Abstract
The use of aquatic model organisms has been greatly diversified in laboratories. Zebrafish is the most advanced aquatic species for the use of Crispr-Cas9 in laboratories. Because of the simplicity and broad applicability of this later system, knock-out is now efficiently performed at medium scale. Forward genetics in zebrafish can now be performed by CRISPR-based F0 screening using high speed and high content phenotyping for example by confocal imaging. As zebrafish, marine model organisms have the prominent advantage to be transparent, all the more at young stages (embryos and larvae) or when fixed samples are cleared by novel methods. The Cripsr-Cas9 system is routinely used in the ascidian Ciona intestinalis. It also starts to be used in many other marine models, such as the medusa Clythia hemispherica. We provide at the end of this review a list of aquatic model species and some examples of questions on the origin of our nervous system that can be coped with these models, where the possibility to perform genome editing would constitute a major advance.
You have full access to this open access chapter, Download chapter PDF
Similar content being viewed by others
Keywords
These keywords were added by machine and not by the authors. This process is experimental and the keywords may be updated as the learning algorithm improves.
Introduction
With the expansion of biochemistry and molecular biology during the twentieth century, researchers focused more and more on a few model organisms such as nematodes, fruit flies, or mice. These models were amenable to many experimental approaches in molecular biology and biochemistry. Recently, a novel species, the zebrafish, has emerged as a major laboratory model. Initially selected because its transparent embryo is an excellent system in which to study development, it has now become the second most used animal in laboratories worldwide. Thus, the current applications of zebrafish studies are now highly diversified in neurobiology, immunology, adult physiology, oncology, and regenerative medicine, exploiting its advantages for in vivo approaches and imaging.
In parallel, due to the explosion of sequencing methods, there has been a clear trend towards the diversification of model organisms, especially those used in neurosciences. In this paper, we will focus on applications of such new models in evolution of development or biomedical research. There are several reasons to use an increasing number of model organisms: first, the classical models are not representative of most branches of the tree of life, and second, many questions now need to be studied in vivo and established models are not always well adapted to many of those biological or medical questions.
We will here elaborate why the use of genome editing in these models offers revolutionary perspectives. Future genome-editing experiments should indeed allow us to unveil the function of critical genes in almost any species; this approach was previously restricted to a few model species, or even restricted to mouse for most precise modifications by homologous recombination. With genome editing, it will become possible with functional data to study the evolutionary origin of highly diversified cell types such as neurons and, in addition, to interrogate how extremely complex cellular organizations such as those found in the brain were built in the course of evolution.
Zebrafish: With the CRiSPR-Cas9 System, Forward Genetic Screens Are Back Again
Zebrafish is a vertebrate, so it has a body plan fundamentally similar to ours (Onai et al. 2014): like humans, zebrafish have a notochord that is a central pile of turgescent cells that confers rigidity to embryo and larvae and serves as the support axis for the development of the spine. Muscles are located on both sides of the notochord. The nervous system is found dorsally and intestine ventrally. Zebrafish embryos, during the so-called “phylotypic stage” (Slack et al. 1993), use the same developmental pattern as human embryos, involving the colinear activation with time and space of Hox gene expressions to build axial structures.
Many organs, including brain, also have similar general organisations in zebrafish and humans. Hence, various aspects of neurobiology can be studied in this species. Zebrafish has a tripartite brain. Brains, as other organs in fish such as fins or hearts, regenerate after a mechanical injury. Some complex behaviours like fear (Amo et al. 2014), social behaviour (Chou et al. 2016) and memory can be studied in adults. Additionally, more basic behaviours can be studied in the 5-day larvae, at a stage most amenable for imaging and for which no authorization for animal experimentation is required (Naumann et al. 2016). This makes the model easy to use for applications such as neurotoxicology performed by academic labs or cosmetology companies.
