome-wide binding of the CRISPR endonuclease Cas9 in mammalian cells. Nat Biotechnol 32:670–676
Xia G, Gao Y, Jin S, Subramony SH, Terada N, Ranum LP, Swanson MS, Ashizawa T (2015) Genome modification leads to phenotype reversal in human myotonic dystrophy type 1 induced pluripotent stem cell-derived neural stem cells. Stem Cells 33:1829–1838
Xu L, Park KH, Zhao L, Xu J, El Refaey M, Gao Y, Zhu H, Ma J, Han R (2016) CRISPR-mediated genome editing restores dystrophin expression and function in mdx mice. Mol Ther 24:564–569
Yin H, Song CQ, Dorkin JR, Zhu LJ, Li Y, Wu Q, Park A, Yang J, Suresh S, Bizhanova A, Gupta A, Bolukbasi MF, Walsh S, Bogorad RL, Gao G, Weng Z, Dong Y, Koteliansky V, Wolfe SA, Langer R, Xue W, Anderson DG (2016) Therapeutic genome editing by combined viral and non-viral delivery of CRISPR system components in vivo. Nat Biotechnol 34:328–333
Zuccato C, Marullo M, Conforti P, MacDonald ME, Tartari M, Cattaneo E (2008) Systematic assessment of BDNF and its receptor levels in human cortices affected by Huntington’s disease. Brain Pathol 18:225–238
Zykovich A, Korf I, Segal DJ (2009) Bind-n-Seq: high-throughput analysis of in vitro protein-DNA interactions using massively parallel sequencing. Nucleic Acids Res 37:e151
Acknowledgments
Support for this work comes from NIH R01s NS094422 and NS100529.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2017 The Author(s)
About this chapter
Cite this chapter
Bailus, B., Zhang, N., Ellerby, L.M. (2017). Using Genome Engineering to Understand Huntington’s Disease. In: Jaenisch, R., Zhang, F., Gage, F. (eds) Genome Editing in Neurosciences. Research and Perspectives in Neurosciences. Springer, Cham. https://doi.org/10.1007/978-3-319-60192-2_9
Download citation
DOI: https://doi.org/10.1007/978-3-319-60192-2_9
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-60191-5
Online ISBN: 978-3-319-60192-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)