At the end of the twentieth century, the zebrafish was suggested to be a promising model for genetic screens relevant to human diseases (Mullins et al. 1994). However, during the evolution of vertebrates, an additional genome duplication occurred in the teleost fish lineage, leading to the presence of many duplicated genes in fish genomes that greatly complicate the analysis of screen data. Another pitfall was that the zebrafish has maternal factors carried by the egg. So, when a gene is mutated, the effect of the mutation is in most cases not visible at early stages because maternal factors are present to insure correct development. Therefore, a disappointingly low number of mutations were identified following large-scale screens by random mutagenesis in embryos. Because many zebrafish screens were performed during early larval stages, most identified mutants failed to exhibit phenotypes similar to human rare diseases, which often appear much later in human life.
After 2000, zebrafish became a very useful model for reverse genetics when phenotypical analyses of mutants were performed with spectacular time-lapse imaging in live fish, for example, during early development (Olivier et al. 2010), neurogenesis (Barbosa et al. 2015), hematopoeisis (Renaud et al. 2011) or immune response (Levraud et al. 2014).
Nowadays, the zebrafish is the most advanced aquatic species for the use of Cripsr-Cas9 in laboratories (Shah and Moens 2016). In this species, however, a challenge remains: efficient insertions of point mutations (for example, to mimic human missense mutations) are still generated at low rates (Renaud et al. 2016). For these applications, the very fast early development of the zebrafish is a drawback, as it probably makes the repair events following DNA cutting by the CRISPR protein highly mosaic and hard to detect in the progeny. Improvements to target the repair construct to the nucleus of the one-cell stage embryo will have to be developed. Modified oligonucleotides could improve KI rates. Alternatively, plasmidic constructs with long homology arms have been used in a recently published method (Hoshijima et al. 2016) to perform KI at large scale in zebrafish; unfortunately, this method remains labor intensive.
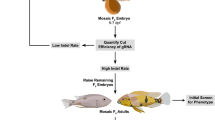
Knock-out is now efficiently performed in zebrafish at medium scale (Shah et al. 2015). Hence so-called forward genetics in zebrafish again seems to have a bright future. To study the molecular basis of a given phenotype in a particular cell type, large gene families can be targeted for mutations, and, importantly, mutations of duplicated genes and of their close paralogs can be performed jointly, due to the possibility of injecting arrays of CRISPR guide RNAs.
For large-scale forward screens, methods of large-scale phenotyping, at first hand by 3D imaging, still need to be optimized. Thus, a current priority for zebrafish researchers is to improve rapid imaging technologies at large scale and at later stages, to make zebrafish a better model. Such a model would provide a perfect context for analyzing large collections of mutants. Tissue-clearing methods (Seo et al. 2016) and high-speed imaging methods, such as those using highly sensitive video cameras, have recently emerged and will certainly be crucial for these approaches.
Optimizing the Cripsr-Cas9 System in Transparent Marine Animals
Most marine model organisms have the obvious advantages, as zebrafish, to have transparent embryos and larvae, a feature selected in water throughout evolution to escape predators. Transparency is crucial for the microscopic analysis of development in these models. Eggs can generally be obtained in huge numbers. Although they are sometimes quite big because of the presence of vitelline reserves, embryos are composed of only a few cells and the lineage analysis is thus easy in these simple and compact embryos. In ascidian embryos, for example, the notochord only has 64 cells. Significant progress in understanding human cardiac developmental gene network was made in ascidian models. This unique insight provided direction for the reprograming of cell lineages in human cell cultures: following the observation that Ci-es1/2 and Ci-mesp generated cardiac progenitors in ascidians, researchers transdifferentiated human dermal fibroblasts into cardiac progenitors (Islas et al. 2012).
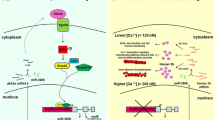
The Cripsr-Cas9 system was used in a study using the ascidian Ciona intestinalis (Stolfi et al. 2014). This study, from Lionel Christiaen’s group, reported the success of tissue-specific genome editing in this species. Optimization of plasmid constructs was performed, in which specific ubiquitous U6 promoters were used to drive guide RNA expression, and tissue-specific promoters were designed to drive the expression of the Cas9 protein. Introducing the CRISPR–Cas9 components in ascidians was quite easy because a large number of eggs could be electroporated with plasmid DNA, producing both the CRISPR protein and the guide RNAs. Nevertheless, while breeding of this species in inland laboratories has been performed (Joly et al. 2007), it remains difficult. Improvements are still required to reliably obtain the culture of stable lines of transgenic animals.
The Cripsr-Cas9 system is beginning to be used in many other aquatic model organisms (for a fascinating example in lampreys, Square et al. 2015). For example, fascinating experiments (unpublished) have been performed in the medusa Clytia hemisphaerica (Tsuyioshi Momose, CNRS Villefranche-sur-Mer, personal communication). This model recently emerged as a remarkable cnidarian species useful for evo-devo studies (Houliston et al. 2010). Experiments were in part supported by the French network named “Etude Fonctionnelle sur les Organismes Modèles (EFOR, www.efor.net)”, promoting research—including genome editing and imaging—in metazoan model organisms.
Success in Clythia is due to obtaining full life cycles in laboratory aquaria, generating quasi-immortal, vegetatively growing colonies. Moreover, adult medusae spawn daily, generating transparent and easy to inject eggs. Rates of Cripsr-Cas9 knock-outs are strikingly high: over 700 embryos, all with potential knock-outs as seen by the loss of fluorescent protein activity, can be generated in a single injection experiment.
Injection of the NLS Cas9 protein/sgRNA into unfertilized eggs can be performed as soon as 1 hour after ovulation and before subsequent fertilization. In this condition, the Cas9 protein probably has time to be targeted to its cutting site before the first division of the embryo occurs. One first exciting application of this method was the deletion of green fluorescent proteins, making newly generated transgenic lines suitable for imaging applications. Indeed, in these species, endogenous fluorescent activity hinders potential observation of GFP in newly generated transgenic animals. The availability of mutants in such species will offer novel routes for fundamental research in evolution and development (Galliot et al. 2009). In such non-marine species, a challenge remains to keep animals alive in captivity long term. Significant investments will also be needed to obtain colonies of inbred lines, which should become reference laboratory lines for the corresponding communities of worldwide researchers.
More and More Aquatic Model Organisms for Diversified Uses
Examples of emerging aquatic models of increasing evolutionary distances from vertebrates are described below. These models are located at several key nodes of the metazoan branches of the tree of life. Alternative fish models to zebrafish (Schartl 2014) allow the study of particular evolutionary processes, such as adaptation to cave life in the Astyanax mexicanus. To study the emergence of synapomorphic (specific and shared) vertebrate features in evolution, lampreys (including the sea lamprey, Petromyzon marinus, an agnathan), lancelets (the cephalochordate amphioxus, Branchiostoma lanceolatum) or tunicates (such as the urochordate Ciona intestinalis) are very relevant models. Other more distantly related bilaterians such as the polychaete annelid Platynereis dumerilii provide insight into which features were already present in the common ancestor of all bilaterian organisms, the so-called “urbilateria.” Even more distant metazoans, with no bilateral symmetry but rather radial symmetry, are also used in laboratories. Thus, Ctenophores (sea gooseberries) and Cnidarians (corals, jellyfish, sea anemones) can be used to study ancient features of nerve cell types. Also, sponges and placozoans constitute fascinating basal Metazoans, with no nervous system.
In Biomedical Research, Why and How Should We Use Aquatic Models to Study Diseases of the Nervous System?
A first obvious use of model organisms is to generate so-called “models of diseases.” In most cases, these models are mutants or transgenic animals that reproduce at best pathological conditions observed in humans, such as neural degeneration. With the advent of precise genome-editing methods, the capacity to generate point mutations in any model by introducing a repair construct bearing the mutations will constitute a true revolution. Indeed, mutations at orthologous positions to variations found in human diseases can be generated in these model animals if genomes can be aligned in the region surrounding the mutation. Then the phenotypical effects of the abnormal protein function can be described, for example, using live imaging to observe abnormal cell behaviors, such as proliferation or migration. Therein resides the great advantage of these aquatic models, with a diversity of developmental and genomic contexts and transparency allowing easy imaging.
Moreover, in an applied perspective in regenerative medicine, understanding how cell types emerged during evolution helps us identify crucial pathways that are, for example, active in stem cells in normal and pathological conditions. The Crispr-Cas9 system applied to aquatic model organisms will offer an unprecedented opportunity to characterize the key genes and pathways that are active following injury or degeneration. They promote regeneration responses in animals with regenerative capacities and could be useful in regenerative medicine, if (re)activated in humans (Karra and Poss 2017).
A Short Natural History of the Nervous System: Several Questions on Its Origin
This chapter provides examples of what marine model organisms bring us in the domain of neuroscience. Many essential questions can indeed be examined with these models, providing us more knowledge about how the human brain was shaped through evolution, and this should help us better understand pathologies and their pleiotropic effects.
Evolution indeed sifts through the noise and allows us to focus on key genes and pathways that have remained crucial for specific cell types throughout evolution. Looking at extant species, and describing common features that are likely to be ancestral and shared, is also a way to “reconstruct” the nervous system of the putative last common ancestor between the two compared species. In this domain, the absence of fossils of nervous system and brains has impeded researchers.
According to Detlev Arendt (Arendt 2008), homology hypotheses are based on the comparison of genes, cytological features and ontological location in the body of the embryo. Functional experiments with genome editing in model organisms will add extremely important cues to this domain.
The Ctenophores are colorful planctonic animals that have a sophisticated nervous net allowing them to swim and to emit beautiful waves of fluorescent flashes. A long-standing debate is whether these animals have neuron-like cells, which would have appeared independently of the neurons of our nervous system during evolution. In this respect, examining the phylogenetic position of these animals is primordial: are they closely related to bilaterians or rahter do they form an out-group of metazoans? If they are more distantly related to us than sponges, this would indeed mean that the nervous system was invented twice in evolution, because it is very unlikely that sponges, which would be closer relatives to bilaterians, secondarily lost neural cells.
Until this controversy is resolved, it will not be possible to know whether there were two independent origins of the nervous system in animals, which would, of course, be a very exciting possibility. However, as argued in a recent review (Jager and Manuel 2016), many lines of evidence now suggest that ctenophores are closely related to bilaterians and that neurons appeared only once in evolution. In favour of a single nervous system type is the presence of the well-known neurogenesis SoxB gene and the presence of acetylcholine and numerous common GPCRs. In any case, ctenophores should provide key insights into deeply conserved features of animal neural cells.
Marine organisms also allow us to examine how the nervous system became condensed and centralized (Arendt et al. 2016) while becoming much more complex in the course of evolution to terrestrial life (Nomaksteinsky et al. 2009). Starting from a diffuse nerve net in a swimming larva, brains became huge, formed from the so-called embryonic neural plate in vertebrates later undergoing neurulation to form the neural tube.
Another question about the origin of the nervous system is how the vertebrate tripartite nervous system arose in the course of evolution. The vertebrate central nervous system is composed of anterior brain, posterior brain and spinal cord. Such organisation occurred with the emergence of borders between brain domains, such as the midbrain/hindbrain boundary. Recently, following studies in an annelid worm, Arendt and colleagues proposed that the nervous system of the common bilaterian ancestor was probably composed of two independent domains (corresponding to the vertebrate forebrain/midbrain and hindbrain) that later fused during evolution (Tosches and Arendt 2013).
Very recently, and in line with Arendt’s hypothesis, Chris Lowe’s group proposed that the adult body plan of an indirect developing hemichordate develops by adding a Hox pattern trunk to an anterior larval territory, confirming the hypothesis that marine larvae are “swimming heads” (Gonzalez et al. 2017).
Concerning the origin of vertebrate synapomorphic characters, ascidians provide evidence of the precraniate or prevertebrate origins of the neural crest. Neural crest cells are stem cells with migratory behaviors and the capacity to differentiate into incredibly diversified tissues and locations. The evolutionary origin of neural crest is obscure. They were first thought to be a vertebrate sinapomorphy. Bill Jeffery identified migratory pigment cells in ascidians (Jeffery et al. 2004; Jeffery 2006) and, more recently, Lionel Christiaen’s group identified migrating neuron precursors in the ascidian Ciona intestinalis. Interestingly, these precursors were shown to arise from the border of the neural plate, a hallmark of the neural crest in vertebrates (Stolfi et al. 2015).
Conclusion
Cripsr-Cas9 system and other genome-editing techniques can now be used in various aquatic model organisms for extremely diversified applications, which will lead to new models of brain diseases for biomedical research. Also, these models will provide strategies to characterize human genetic variations linked to diseases at large scale and suggest new avenues for regenerative medicine, because of the exceptional abilities of most aquatic models to regenerate. Basic biological knowledge will, of course, benefit from this revolution, and one might expect many more revolutionary discoveries from the exploration of the genomes of multiple animal aquatic species using Cripsr-Cas9 approaches.
References
Amo R, Fredes F, Kinoshita M, Aoki R, Aizawa H, Agetsuma M, Aoki T, Shiraki T, Kakinuma H, Matsuda M, Yamazaki M, Takahoko M, Tsuboi T, Higashijima S, Miyasaka N, Koide T, Yabuki Y, Yoshihara Y, Fukai T, Okamoto H (2014) The habenulo-raphe serotonergic circuit encodes an aversive expectation value essential for adaptive active avoidance of danger. Neuron 84:1034–1048
Arendt D (2008) The evolution of cell types in animals: emerging principles from molecular studies. Nat Rev Genet 9:868–882
Arendt D, Tosches MA, Marlow H (2016) From nerve net to nerve ring, nerve cord and brain evolution of the nervous system. Nat Rev Neurosci 17:61–72
Barbosa JS, Sanchez-Gonzalez R, Di Giaimo R, Baumgart EV, Theis FJ, Götz M, Neurodevelopment NJ (2015) Live imaging of adult neural stem cell behavior in the intact and injured zebrafish brain. Science 348:789–793
Chou MY, Amo R, Kinoshita M, Cherng BW, Shimazaki H, Agetsuma M, Shiraki T, Aoki T, Takahoko M, Yamazaki M, Higashijima S, Okamoto H (2016) Social conflict resolution regulated by two dorsal habenular subregions in zebrafish. Science 352:87–90
Galliot B, Quiquand M, Ghila L, de Rosa R, Miljkovic-Licina M, Chera S (2009) Origins of neurogenesis, a cnidarian view. Dev Biol 332:2–24
Gonzalez P, Uhlinger KR, Lowe CJ (2017) The adult body plan of indirect developing hemichordates develops by adding a Hox-patterned trunk to an anterior larval territory. Curr Biol 27:87–95
Hoshijima K, Jurynec MJ, Grunwald DJ (2016) Precise genome editing by homologous recombination. Meth Cell Biol 135:121–147
Houliston E, Momose T, Manuel M (2010) Clytia hemisphaerica: a jellyfish cousin joins the laboratory. Trends Genet 26:159–167
Islas JF, Liu Y, Weng KC, Robertson MJ, Zhang S, Prejusa A, Harger J, Tikhomirova D, Chopra M, Iyer D, Mercola M, Oshima RG, Willerson JT, Potaman VN, Schwartz RJ (2012) Transcription factors ETS2 and MESP1 transdifferentiate human dermal fibroblasts into cardiac progenitors. Proc Natl Acad Sci USA 109:13016–13021
Jager M, Manuel M (2016) Ctenophores: an evolutionary-developmental perspective. Curr Opin Genet Dev 39:85–92
Jeffery WR (2006) Ascidian neural crest-like cells: phylogenetic distribution, relationship to larval complexity, and pigment cell fate. J Exp Zool B Mol Dev Evol 306:470–480
Jeffery WR, Strickler AG, Yamamoto Y (2004) Migratory neural crest-like cells form body pigmentation in a urochordate embryo. Nature 431:696–699
Joly JS, Kano S, Matsuoka T, Auger H, Hirayama K, Satoh N, Awazu S, Legendre L, Sasakura Y (2007) Culture of Ciona intestinalis in closed systems. Dev Dyn 236:1832–1840
Karra R, Poss KD (2017) Redirecting cardiac growth mechanisms for therapeutic regeneration. J Clin Invest 127:427–436
Levraud JP, Palha N, Langevin C, Boudinot P (2014) Through the looking glass: witnessing host-virus interplay in zebrafish. Trends Microbiol 22:490–497
Mullins MC, Hammerschmidt M, Haffter P, Nüsslein-Volhard C (1994) Large-scale mutagenesis in the zebrafish: in search of genes controlling development in a vertebrate. Curr Biol 4:189–202
Naumann EA, Fitzgerald JE, Dunn TW, Rihel J, Sompolinsky H, Engert F (2016) From whole-brain data to functional circuit models: the zebrafish optomotor response. Cell 167:947–960
Nomaksteinsky M, Röttinger E, Dufour HD, Chettouh Z, Lowe CJ, Martindale MQ, Brunet JF (2009) Centralization of the deuterostome nervous system predates chordates. Curr Biol 19:1264–1269
Olivier N, Luengo-Oroz MA, Duloquin L, Faure E, Savy T, Veilleux I, Solinas X, Débarre D, Bourgine P, Santos A, Peyriéras N, Beaurepaire E (2010) Cell lineage reconstruction of early zebrafish embryos using label-free nonlinear microscopy. Science 329:967–971
Onai T, Irie N, Kuratani S (2014) The evolutionary origin of the vertebrate body plan: the problem of head segmentation. Annu Rev Genomics Hum Genet 15:443–459
Renaud O, Herbomel P, Kissa K (2011) Studying cell behavior in whole zebrafish embryos by confocal live imaging: application to hematopoietic stem cells. Nat Protoc 6:1897–1904
Renaud JB, Boix C, Charpentier M, De Cian A, Cochennec J, Duvernois-Berthet E, Perrouault L, Tesson L, Edouard J, Thinard R, Cherifi Y, Menoret S, Fontanière S, de Crozé N, Fraichard A, Sohm F, Anegon I, Concordet JP, Giovannangeli C (2016) Improved genome editing efficiency and flexibility using modified oligonucleotides with TALEN and CRISPR-Cas9 nucleases. Cell Rep 14:2263–2272
Schartl M (2014) Beyond the zebrafish: diverse fish species for modeling human disease. Dis Model Mech 7:181–192
Seo J, Choe M, Kim SY (2016) Clearing and labeling techniques for large-scale biological tissues. Mol Cells 39:439–446
Shah AN, Moens CB (2016) Approaching perfection: new developments in zebrafish genome engineering. Dev Cell 36:595–596
Shah AN, Davey CF, Whitebirch AC, Miller AC, Moens CB (2015) Rapid reverse genetic screening using CRISPR in zebrafish. Nat Methods 12:535–540
Slack JMW, Holland PWH, Graham CF (1993) The zootype and the phylotypic stage. Nature 361:490–492
Stolfi A, Ryan K, Meinertzhagen IA, Christiaen L (2015) Migratory neuronal progenitors arise from the neural plate borders in tunicates. Nature 527:371–374
Square T, Romášek M, Jandzik D, Cattell MV, Klymkowsky M, Medeiros DM (2015) CRISPR/Cas9-mediated mutagenesis in the sea lamprey Petromyzon marinus: a powerful tool for understanding ancestral gene functions in vertebrates. Development 142(23):4180–4187
Stolfi A, Gandhi S, Salek F, Christiaen L (2014) Tissue-specific genome editing in Ciona embryos by CRISPR/Cas9. Development 141(21):4115–4120
Tosches MA, Arendt D (2013) The bilaterian forebrain: an evolutionary chimaera. Curr Opin Neurobiol 23:1080–1089
Acknowledgments
Marine Joly is warmly acknowledged for help and discussion on the talk and manuscript. Pierre Boudinot contributed a lot to the improvement of earlier versions of this manuscript. I wish to thank T. Momose for sharing unpublished data and M. Manuel for discussion.
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2017 The Author(s)
About this chapter
Cite this chapter
Joly, JS. (2017). Aquatic Model Organisms in Neurosciences: The Genome-Editing Revolution. In: Jaenisch, R., Zhang, F., Gage, F. (eds) Genome Editing in Neurosciences. Research and Perspectives in Neurosciences. Springer, Cham. https://doi.org/10.1007/978-3-319-60192-2_2
Download citation
DOI: https://doi.org/10.1007/978-3-319-60192-2_2
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-319-60191-5
Online ISBN: 978-3-319-60192-2
eBook Packages: Biomedical and Life SciencesBiomedical and Life Sciences (R0)